Methods For Targeting Membrane Steroid Transporters
YAMANAKA; Naoki ; et al.
U.S. patent application number 15/902948 was filed with the patent office on 2018-12-27 for methods for targeting membrane steroid transporters. This patent application is currently assigned to The Regents of the University of California. The applicant listed for this patent is The Regents of the University of California. Invention is credited to Riyan BITTAR, Naoki OKAMOTO, Naoki YAMANAKA, Sachiko YAMANAKA.
| Application Number | 20180372760 15/902948 |
| Document ID | / |
| Family ID | 64693059 |
| Filed Date | 2018-12-27 |






| United States Patent Application | 20180372760 |
| Kind Code | A1 |
| YAMANAKA; Naoki ; et al. | December 27, 2018 |
METHODS FOR TARGETING MEMBRANE STEROID TRANSPORTERS
Abstract
Described herein are methods of identifying compounds that can modulate the transport of a steroid hormone across a phospholipid membrane. Also described herein are methods of identifying compounds that can affect the transcriptional activity of a steroid hormone nuclear receptor.
| Inventors: | YAMANAKA; Naoki; (Riverside, CA) ; YAMANAKA; Sachiko; (Riverside, CA) ; OKAMOTO; Naoki; (Riverside, CA) ; BITTAR; Riyan; (South Pasadena, CA) | ||||||||||
| Applicant: |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Assignee: | The Regents of the University of
California Oakland CA |
||||||||||
| Family ID: | 64693059 | ||||||||||
| Appl. No.: | 15/902948 | ||||||||||
| Filed: | February 22, 2018 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 15883809 | ||||
| 15902948 | ||||
| 62453172 | Feb 1, 2017 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C12Q 1/66 20130101; G01N 33/5023 20130101; G01N 33/743 20130101; G01N 2500/04 20130101 |
| International Class: | G01N 33/74 20060101 G01N033/74 |
Goverment Interests
STATEMENT OF RIGHTS TO INVENTIONS MADE UNDER FEDERALLY SPONSORED RESEARCH
[0002] This invention was made with government support under Contract R00HD073239 awarded by the National Institutes of Health (NIH). The government has certain rights in the invention.
Claims
1. A method of identifying a compound that can modulate the transport of a steroid hormone across a phospholipid membrane, comprising: providing a steroid hormone to an initial cell and a comparative cell in either the presence or absence of a candidate compound, wherein the initial cell expresses a steroid hormone transporter gene, and the expression of the steroid hormone transporter gene in the comparative cell is absent or lower than in the initial cell; determining the amount of the steroid hormone that is transported across the phospholipid membrane of the initial cell and the comparative cell; observing a difference between the amount of steroid hormone that is transported across the phospholipid membrane of the initial cell in the presence of the candidate compound compared to the absence of the candidate compound; comparing the difference to the amount of steroid hormone that is transported across the phospholipid membrane of the comparative cell in the presence or absence of the candidate compound, and determining that the candidate compound can modulate the transport of a steroid hormone across a phospholipid membrane if the amount of steroid hormone that is transported across the phospholipid membrane is different in the initial cell than in the comparative cell, wherein the steroid hormone transporter gene is ecdysone importer (EcI).
2. The method of claim 1, wherein the difference in the amount of steroid hormone transported across the phospholipid membrane is determined using a luciferase reporter assay.
3. The method of claim 1, wherein the expression of the steroid hormone transporter gene in the comparative cell is absent, and the method further comprises transfecting the initial cell, the comparative cell, or both with cDNA comprising the steroid hormone transporter gene.
4. The method of claim 1, further comprising transfecting the initial cell, the comparative cell, or both with cDNA comprising a steroid hormone nuclear receptor gene.
5-8. (canceled)
9. The method of claim 4, wherein the steroid hormone nuclear receptor gene is ecdysone receptor (EcR).
10. The method of claim 1, wherein the initial cell and the comparative cell are from an arthropod cell line or a mammalian cell line.
11. (canceled)
12. The method of claim 1, wherein the steroid hormone is an ecdysteroid.
13. The method of claim 1, wherein the steroid hormone is 20-hydroxyecdysone.
14. A method of identifying a compound that can affect the transcriptional activity of a steroid hormone nuclear receptor, comprising: providing a steroid hormone to an initial cell and a comparative cell in either the presence or absence of a candidate compound, wherein the initial cell expresses a steroid hormone transporter gene, and the expression of the steroid hormone transporter gene in the comparative cell is absent or lower than in the initial cell; determining the transcriptional activity of the steroid hormone nuclear receptor in the initial cell and the comparative cell; observing a difference between the transcriptional activity in the initial cell in the presence of the candidate compound compared to the absence of the candidate compound; comparing the difference to the transcriptional activity of the comparative cell in the presence or absence of the candidate compound, and determining that the candidate compound can affect the transcriptional activity of a steroid hormone nuclear receptor if the amount of transcriptional activity is different in the initial cell than in the comparative cell, wherein the steroid hormone transporter gene is ecdysone importer (EcI).
15. The method of claim 14, wherein the difference in the transcriptional activity is determined using a luciferase reporter assay.
16. The method of claim 14, wherein the expression of the steroid hormone transporter gene in the comparative cell is absent, and the method further comprises transfecting the initial cell, the comparative cell, or both with cDNA comprising the steroid hormone transporter gene.
17. The method of claim 14, further comprising transfecting the initial cell, the comparative cell, or both with cDNA comprising the steroid hormone nuclear receptor gene.
18-21. (canceled)
22. The method of claim 14, wherein the steroid hormone nuclear receptor gene is ecdysone receptor (EcR).
23. The method of claim 14, wherein the initial cell and the comparative cell are from an arthropod cell line or a mammalian cell line.
24. (canceled)
25. The method of claim 14, wherein the steroid hormone is an ecdysteroid.
26. The method of claim 14, wherein the steroid hormone is 20-hydroxyecdysone.
27. The method of claim 1, wherein the expression of the steroid hormone transporter gene in the comparative cell is lower than in the initial cell.
28. The method of claim 27, wherein the expression of the steroid hormone transporter gene in the comparative cell is reduced using RNAi.
29. The method of claim 14, wherein the expression of the steroid hormone transporter gene in the comparative cell is lower than in the initial cell.
30. The method of claim 29, wherein the expression of the steroid hormone transporter gene in the comparative cell is reduced using RNAi.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation application of U.S. patent application Ser. No. 15/883,809, filed Jan. 30, 2018, which claims priority to and the benefit of U.S. Provisional Patent Application No. 62/453,172, filed Feb. 1, 2017, the disclosures of each of which are incorporated herein by reference in their entireties.
SUBMISSION OF SEQUENCE LISTING ON ASCII TEXT FILE
[0003] The content of the following submission on ASCII text file is incorporated herein by reference in its entirety: a computer readable form (CRF) of the Sequence Listing (file name: 677032001001SEQLIST.txt, date recorded: Feb. 13, 2018, size: 15 KB).
FIELD
[0004] The present disclosure relates generally to chemical compounds that influence the cellular uptake of steroid hormones, and more specifically to screening methods for identifying chemical compounds that can influence the transport of steroid hormones across cell membranes by steroid hormone transporters.
BACKGROUND
[0005] Steroid hormones are a group of hormones that regulate diverse physiological and pathological processes, including immune response, energy homeostasis, sexual maturation and cancer progression. They enter target cells and bind to intracellular nuclear receptors, which then function as transcription factors and regulate gene expression in the nucleus. Steroid hormones are lipophilic, and are generally believed to enter cells by simple diffusion across the lipid bilayer.
[0006] There is interest in controlling the effects of steroid hormones, for example to regulate insect growth, to control inflammation, for hormone replacement therapy, or for the treatment of certain cancers. This is often done through the administration of synthetic or natural steroid hormones or steroid hormone antagonists to a subject. However, the administration of steroid hormones or steroid hormone antagonists often leads to unwanted side effects in the subject. Thus, what is desired in the art is methods to regulate the effect of steroid hormones and steroid hormone signaling in organisms, such as arthropods and humans.
BRIEF SUMMARY
[0007] In one aspect, provided is a method of identifying a compound that can modulate the transport of a steroid hormone across a phospholipid membrane, by: [0008] providing a steroid hormone to an initial cell and a comparative cell in either the presence or absence of a candidate compound, wherein the initial cell expresses a steroid hormone transporter gene, and the expression of the steroid hormone transporter gene in the comparative cell is absent or lower than in the initial cell; [0009] determining the amount of the steroid hormone that is transported across the phospholipid membrane of the initial cell and the comparative cell; [0010] observing a difference between the amount of steroid hormone that is transported across the phospholipid membrane of the initial cell in the presence of the candidate compound compared to the absence of the candidate compound; and [0011] comparing the difference to the amount of steroid hormone that is transported across the phospholipid membrane of the comparative cell in the presence or absence of the candidate compound.
[0012] In some variations, the difference in the amount of steroid hormone transported across the phospholipid membrane is determined using a luciferase reporter assay.
[0013] In another aspect, provided is a method of identifying a compound that can affect the transcriptional activity of a steroid hormone nuclear receptor, by: [0014] providing a steroid hormone to an initial cell and a comparative cell in either the presence or absence of a candidate compound, wherein the initial cell expresses a steroid hormone transporter gene, and the expression of the steroid hormone transporter gene in the comparative cell is absent or lower than in the initial cell; [0015] determining the transcriptional activity of the steroid hormone nuclear receptor in the initial cell and the comparative cell; [0016] observing a difference between the transcriptional activity in the initial cell in the presence of the candidate compound compared to the absence of the candidate compound; and [0017] comparing the difference to the transcriptional activity of the comparative cell in the presence or absence of the candidate compound.
[0018] In some variations, the difference in the transcriptional activity is determined using a luciferase reporter assay.
[0019] In certain variations, any of the preceding methods further include transfecting the initial cell, the comparative cell, or both with cDNA which includes the steroid hormone transporter gene. In some variations, the methods further include transfecting the initial cell, the comparative cell, or both with cDNA which includes the steroid hormone nuclear receptor gene.
[0020] In some variations, the steroid hormone transporter gene is of the solute carrier (SLC) family, for example of the solute carrier organic anion (SLCO) family. In certain variations, the gene encodes an organic anion-transporting polypeptide (OATP). In certain variations, the gene has the cDNA sequence of SEQ ID NO: 1.
[0021] In some variations, the steroid hormone is an ecdysteroid, estrogen, corticosteroid, progestogen, or androgen.
DESCRIPTION OF THE FIGURES
[0022] The present application can be understood by reference to the following description taken in conjunction with the accompanying drawing figures, in which like parts may be referred to by like numerals.
[0023] FIG. 1A is a scheme of in vivo RNAi screening for a gene encoding a membrane transporter for 20-hydroxyecdysone (20E).
[0024] FIG. 1B is an image of in vivo RNAi screening in wandering stage larvae of D. melanogaster, showing green-fluorescent protein (GFP) signal in the salivary gland in the presence and absence of ecdysone importer (EcI) and ecdysone reporter (EcR) RNAi. The GFP signal is from glue-GFP, the expression of which in the salivary gland is typically induced by 20E during the wandering stage. fkh-Gal4 is a salivary gland-specific Gal4 driver.
[0025] FIG. 1C is an image of in vivo RNAi screening in the white prepupal stage of D. melanogaster, showing the distribution of fat body (FB) cells that typically migrate into the pupal head in response to 20E. FB cells are labeled with GFP; lack of FB cell migration into the pupal head is indicated by the arrowheads.
[0026] FIG. 1D is a graph showing the level of ecdysteroids in the FB or hemolymph of white prepupae with knockdown of EcI, knockdown of EcR, or no knockdown in the FB. Each bar represents mean.+-.SEM; * indicates p<0.05 from Student's t-test compared to control; ** indicates p<0.01 from Student's t-test compared to control.
[0027] FIG. 2A is an image of overexpression of Flag-HA-tagged EcI (CG-Gal4>UAS-Oatp74D-Flag-HA) in D. melanogaster FB cells. The middle panel depicts the signal from the cell nuclei (stained using Hoechst 33342), while the right panel depicts the signal from .alpha.-HA.
[0028] FIG. 2B is an image of overexpression of Flag-HA-tagged EcI (fkh-Gal4>UAS-Oatp74D-Flag-HA) in D. melanogaster salivary gland cells. The middle panel depicts the signal from the cell nuclei (stained using Hoechst 33342), while the right panel depicts the signal from .alpha.-HA.
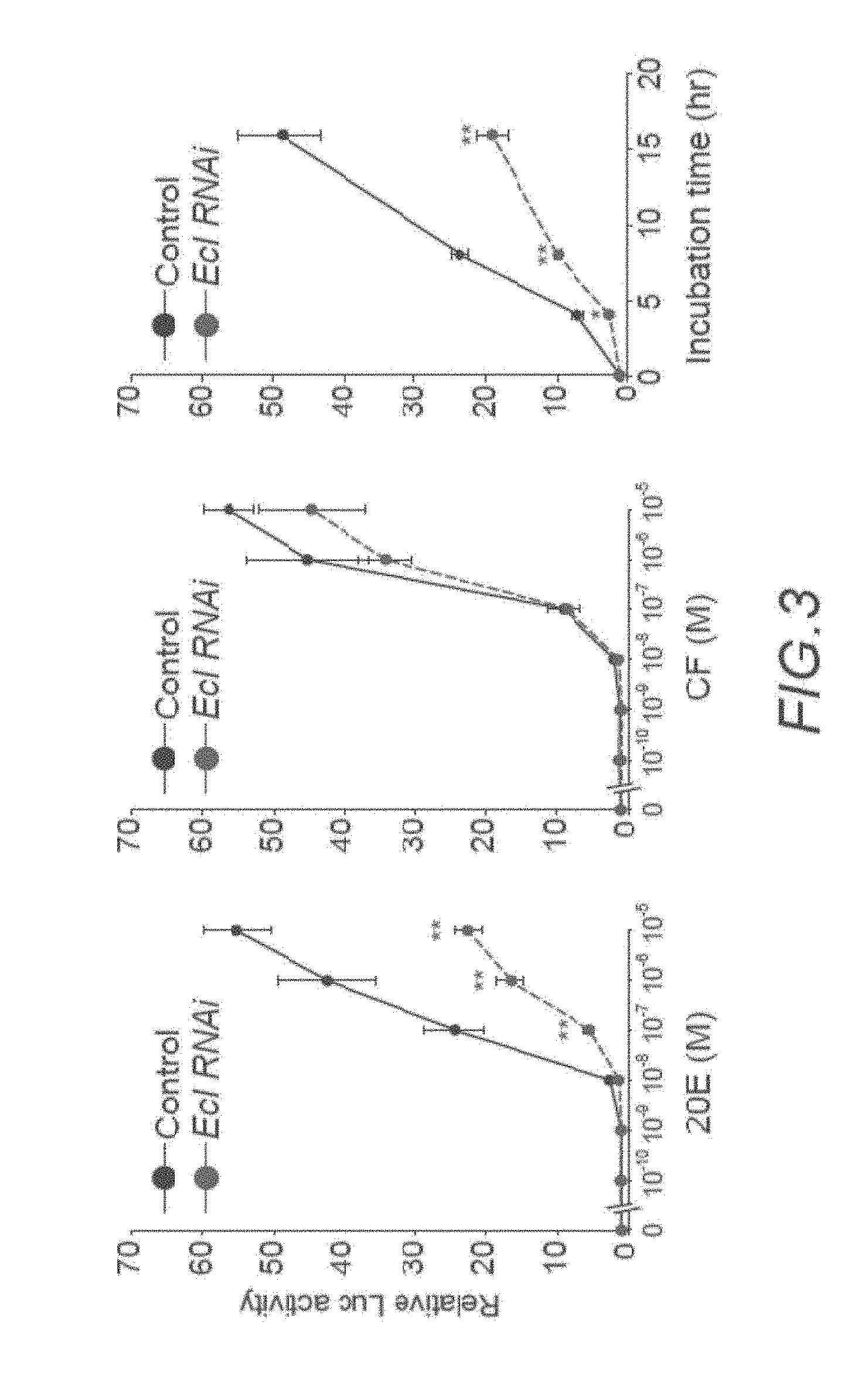
[0029] FIG. 3 is a graph of the fold change of D. melanogaster EcR response after incubation of S2 culture cells for 24 hours in varying concentrations of 20E (left graph) or chromafenozide (CF; middle graph), as measured by luciferase reporter assay. The right graph is a time-course analysis of EcR-mediated luciferase activity in S2 culture cells treated with 10.sup.-7 M of 20E. In each graph, the control data is the solid top line, and the dashed bottom line is data of cells treated with EcI RNAi. Each point represents mean.+-.SD; * indicates p<0.05 from Student's t-test compared to control; ** indicates p<0.01 from Student's t-test compared to control.
[0030] FIG. 4 is a relative luciferase reporter activity in HEK culture cells treated for 24 h with 20E. Response of the cells transfected with EcR is the solid bottom line; cells transfected with EcR and EcI is the dashed top line. Each point represents mean.+-.SEM; ** indicates p<0.01 from Student's t-test compared to control.
[0031] FIG. 5A depicts a schematic of a luciferase reporter assay in a cell derived from an arthropod cell line, for determining transport of 20E by the membrane steroid hormone transporter encoded by ecdysone importer (EcI).
[0032] FIG. 5B depicts a schematic of a luciferase reporter assay in a cell derived from a mammalian cell line, for determining transport of 20E by the membrane steroid hormone transporter encoded by ecdysone importer (EcI).
DETAILED DESCRIPTION
[0033] The following description sets forth numerous exemplary configurations, methods, parameters, and the like. It should be recognized, however, that such description is not intended as a limitation on the scope of the present disclosure, but is instead provided as a description of exemplary embodiments.
[0034] In some aspects, provided herein are methods of identifying chemical compounds that can modulate the effect of steroid hormones in organisms. After crossing the phospholipid bilayer into the cell, certain steroid hormones bind to nuclear receptors, which in turn affect the expression of genes in the organism. It has been surprisingly found that instead of passively diffusing through the cell membrane, cellular uptake of certain steroid hormones is mediated by steroid hormone transporter polypeptides. Modulating the transport of a steroid hormone across a cell membrane by a polypeptide transporter may provide an alternative method of controlling the effect of the steroid hormones and steroid hormone signaling in an organism. Thus, in certain aspects, provided herein are methods of identifying chemical compounds that can modulate the transport of a steroid hormone across a phospholipid membrane by a steroid hormone transporter. In other aspects, provided herein are also methods of identifying chemical compounds that can modulate the transcriptional activity of a steroid hormone nuclear receptor, for example by modulating the transport of a steroid hormone across a membrane by a steroid hormone transporter. In yet other aspects, provided are compounds identified by the methods described herein, suitable for use as hormone agonists or hormone antagonists (e.g., in hormone therapy).
Methods of Screening Candidate Compounds
[0035] In some aspects, the methods described herein include screening candidate compounds to identify one or more compounds that affect the transport of a steroid hormone across a cell membrane by a steroid hormone transporter.
[0036] In some embodiments, the transport of the steroid hormone is evaluated in an initial cell in the presence and absence of a candidate compound. In certain embodiments, the initial cell expresses a steroid hormone transporter gene. For example, a compound may be identified if the transport of a steroid hormone by a steroid hormone transporter in an initial cell is increased or decreased in the presence of the compound, compared to transport in the absence of the compound.
[0037] Thus, in one embodiment, the screening method includes providing a steroid hormone to an initial cell in the presence and absence of a candidate compound, wherein the initial cell expresses a steroid hormone transporter gene; determining the amount of the steroid hormone that is transported across the phospholipid membrane of the initial cell in the presence and absence of the candidate compound; and observing a difference between the amount of steroid hormone that is transported across the phospholipid membrane of the initial cell in the presence of the candidate compound compared to the absence of the candidate compound.
[0038] In some variations, the transport of the steroid hormone in the initial cell is compared to the transport of the steroid hormone in a comparative cell. In some embodiments, both the initial cell and the comparative cell express the same steroid hormone transporter gene. In certain embodiments, the expression of the transporter gene in the comparative cell is lower than the expression of the transporter gene in the initial cell. In other embodiments, the initial cell expresses a steroid hormone transporter gene, while the comparative cell does not express the same transporter gene.
[0039] In some embodiments, a chemical compound is identified if the transport of a steroid hormone by a steroid hormone transporter in the presence of a candidate compound is different in an initial cell (e.g., higher or lower) than in a comparative cell.
[0040] In other embodiments, a chemical compound is identified if the presence of the compound has a greater effect on the transport by a steroid hormone transporter of a steroid hormone in an initial cell than in the comparative cell.
[0041] In some embodiments, the transport of the steroid hormone is evaluated by measuring the transcriptional activity of a steroid hormone nuclear receptor. For example, in some variations, the transcriptional activity of a steroid hormone nuclear receptor is increased or decreased in the presence of a candidate compound, compared to the transcriptional activity in the absence of the candidate compound.
[0042] Thus, in one embodiment, the screening method includes providing a steroid hormone to an initial cell in the presence and absence of a candidate compound, wherein the initial cell expresses a steroid hormone transporter gene; determining the transcriptional activity of the steroid hormone nuclear receptor in the presence and absence of the candidate compound; and observing a difference between the transcriptional activity of the initial cell in the presence of the candidate compound compared to the absence of the candidate compound.
[0043] Thus, in certain embodiments, the initial cell expresses a steroid hormone nuclear receptor gene. In some embodiments, the comparative cell expresses the same steroid hormone nuclear receptor gene. In still other embodiments, the initial cell and the comparative cell, if present, express the same steroid hormone transporter gene and the same steroid hormone nuclear receptor gene.
Steroid Hormone Transporter Gene
[0044] The initial cell or comparative cell used in the methods described herein may express any suitable steroid hormone transporter gene. In some embodiments, the initial cell and comparative cell both express the same steroid hormone transporter gene.
[0045] In some embodiments, the steroid hormone transporter gene is a mammalian gene, such as a human gene. In other embodiments, the steroid hormone transporter gene is an arthropod gene, such as a D. melanogaster gene. Thus, included herein are methods of screening candidate compounds to identify one or more compounds that affect the transport of a steroid hormone by a steroid hormone transporter into a mammalian or arthropod cell. In some embodiments, the mammal is a human.
[0046] In some embodiments, the steroid hormone transporter gene is of the solute carrier (SLC) family. In some embodiments, the steroid hormone transporter gene is of the solute carrier organic anion (SLCO) family. In still other embodiments, the steroid hormone transporter gene encodes an organic anion-transporting polypeptide (OATP). For example, in one embodiment, the steroid hormone transporter gene is Oatp74D. In one embodiment, the steroid hormone transporter gene is ecdosyne importer (EcI). In one embodiment, the steroid hormone transporter gene Oatp74D is identical to the steroid hormone transporter gene ecdosyne importer (EcI). In some embodiments, the steroid hormone transporter gene has the cDNA sequence of SEQ ID NO: 1. In one embodiment, the steroid hormone transporter gene has a cDNA sequence as shown in Table 1A.
TABLE-US-00001 TABLE 1A SEQ ID cDNA Sequence No ecdosyne importer ("EcI", Oatp74D) of D. melanogaster 1 TTACCGTGTGCGGACTGTGCCGACGGGCAGCGGAACTCAGGCGGTGCAACTTGA GTGCTTGATAAATTTGTTGTATTGAAACCCAACCGCAAATACAATTTTCAATTGG CTGAGTGCGTGTCTTCAACGCACAAACATACGAATATAGCATTTAATATTGTTCA TTATGTGTGCAATTAAATAAAAAAAATTGTGTGTGAAATAACAGAAAAACGAGT GCGGCGAACAGAGGCAGCAACAAATTAAACGCGTAAATTGCGTAGCAAATTCTG AATGAAAAAATTCAACAAAACCAGAAAGCGAAACAGGAGCGATAAGCTTACCA ACTAAAGAAACTCACGCCTAACAGAAATACACACCACTGCCATATTAAAGCAAA GAATATAGAACTGTTTTACAGACCAGATCAACCAATCTACCTCGACTTCTGGGCC AACAAGCAAAATAAAACTAAACTCAATGCCAGAACCCAATTTCCCGATAGGAAA ATGGAACAGGCAATAAGCAAGACCACATAGAGGAGATTAACTAAACGGATCCG TTGAGATTACACAGTTGAGATCAATAAAACCCATTGAAGATCGATCAAACCATC TAGTTTTTTAGAGTTCCATTAGAAGCCAAAATGACGAAGAGCAATGGCGATGTG GAGGCGGCAGCCCAGGTGCAATCTCTGGGCGGAAAGCCCAGCAACGGACATGG CCAGCTGAATGGGAATGGCTATCATCAGAACGGCGGACGCAGGGATTCCAGTCA AGCCTTCACACCACTGCTGTCGCAGCACAATAATGGCACCACCAATGGCGAGGT GACCACTCCGCCTCCGAGTACAGTGCTCTACGAGAGCACCCCGAGCAATAATAA CGAGTGGAAGGCCCCGGAGGATCTGGGACACCTGAAGAATGGCCTGGGCAACAT ACTGAGCAGTAATAATAATGGCACCGGCAATGGGCACAGTCTGAGCGAAAAGTA TGCCCATGAACAGGCTCCCCTGACCGGAGGTTACAAGTTGCCACCTCGCTCCAGT GAGTCCGAGGAATCCGATTTCGATTCCGACCTCAATGGCGGCTCCTCCGCGGAAT CCAGCTCCAGTTGCGGCCTTTTCGGCTGTCGGCCCAGGTGGGCCAGAAGATTCGC GTCCACGCACGTCTTCATGGTGGTCTTCCTGCTGGCCTACATCCTGCAGGGCATG TACATGACCTACTTCGTCTCTGTGATCACCACCATTGAGAAGCTATTCCAGATCA AGTCGAAGACCACGGGAATTCTGCTGAGCGCCAGTGAGATGGGTCAGATATGCA CGGCCATGTTGCTGACCTACTTCGCCGGCAGAGGACATCGTCCGAGATGGATTGC CTGCGGCATGGTCCTGTTCTCAATCGCCGCCTTCTCCTGCGCCCTGCCGCATTTCA TCTTCGGCGAGCAGCTGATGCACTCCAGTGTAATTCTGCAGCAGACGCAGGTCTC CCCCCCCAATAATTCCTTCTCATCACACTGGCTGAATGCCAGCAGTGAACAGGTT AATCCCAATTTGTGCATTTTGGGTGGCAACCAAACCCATTCGGGCAGCGAGTGCA ACGAGGAGCGCCAGCTGGAACAGGCCTCCCACTCTAAGATCACCGTCATCGTGC TGTGCATCTTCTTTGGCAGCCTGCTCAGCTCGGGCATTGGCCAGACCGCCGTGGC CACACTGGGCATACCCTACATCGATGACAATGTAGGCAGCAAGCAGTCGCCCAT GTACATGGCCGTCACCATTGGCATGAGGATCCTGGGACCGGCATCCGGTTTCATT TTTGGCAGCTTCTGCACTCGCTGGTATGTCAACTTCTCGAATCCCGGCTTCGACG CCACGGATCCGCGCTGGATTGGCGCCTGGTGGCTGGGGCCTGTGGCCATTGGCA GCCTCATGCTGCTGGCCTCCATCGCCATGTTCTCGTTTCCCAAGCAGTTGAGAGG CAAGCAGAAGCCGCCGGGGCAGACAGCAACTCCAGCAGCTCCAGTTGAGCCGG AGGAGAAGCCCAAGCTAAAAGATTTTCCCAAGACAGTCCGTCGCCAGCTGAGCA ACGACATCCTGATGTTCCGCACCGCCTCGTGCGTGTTCCACCTGCTGCCCATCGC CGGTCTCTATACGTTCCTGCCCAAGTATCTGGAGACGCAGTTCCGGCTGGCCACC TATGATGCCAACATGATCGCCGCCTTCTGTGGCATCCTGGTCATGGGCATAGGTA TTGTCATTTCCGGGCTCTTCATCCTGAAGCGAAAGCCCACTGCCAGGGGCGTGGC CGCCTGGATCGCCTTTACAGCCCTCGTCTACTCGGCGGGCATGATCATCCTGATG TTCATCGGCTGCAGCATGAACGACTTTGCCGGCTACAAGCCAAGCGATGGCAAC AGTCCCGCCTTGATCGAGCCCACGTGCAGTGCCGCTCTCAACTGTACCTGTGATA AGGAGAACTTCGCGCCCATCTGCGCCGACGGCAAAATGTACATCTCGGCCTGCC ACGCCGGATGCAGCAGCTCCTCACTGCGGCCCAGCGACAATCGCACACTCTACT CCGATTGTGCTTGCATTCCAGATGCTCCGGAGGCGGTCAACGGTTACTGTGATAA TAACTGCAAGAACTTCATCTACTTTATACTGATCTTTGCCATTTGCGTATTTATGC ATTCCACCTCCGAGGTGGGCAGCATGCTGCTCGTCATGCGCTGCACACACCCCAA AGATAAGGCCATGGCCATGGGTGTGATACAGTCGGCCATCGGCCTGTTCGGCAA CGTTCCCTGTCCCATCATCTACGGCGCAGTGGTGGACTCCGCCTGCCTCATCTGG AAGTCGGTGTGCGGCAAGCACGGCGCCTGTTCACTCTACGATGCGGACACTTTCC GGCAATATTTCCTAGGAATCACGGCTGGCATTATGTTCCTGGCATTCCTGATGGA TCTGGTGGTGTGGCGCAAGGCGCATCGCATTGACATCGCGCCCGAGGATCCGCA GGAGGGCGGGCCCGCTTCCAACGGAAGGACCTTGGAGGTGTCCGAGTCCAAGCA GCCCATCACCCCGGCGCCGGACACGACGGTCTAGGAGGAGCGGGTCGGCGACGA GCCTTGCACAAGCTGCAGGATTTCCAATAAACGTTTACCTTAATTGTTAATTAGT TATCATTTGTCTAGTTTGTTAGCCTAGTGCAAGAGTTATGTATTTAGTTAAGTGGC ATCTTCGAGCGTCGGGAGACCTCATTCAAATCCACATTAGAACGCGCTCGTCCAG CTCCCGCTCCCGATCCTGCTCCAAATCCCAGCACCATATTCTACATGAAGCCCAT GGATTGCGATTTGAATCCTTGTAAATCTCAACGCGAAACAATGAAATGAAATTCT AGACATTATCTAACGTCATGCATGAGCGTAGTTAATCGACGAGCTAATACTACA AACTGATCGACAATTGTGCAAAGTGCAAGAAAATTTATGAATCCTTATGTGTAA ACTATATGTAACAGTTATATCGCGATGTATGTAATCTATAATTAATATATTAGAC ATACACTTACATGTATGTATGTAATTGCAACCTATCTGTGGTAGTTAAATTTTAGT TAAAATTCAAATTAATTGTCTAAGTTTGTCATACAACAAATATATACGAAAAGAA ATTAAATGGAAAACTATATACAAGTAAAAAAAAAAAAAAAAAAA
[0047] In some embodiments, the steroid hormone transporter gene has the coding sequence of SEQ ID NO: 2. In one embodiment, the steroid hormone transporter gene has a coding sequence as shown in Table 1B.
TABLE-US-00002 TABLE 1B SEQ ID Coding Sequence No ecdosyne importer ("EcI", Oatp74D) of D. melanogaster 2 ATGACGAAGAGCAATGGCGATGTGGAGGCGGCAGCCCAGGTGCAATCTCTGGGC GGAAAGCCCAGCAACGGACATGGCCAGCTGAATGGGAATGGCTATCATCAGAAC GGCGGACGCAGGGATTCCAGTCAAGCCTTCACACCACTGCTGTCGCAGCACAAT AATGGCACCACCAATGGCGAGGTGACCACTCCGCCTCCGAGTACAGTGCTCTAC GAGAGCACCCCGAGCAATAATAACGAGTGGAAGGCCCCGGAGGATCTGGGACA CCTGAAGAATGGCCTGGGCAACATACTGAGCAGTAATAATAATGGCACCGGCAA TGGGCACAGTCTGAGCGAAAAGTATGCCCATGAACAGGCTCCCCTGACCGGAGG TTACAAGTTGCCACCTCGCTCCAGTGAGTCCGAGGAATCCGATTTCGATTCCGAC CTCAATGGCGGCTCCTCCGCGGAATCCAGCTCCAGTTGCGGCCTTTTCGGCTGTC GGCCCAGGTGGGCCAGAAGATTCGCGTCCACGCACGTCTTCATGGTGGTCTTCCT GCTGGCCTACATCCTGCAGGGCATGTACATGACCTACTTCGTCTCTGTGATCACC ACCATTGAGAAGCTATTCCAGATCAAGTCGAAGACCACGGGAATTCTGCTGAGC GCCAGTGAGATGGGTCAGATATGCACGGCCATGTTGCTGACCTACTTCGCCGGC AGAGGACATCGTCCGAGATGGATTGCCTGCGGCATGGTCCTGTTCTCAATCGCCG CCTTCTCCTGCGCCCTGCCGCATTTCATCTTCGGCGAGCAGCTGATGCACTCCAG TGTAATTCTGCAGCAGACGCAGGTCTCCCCCCCCAATAATTCCTTCTCATCACAC TGGCTGAATGCCAGCAGTGAACAGGTTAATCCCAATTTGTGCATTTTGGGTGGCA ACCAAACCCATTCGGGCAGCGAGTGCAACGAGGAGCGCCAGCTGGAACAGGCCT CCCACTCTAAGATCACCGTCATCGTGCTGTGCATCTTCTTTGGCAGCCTGCTCAG CTCGGGCATTGGCCAGACCGCCGTGGCCACACTGGGCATACCCTACATCGATGA CAATGTAGGCAGCAAGCAGTCGCCCATGTACATGGCCGTCACCATTGGCATGAG GATCCTGGGACCGGCATCCGGTTTCATTTTTGGCAGCTTCTGCACTCGCTGGTAT GTCAACTTCTCGAATCCCGGCTTCGACGCCACGGATCCGCGCTGGATTGGCGCCT GGTGGCTGGGGCCTGTGGCCATTGGCAGCCTCATGCTGCTGGCCTCCATCGCCAT GTTCTCGTTTCCCAAGCAGTTGAGAGGCAAGCAGAAGCCGCCGGGGCAGACAGC AACTCCAGCAGCTCCAGTTGAGCCGGAGGAGAAGCCCAAGCTAAAAGATTTTCC CAAGACAGTCCGTCGCCAGCTGAGCAACGACATCCTGATGTTCCGCACCGCCTC GTGCGTGTTCCACCTGCTGCCCATCGCCGGTCTCTATACGTTCCTGCCCAAGTATC TGGAGACGCAGTTCCGGCTGGCCACCTATGATGCCAACATGATCGCCGCCTTCTG TGGCATCCTGGTCATGGGCATAGGTATTGTCATTTCCGGGCTCTTCATCCTGAAG CGAAAGCCCACTGCCAGGGGCGTGGCCGCCTGGATCGCCTTTACAGCCCTCGTCT ACTCGGCGGGCATGATCATCCTGATGTTCATCGGCTGCAGCATGAACGACTTTGC CGGCTACAAGCCAAGCGATGGCAACAGTCCCGCCTTGATCGAGCCCACGTGCAG TGCCGCTCTCAACTGTACCTGTGATAAGGAGAACTTCGCGCCCATCTGCGCCGAC GGCAAAATGTACATCTCGGCCTGCCACGCCGGATGCAGCAGCTCCTCACTGCGG CCCAGCGACAATCGCACACTCTACTCCGATTGTGCTTGCATTCCAGATGCTCCGG AGGCGGTCAACGGTTACTGTGATAATAACTGCAAGAACTTCATCTACTTTATACT GATCTTTGCCATTTGCGTATTTATGCATTCCACCTCCGAGGTGGGCAGCATGCTG CTCGTCATGCGCTGCACACACCCCAAAGATAAGGCCATGGCCATGGGTGTGATA CAGTCGGCCATCGGCCTGTTCGGCAACGTTCCCTGTCCCATCATCTACGGCGCAG TGGTGGACTCCGCCTGCCTCATCTGGAAGTCGGTGTGCGGCAAGCACGGCGCCT GTTCACTCTACGATGCGGACACTTTCCGGCAATATTTCCTAGGAATCACGGCTGG CATTATGTTCCTGGCATTCCTGATGGATCTGGTGGTGTGGCGCAAGGCGCATCGC ATTGACATCGCGCCCGAGGATCCGCAGGAGGGCGGGCCCGCTTCCAACGGAAGG ACCTTGGAGGTGTCCGAGTCCAAGCAGCCCATCACCCCGGCGCCGGACACGACG GTCTAG
[0048] In some embodiments, the steroid hormone transporter gene has the amino acid sequence of SEQ ID NO: 3. In one embodiment, the steroid hormone transporter gene has an amino acid sequence as shown in Table 1C.
TABLE-US-00003 TABLE 1C SEQ ID Amino Acid Sequence No ecdosyne importer ("EcI", Oatp74D) of D. melanogaster 3 MTKSNGDVEAAAQVQSLGGKPSNGHGQLNGNGYHQNGGRRDSSQAFTPLLSQHNN GTTNGEVTTPPPSTVLYESTPSNNNEWKAPEDLGHLKNGLGNILSSNNNGTGNGHSL SEKYAHEQAPLTGGYKLPPRSSESEESDFDSDLNGGSSAESSSSCGLFGCRPRWARRF ASTHVFMVVFLLAYILQGMYMTYFVSVITTIEKLFQIKSKTTGILLSASEMGQICTAM LLTYFAGRGHRPRWIACGMVLFSIAAFSCALPHFIFGEQLMHSSVILQQTQVSPPNNS FSSHWLNASSEQVNPNLCILGGNQTHSGSECNEERQLEQASHSKITVIVLCIFFGSLLS SGIGQTAVATLGIPYIDDNVGSKQSPMYMAVTIGMRILGPASGFIFGSFCTRWYVNFS NPGFDATDPRWIGAWWLGPVAIGSLMLLASIAMFSFPKQLRGKQKPPGQTATPAAP VEPEEKPKLKDFPKTVRRQLSNDILMFRTASCVFHLLPIAGLYTFLPKYLETQFRLAT YDANMIAAFCGILVMGIGIVISGLFILKRKPTARGVAAWIAFTALVYSAGMIILMFIGC SMNDFAGYKPSDGNSPALIEPTCSAALNCTCDKENFAPICADGKMYISACHAGCSSS SLRPSDNRTLYSDCACIPDAPEAVNGYCDNNCKNFIYFILIFAICVFMHSTSEVGSMLL VMRCTHPKDKAMAMGVIQSAIGLFGNVPCPIIYGAVVDSACLIWKSVCGKHGACSL YDADTFRQYFLGITAGIMFLAFLMDLVVWRKAHRIDIAPEDPQEGGPASNGRTLEVS ESKQPITPAPDTTV
[0049] In some embodiments, the steroid hormone transporter gene is introduced into the initial cell, or the comparative cell, or both. The gene may be introduced using any suitable methods, such as transfection. In some embodiments, the initial cell is transfected with cDNA encoding the steroid hormone transporter gene. In some embodiments, the comparative cell is transfected with cDNA encoding the steroid hormone transporter gene.
[0050] In some embodiments, the comparative cell has lower expression or no expression of the steroid hormone transporter gene. In one embodiment, expression of the transporter gene in the comparative cell is decreased through RNA interference (RNAi).
Steroid Hormone Nuclear Receptor
[0051] In some variations, the methods include measuring the transcriptional activity of a steroid hormone nuclear receptor. Thus, in certain embodiments, the initial cell, the comparative cell (if present), or both cells express a steroid hormone nuclear receptor gene. In some embodiments, the steroid hormone nuclear receptor gene encodes a receptor that binds to one or more ecdysteroids (such as 20E), estrogens (such as estradiol), corticosteroids (such as cortisol), progestogens (such as progesterone), or androgens (such as testosterone), or any combinations thereof. In some embodiments, the steroid hormone nuclear receptor gene encodes a receptor that binds to 20-hydroxyecdysone, estradiol, cortisol, progesterone, or testosterone.
[0052] In one embodiment, the steroid hormone nuclear receptor gene is ecdysone receptor (EcR), estrogen receptor 1 (ESR1 or NR3A1), estrogen receptor 2 (ESR2 or NR3A2), glucocorticoid receptor (GR or NR3C1), progesterone receptor (PR or NR3C3) or androgen receptor (AR or NR3C4). In one embodiment, the steroid hormone nuclear receptor gene is ecdysone receptor (EcR).
[0053] In some embodiments, the steroid hormone nuclear receptor gene is introduced into the initial cell, or the comparative cell, or both. The gene may be introduced using any suitable methods, such as transfection. In some embodiments, the initial cell is transfected with cDNA encoding the steroid hormone nuclear receptor gene. In some embodiments, the comparative cell is transfected with cDNA encoding the steroid hormone nuclear receptor gene.
[0054] In some embodiments, the comparative cell has lower expression or no expression of the steroid hormone nuclear receptor gene. In one embodiment, expression of the nuclear receptor gene in the comparative cell is decreased through RNA interference (RNAi).
Transcriptional Activity
[0055] As described above, in some embodiments, the transcriptional activity of a steroid hormone nuclear receptor is used to evaluate the transport of a steroid hormone across the cell membrane of an initial cell, or a comparative cell. In some variations, the luciferase reporter assay is used to measure the transcriptional activity of the steroid hormone nuclear receptor. For example, with reference to FIG. 5A and FIG. 5B, depicted are schemes showing the use of a luciferase reporter assay in an S2 cell (FIG. 5A) or in a HEK cell (FIG. 5B) to evaluate the transcriptional activity of the nuclear receptor encoded by the gene ecdysone receptor (EcR).
Cell Types
[0056] The screening methods described herein may be carried out using any appropriate type of cell or cell line. In some embodiments, a heterologous expression system is used.
[0057] In certain embodiments, one or more cells or cell culture lines derived from an animal are used. In some embodiments, the one or more cells or cell culture lines are derived from a mammal, an arthropod, or an amphibian. For example, in some variations, the screening methods include the use of one or more cells or cell culture lines derived from humans, an organism of the genus Drosophila, or an organism of the genus Xenopus. In some embodiments, the methods include the use of human embryonic kidney (HEK) cells, CHO cells, HeLa cells, cells from D. melanogaster (such as Schneider 2 (S2) cells), or Xenopus oocytes. Thus, for example, in some embodiments the initial cell is a HEK cell, CHO cell, HeLa cell, S2 cell, or Xenopus oocyte. In some embodiments, the comparative cell is a HEK cell, CHO cell, HeLa cell, S2 cell, or Xenopus oocyte. In some embodiments, the initial cell and the comparative cell are the same type of cell. In some embodiments, the initial cell is from D. melanogaster. In some embodiments, the comparative cell is from D. melanogaster.
[0058] In some embodiments, the initial cell and the comparative cell are the same type of cell, but differ in expression of the steroid hormone transporter gene. In some embodiments, the initial cell and the comparative cell are cells or cell culture lines derived from the same species (for example, cells or cell culture lines derived from humans, an organism of the genus Drosophila, or an organism of the genus Xenopus), and the comparative cell has lower expression of the steroid hormone transporter gene than the initial cell. In some embodiments, the initial cell and the comparative cell are cells or cell culture lines derived from the same species (for example, cells or cell culture lines derived from humans, an organism of the genus Drosophila, or an organism of the genus Xenopus), the initial cell expresses the steroid hormone transporter gene, and the comparative cell has no expression of the steroid hormone transporter gene. In some embodiments, the initial cell and the comparative cell are derived from the same cells or cell culture line (for example, human embryonic kidney (HEK) cells, CHO cells, HeLa cells, cells from D. melanogaster (such as Schneider 2 (S2) cells), or Xenopus oocytes), and the comparative cell has lower expression of the steroid hormone transporter gene than the initial cell. In some embodiments, the initial cell and the comparative cell are derived from the same cell or cell culture line (for example, human embryonic kidney (HEK) cells, CHO cells, HeLa cells, cells from D. melanogaster (such as Schneider 2 (S2) cells), or Xenopus oocytes), the initial cell expresses the steroid hormone transporter gene, and the comparative cell has no expression of the steroid hormone transporter gene.
Steroid Hormones
[0059] The methods described herein may be used to evaluate the transport of any suitable steroid hormone. In some embodiments, the steroid is an ecdysteroid (such as 20E), estrogen (such as estradiol), corticosteroid (such as cortisol), progestogen (such as progesterone), or androgen (such as testosterone). In some embodiments, the steroid hormone is 20-hydroxyecdysone, estradiol, cortisol, progesterone or testosterone.
EXAMPLES
[0060] The following Examples are merely illustrative and are not meant to limit any aspects of the present disclosure in any way.
Example 1
Identification of Ecdysone Importer (EcI)
[0061] The gene ecdysone importer was identified using an in vivo RNAi screen (FIG. 1A).
[0062] 537 candidate genes were selected based on "gene ontology" (GO) terms, and screened for their knockdown effects on ecdysone-dependent glue-GFP expression in the salivary gland (SG) of D. melanogaster. From this screen, 51 RNAi lines were found to diminish GFP signal in the SG. These were further tested for their effect on ecdysone-dependent cell migration in the fat body (FB) of flies, which yielded a single hit, Oatp74D (CG7571). This gene was renamed ecdysone importer (EcI). The 537 candidate genes and the screening outcome are shown in Table 2.
TABLE-US-00004 TABLE 2 fat body glue-GFP cell Stock RNAi Stock expression migration # Symbol Center ID# defect* defect** 459 Oatp74D VDRC 37295 YES YES 7 Ant2 VDRC 102533 YES NO 40 CG10804 BDSC 29599 YES NO 43 CG10920 BDSC 44650 YES NO 84 CG13743 VDRC 40974 YES NO 166 CG31229 VDRC 106764 YES NO 243 CG4797 VDRC 10598 YES NO 277 CG6356 BDSC 28745 YES NO 361 Cog3 VDRC 108574 YES NO 385 emb BDSC 34021 YES NO 390 exo70 BDSC 55234 YES NO 392 fabp/sea BDSC 34685 YES NO 437 msk BDSC 33626 YES NO 483 Rph BDSC 25950 YES NO 506 Surf6 BDSC 34563 YES NO 527 vib VDRC 26848 YES NO 395 foi BDSC 55281 YES NO 528 Vmat BDSC 44471 YES NO 73 CG12858 BDSC 43284 YES NO 190 CG32091 BDSC 57783 YES NO 8 AP-1gamma BDSC 27533 YES NO 520 Ucp4B VDRC 33130 YES NO 27 cdm BDSC 44551 YES NO 176 CG3168 BDSC 29301 YES NO 258 CG5549 VDRC 8222 YES NO 326 CG8713 BDSC 25853 YES NO 118 CG1703 VDRC 105998 YES NO 10 AP-2sigma BDSC 27322 YES NO 59 CG11779 BDSC 43155 YES NO 278 CG6404 VDRC 108091 YES NO 219 CG42260 BDSC 26723 YES NO 170 CG3149 BDSC 53324 YES NO 174 CG3164 BDSC 57478 YES NO 391 Exp6 VDRC 34737 YES NO 151 CG2930 BDSC 54827 YES NO 431 mge BDSC 57430 YES NO 209 CG34120 BDSC 34596 YES NO 269 CG6006 BDSC 55282 YES NO 462 Orct BDSC 60125 YES NO 126 CG17646 BDSC 26008 YES NO 364 cpx BDSC 42017 YES NO 432 mnd VDRC 110217 YES NO 41 CG10864 BDSC 25878 YES NO 242 CG4743 BDSC 56025 YES NO 294 CG7309 BDSC 44653 YES NO 367 Ctr1A BDSC 44000 YES NO 393 Fatp BDSC 55919 YES NO 398 g VDRC 41369 YES NO 56 CG11659 BDSC 55268 YES NO 363 Cpr47Eg BDSC 56004 YES NO 470 Pen BDSC 27692 YES NO 421 Mdr49 BDSC 32405 NO ND 1 ABCB7 BDSC 51696 NO ND 2 AChT BDSC 27684 NO ND 3 Afti VDRC 104750 NO ND 4 alpha-Adaptin VDRC 15565 NO ND 5 alphaKap4 VDRC 108143 NO ND 6 Ank2 BDSC 33414 NO ND 9 AP-1sigma BDSC 40895 NO ND 11 AP-47 VDRC 24017 NO ND 12 AP-50 VDRC 103390 NO ND 13 AQP BDSC 51504 NO ND 14 aralar1 BDSC 56884 NO ND 15 Atet BDSC 57796 NO ND 16 ATP7 BDSC 31083 NO ND 17 Atpalpha BDSC 32913 NO ND 18 att-ORFA VDRC 50707 NO ND 19 bai VDRC 100612 NO ND 20 bap BDSC 27061 NO ND 21 bdg VDRC 110054 NO ND 22 blot VDRC 101083 NO ND 23 Bmcp BDSC 37498 NO ND 24 btsz BDSC 40898 NO ND 25 bw VDRC 101584 NO ND 26 Catsup BDSC 55396 NO ND 28 cert BDSC 60080 NO ND 29 CG10006 VDRC 101031 NO ND 30 CG10019 VDRC 7292 NO ND 31 CG10026 VDRC 107452 NO ND 32 CG10226 VDRC 108196 NO ND 33 CG10237 VDRC 108388 NO ND 34 CG10300 VDRC 109472 NO ND 35 CG10413 BDSC 38364 NO ND 36 CG10444 VDRC 107008 NO ND 37 CG10486 VDRC 107903 NO ND 38 CG10505 BDSC 38317 NO ND 39 CG10657 VDRC 103376 NO ND 42 CG1090 BDSC 25806 NO ND 44 CG10950 VDRC 41462 NO ND 45 CG10960 BDSC 34598 NO ND 46 CG11069 VDRC 101080 NO ND 47 CG11147 BDSC 57741 NO ND 48 CG11163 VDRC 107931 NO ND 49 CG11262 VDRC 105920 NO ND 50 CG11374 VDRC 15562 NO ND 51 CG1139 VDRC 102363 NO ND 52 CG11391 BDSC 56947 NO ND 53 CG11407 VDRC 107610 NO ND 54 CG11537 BDSC 43180 NO ND 55 CG11655 VDRC 9131 NO ND 57 CG11665 BDSC 52902 NO ND 58 CG11739 BDSC 38230 NO ND 60 CG11897 BDSC 38318 NO ND 61 CG11898 VDRC 100660 NO ND 62 CG12004 VDRC 101732 NO ND 63 CG12027 VDRC 108418 NO ND 64 CG12061 BDSC 26248 NO ND 65 CG1213 VDRC 6487 NO ND 66 CG12194 VDRC 109287 NO ND 67 CG12201 VDRC 109013 NO ND 68 CG12376 VDRC 22882 NO ND 69 CG12490 BDSC 44511 NO ND 70 CG12531 VDRC 105771 NO ND 71 CG12773 VDRC 108667 NO ND 72 CG12783 BDSC 27678 NO ND 74 CG12866 BDSC 57427 NO ND 75 CG12926 VDRC 109580 NO ND 76 CG12943 BDSC 56990 NO ND 77 CG13189 VDRC 105650 NO ND 78 CG13223 VDRC 1683 NO ND 79 CG13248 BDSC 28761 NO ND 80 CG13384 BDSC 41703 NO ND 81 CG13426 VDRC 107528 NO ND 82 CG1358 VDRC 101453 NO ND 83 CG13646 BDSC 53701 NO ND 85 CG13793 VDRC 50167 NO ND 86 CG13794 VDRC 102583 NO ND 87 CG13795 VDRC 102250 NO ND 88 CG13796 BDSC 57251 NO ND 89 CG13801 VDRC 104206 NO ND 90 CG13893 VDRC 108011 NO ND 91 CG13907 VDRC 110059 NO ND 92 CG14040 VDRC 110668 NO ND 93 CG14160 VDRC 104744 NO ND 94 CG14511 VDRC 107875 NO ND 95 CG14605 VDRC 105310 NO ND 96 CG14606 VDRC 100903 NO ND 97 CG14691 BDSC 31986 NO ND 98 CG14743 VDRC 105880 NO ND 99 CG14744 VDRC 108697 NO ND 100 CG14767 BDSC 34679 NO ND 101 CG14855 BDSC 57569 NO ND 102 CG14856 VDRC 101004 NO ND 103 CG1494 BDSC 57777 NO ND 104 CG15096 VDRC 107275 NO ND 105 CG15221 BDSC 31888 NO ND 106 CG15279 BDSC 50583 NO ND 107 CG15406 VDRC 105077 NO ND 108 CG15408 BDSC 58239 NO ND 109 CG15553 BDSC 28016 NO ND 110 CG15828 VDRC 51937 NO ND 111 CG15890 BDSC 28768 NO ND 112 CG1607 BDSC 57797 NO ND 113 CG1628 BDSC 54464 NO ND 114 CG16700 VDRC 110058 NO ND 115 CG16727 BDSC 57434 NO ND 116 CG1688 BDSC 25809 NO ND 117 CG1698 VDRC 101947 NO ND 119 CG17119 BDSC 40823 NO ND 120 CG17139/40 BDSC 26007 NO ND 121 CG17167 BDSC 27245 NO ND 122 CG1718 BDSC 38329 NO ND 123 CG1724 VDRC 30315 NO ND 124 CG17300 VDRC 6521 NO ND 125 CG17637 VDRC 102424 NO ND 127 CG17662 VDRC 104067 NO ND 128 CG17664 BDSC 25924 NO ND 129 CG17751 VDRC 50914 NO ND 130 CG17752 VDRC 106787 NO ND 131 CG17922 VDRC 100882 NO ND 132 CG17929 VDRC 100093 NO ND 133 CG17930 VDRC 108912 NO ND 134 CG1801 VDRC 100868 NO ND 135 CG1824 VDRC 106935 NO ND 136 CG18281 BDSC 58222 NO ND 137 CG18317 VDRC 100807 NO ND 138 CG18324 BDSC 34720 NO ND 139 CG18327 VDRC 100920 NO ND 140 CG18347 VDRC 106319 NO ND 141 CG18418 VDRC 102109 NO ND 142 CG18788 VDRC 101610 NO ND 143 CG1907 BDSC 38998 NO ND 144 CG2003 BDSC 51699 NO ND 145 CG2177 VDRC 51083 NO ND 146 CG2187 VDRC 101065 NO ND 147 CG2316 BDSC 41984 NO ND 148 CG2616 VDRC 102630 NO ND 149 CG2663 VDRC 33548 NO ND 150 CG2893 VDRC 40987 NO ND 152 CG30194 VDRC 106140 NO ND 153 CG30265 BDSC 50731 NO ND 154 CG30272 VDRC 105475 NO ND 155 CG30339 VDRC 104772 NO ND 156 CG30344 VDRC 106870 NO ND 157 CG30345 BDSC 33994 NO ND 158 CG3036 VDRC 43179 NO ND 159 CG30392 VDRC 103584 NO ND 160 CG30394 VDRC 3470 NO ND 161 CG3091 VDRC 109832 NO ND 162 CG31100 VDRC 42627 NO ND 163 CG31103 BDSC 28359 NO ND 164 CG31106 BDSC 29582 NO ND 165 CG31121 VDRC 100046 NO ND 167 CG31262 BDSC 53970 NO ND 168 CG31272 BDSC 28030 NO ND 169 CG31321 VDRC 101856 NO ND 171 CG31547 VDRC 8552 NO ND 172 CG3156 BDSC 57727 NO ND 173 CG31636 VDRC 105018 NO ND 175 CG31668 VDRC 104093 NO ND 177 CG31693 VDRC 105558 NO ND 178 CG31731 VDRC 107135 NO ND 179 CG31787 VDRC 6372 NO ND 180 CG31792 BDSC 34942 NO ND 181 CG31793 BDSC 38319 NO ND 182 CG31826 VDRC 100639 NO ND 183 CG31860 VDRC 103398 NO ND 184 CG31904 BDSC 34677 NO ND 185 CG3191 VDRC 109512 NO ND 186 CG32053 VDRC 107225 NO ND 188 CG32079 VDRC 104454 NO ND 189 CG32081 VDRC 107023 NO ND 191 CG32103 VDRC 108078 NO ND 192 CG32164/65 BDSC 60487 NO ND 193 CG32250 VDRC 108323 NO ND 194 CG32669 VDRC 102859 NO ND 195 CG3285 VDRC 51060 NO ND 196 CG33124 VDRC 105570 NO ND 197 CG33181 BDSC 38334 NO ND 198 CG33233 VDRC 106897 NO ND 199 CG33234 VDRC 106792 NO ND 200 CG33281 VDRC 7273 NO ND 201 CG33282 VDRC 100325 NO ND 202 CG33514 VDRC 47462 NO ND 203 CG33934/Indy-2 BDSC 34891 NO ND 204 CG3394 BDSC 56032 NO ND 205 CG33965 VDRC 61198 NO ND 206 CG33966 VDRC 104101 NO ND 207 CG33970 BDSC 60135 NO ND 208 CG3409 VDRC 37139 NO ND 210 CG34396 BDSC 26011 NO ND 211 CG3476 BDSC 60107 NO ND
212 CG3649 BDSC 44549 NO ND 213 CG3690 BDSC 27688 NO ND 214 CG3790 VDRC 108223 NO ND 215 CG3823 VDRC 107872 NO ND 216 CG3876 VDRC 44201 NO ND 217 CG4019 VDRC 107980 NO ND 218 CG42235 BDSC 50724 NO ND 220 CG42269 VDRC 101600 NO ND 221 CG42340 BDSC 27257 NO ND 222 CG42514 VDRC 45643 NO ND 223 CG42575 VDRC 107624 NO ND 224 CG42594 BDSC 35006 NO ND 225 CG42816 VDRC 106262 NO ND 226 CG42825 VDRC 5203 NO ND 227 CG4288 VDRC 104145 NO ND 228 CG43066 BDSC 28069 NO ND 229 CG43155 BDSC 55299 NO ND 230 CG4324 VDRC 109433 NO ND 231 CG4334 VDRC 106785 NO ND 232 CG43672 VDRC 48781 NO ND 233 CG44098 VDRC 100059 NO ND 234 CG4459 VDRC 107022 NO ND 235 CG4462 VDRC 105566 NO ND 236 CG4465 VDRC 100202 NO ND 237 CG4476 BDSC 38930 NO ND 238 CG4520 VDRC 34862 NO ND 239 CG4562 BDSC 32842 NO ND 240 CG4607 VDRC 107219 NO ND 241 CG4630 VDRC 101254 NO ND 244 CG4822 BDSC 25894 NO ND 245 CG4830 BDSC 57155 NO ND 246 CG4991 BDSC 44450 NO ND 247 CG4995 VDRC 106173 NO ND 248 CG5002 BDSC 55359 NO ND 249 CG5078 BDSC 57852 NO ND 250 CG5130 VDRC 5390 NO ND 251 CG5142 VDRC 22017 NO ND 252 CG5254 BDSC 50623 NO ND 253 CG5348 VDRC 101003 NO ND 254 CG5398 VDRC 109717 NO ND 255 CG5404 VDRC 100953 NO ND 256 CG5498 VDRC 103711 NO ND 257 CG5535 VDRC 107030 NO ND 259 CG5592 VDRC 110489 NO ND 260 CG5646 BDSC 33360 NO ND 261 CG5687 VDRC 33262 NO ND 262 CG5789 BDSC 57570 NO ND 263 CG5802 VDRC 103753 NO ND 264 CG5805 BDSC 57571 NO ND 265 CG5850 VDRC 1456 NO ND 266 CG5853 BDSC 27668 NO ND 267 CG5958 VDRC 100038 NO ND 268 CG5973 VDRC 24998 NO ND 270 CG6052 VDRC 106738 NO ND 271 CG6125 VDRC 107894 NO ND 272 CG6126 BDSC 56038 NO ND 273 CG6231 VDRC 105194 NO ND 274 CG6293 VDRC 108619 NO ND 275 CG6299 VDRC 109737 NO ND 276 CG6300 BDSC 55264 NO ND 279 CG6484 BDSC 60372 NO ND 280 CG6499 VDRC 100416 NO ND 281 CG6565 VDRC 39006 NO ND 282 CG6672 VDRC 107388 NO ND 283 CG6723 BDSC 29425 NO ND 284 CG6812 VDRC 103925 NO ND 285 CG6836 VDRC 108502 NO ND 286 CG6893 VDRC 102093 NO ND 287 CG6901 VDRC 104673 NO ND 288 CG6928 BDSC 50552 NO ND 289 CG6978 VDRC 7041 NO ND 290 CG7084 BDSC 42767 NO ND 291 CG7091 VDRC 102183 NO ND 292 CG7255 BDSC 58352 NO ND 293 CG7272 BDSC 41950 NO ND 295 CG7333 BDSC 57433 NO ND 296 CG7342 VDRC 48001 NO ND 297 CG7442 BDSC 35817 NO ND 298 CG7448 VDRC 102236 NO ND 299 CG7458 BDSC 35237 NO ND 300 CG7514 VDRC 103023 NO ND 301 CG7627 BDSC 32337 NO ND 302 CG7708 BDSC 28613 NO ND 303 CG7777 BDSC 50695 NO ND 304 CG7806 BDSC 38997 NO ND 305 CG7816 VDRC 1364 NO ND 306 CG7881 BDSC 44505 NO ND 307 CG7882 VDRC 109918 NO ND 308 CG7888 VDRC 37263 NO ND 309 CG7912 VDRC 1378 NO ND 310 CG7943 VDRC 28632 NO ND 311 CG8008 VDRC 4159 NO ND 312 CG8026 VDRC 105681 NO ND 313 CG8028 VDRC 102317 NO ND 314 CG8034 BDSC 32340 NO ND 315 CG8046 VDRC 7380 NO ND 316 CG8219 VDRC 103487 NO ND 317 CG8249 VDRC 106077 NO ND 318 CG8323 VDRC 4861 NO ND 319 CG8389 VDRC 107639 NO ND 320 CG8451 VDRC 104177 NO ND 321 CG8468 VDRC 6452 NO ND 322 CG8596 BDSC 55664 NO ND 323 CG8602 VDRC 101575 NO ND 324 CG8632 VDRC 108929 NO ND 325 CG8654 BDSC 57428 NO ND 327 CG8785 VDRC 4651 NO ND 328 CG8791 VDRC 7945 NO ND 329 CG8837 VDRC 100670 NO ND 330 CG8850 VDRC 104098 NO ND 331 CG8860 BDSC 60127 NO ND 332 CG8908 BDSC 41850 NO ND 333 CG8925 VDRC 101128 NO ND 334 CG9090 BDSC 44495 NO ND 335 CG9194 BDSC 25921 NO ND 336 CG9254 VDRC 13409 NO ND 337 CG9270 BDSC 34029 NO ND 338 CG9281 BDSC 58189 NO ND 339 CG9308 VDRC 6606 NO ND 340 CG9317 BDSC 57730 NO ND 341 CG9330 BDSC 57795 NO ND 342 CG9396 VDRC 104068 NO ND 343 CG9399 VDRC 101455 NO ND 344 CG9413 BDSC 51791 NO ND 345 CG9430 VDRC 110047 NO ND 346 CG9444 VDRC 104529 NO ND 347 CG9582 VDRC 100256 NO ND 348 CG9657 BDSC 28384 NO ND 349 CG9663 VDRC 62192 NO ND 350 CG9664 VDRC 42467 NO ND 351 CG9702 VDRC 6860 NO ND 352 CG9706 BDSC 51908 NO ND 354 CG9825 BDSC 50639 NO ND 355 CG9826 BDSC 51728 NO ND 356 CG9864 VDRC 103970 NO ND 357 CG9903 VDRC 42689 NO ND 358 CG9990 VDRC 107544 NO ND 359 CHOp24 VDRC 100274 NO ND 360 cm BDSC 27282 NO ND 362 colt BDSC 51798 NO ND 365 Cralbp VDRC 31258 NO ND 366 Csat BDSC 44488 NO ND 368 Ctr1B BDSC 57710 NO ND 369 Ctr1C VDRC 102136 NO ND 370 cv-d VDRC 3975 NO ND 371 DAT BDSC 50619 NO ND 372 Dic1 VDRC 103757 NO ND 373 Dic2 VDRC 104152 NO ND 374 Dic3 VDRC 100255 NO ND 375 Dic4 VDRC 8416 NO ND 376 dmGlut BDSC 36724 NO ND 377 Drip BDSC 44661 NO ND 378 E23 BDSC 57782 NO ND 379 Eaat1 BDSC 43287 NO ND 380 Eaat2 BDSC 40832 NO ND 381 eag BDSC 31678 NO ND 382 eca VDRC 101388 NO ND 383 Efr VDRC 30238 NO ND 384 Elk BDSC 25821 NO ND 386 Ent1 BDSC 51055 NO ND 387 Ent2 VDRC 100464 NO ND 388 Ent3 VDRC 47537 NO ND 389 Esp VDRC 9795 NO ND 394 Fbp1 VDRC 37881 NO ND 396 frc BDSC 42007 NO ND 397 Fs(2)Ket BDSC 41845 NO ND 399 G31689 BDSC 57778 NO ND 400 galectin BDSC 34880 NO ND 401 gb VDRC 1262 NO ND 402 Gfr VDRC 105410 NO ND 403 Gga BDSC 51170 NO ND 404 glob1 BDSC 36668 NO ND 405 Glut1 BDSC 40904 NO ND 406 Glut3 VDRC 100253 NO ND 407 Hmt-1 BDSC 53284 NO ND 408 hoe1 BDSC 33377 NO ND 409 hoe2 BDSC 28661 NO ND 410 Ih BDSC 58089 NO ND 411 Indy VDRC 9981 NO ND 413 Jwa VDRC 6375 NO ND 414 kar VDRC 105429 NO ND 416 l(2)03659 VDRC 100105 NO ND 417 l(2)08717 VDRC 11117 NO ND 418 List VDRC 109791 NO ND 419 loj BDSC 57702 NO ND 420 Mct1 VDRC 106773 NO ND 422 Mdr50 BDSC 35034 NO ND 423 Mdr65 BDSC 35035 NO ND 424 mfrn BDSC 34038 NO ND 425 MFS10 BDSC 34562 NO ND 426 MFS15 VDRC 106207 NO ND 427 MFS16 VDRC 108635 NO ND 428 MFS17 BDSC 44033 NO ND 429 MFS18 BDSC 33998 NO ND 430 MFS3 VDRC 107656 NO ND 433 Mpcp BDSC 44508 NO ND 434 MRP BDSC 38316 NO ND 435 Mrp4 BDSC 60136 NO ND 436 MSBP VDRC 45185 NO ND 438 Mtch BDSC 38986 NO ND 439 Mtp BDSC 51872 NO ND 440 Mv1 BDSC 55316 NO ND 441 na BDSC 25808 NO ND 442 NAAT1 VDRC 50064 NO ND 443 NaPi-T BDSC 34003 NO ND 444 Ncc69 BDSC 28682 NO ND 445 Nckx30C BDSC 42581 NO ND 446 Ndae1 VDRC 102707 NO ND 447 Nha1 BDSC 42648 NO ND 448 Nhe1 BDSC 28589 NO ND 449 Nhe2 BDSC 51491 NO ND 450 Nhe3 BDSC 50513 NO ND 451 Ntl VDRC 102776 NO ND 452 Oatp26F VDRC 2650 NO ND 453 Oatp30B VDRC 22983 NO ND 454 Oatp33Ea BDSC 50736 NO ND 455 Oatp33Eb VDRC 100431 NO ND 456 Oatp58Da BDSC 44122 NO ND 457 Oatp58Db VDRC 100348 NO ND 458 Oatp58Dc BDSC 44583 NO ND 460 opm BDSC 43280 NO ND 461 or BDSC 55218 NO ND 463 Orct2 BDSC 57583 NO ND 464 out VDRC 108364 NO ND 465 p115 VDRC 44510 NO ND 466 p24-1 BDSC 57776 NO ND 467 p24-2 VDRC 109179 NO ND 468 para BDSC 33923 NO ND 469 path VDRC 100519 NO ND 471 PGRP-LC BDSC 33383 NO ND 472 Picot BDSC 25920 NO ND 473 PMCA BDSC 31572 NO ND 474 Pmp70 BDSC 34349 NO ND 475 Prestin BDSC 50706 NO ND 476 prt VDRC 104763 NO ND 477 rb BDSC 32477 NO ND 478 rdgB BDSC 28796 NO ND 479 rdgBbeta BDSC 44523 NO ND 480 retm VDRC 44687 NO ND 481 Rfabg BDSC 28946 NO ND 482 Rh50 BDSC 50666 NO ND 484 rtet VDRC 110473 NO ND 485 salt VDRC 108782 NO ND 486 ScpX BDSC 51479 NO ND 487 Sec24AB VDRC 107154 NO ND 489 SerT VDRC 100584 NO ND 490 sesB BDSC 36661 NO ND 491 Shawn VDRC 109948 NO ND 492 Slc45-1 VDRC 105439 NO ND 493 slif VDRC 45590 NO ND 494 sl1 VDRC 12148 NO ND
495 Sln VDRC 109464 NO ND 496 slo BDSC 55405 NO ND 497 slv BDSC 29388 NO ND 498 Smvt VDRC 102662 NO ND 499 spict BDSC 37505 NO ND 500 spin BDSC 27702 NO ND 501 st BDSC 60134 NO ND 502 Start1 VDRC 4053 NO ND 503 sug BDSC 55182 NO ND 504 Sur BDSC 36087 NO ND 505 Surf1 BDSC 51783 NO ND 507 sut1 VDRC 104983 NO ND 508 sut2 VDRC 28207 NO ND 509 sut3 VDRC 4009 NO ND 510 sut4 VDRC 44935 NO ND 511 TM9SF4 BDSC 54019 NO ND 512 Tpc1 BDSC 31749 NO ND 513 Tpc2 VDRC 107215 NO ND 514 Tret1-1 BDSC 42880 NO ND 515 Tret1-2 VDRC 49889 NO ND 516 Trn BDSC 50732 NO ND 517 Trn-SR BDSC 56974 NO ND 518 Tyler VDRC 50573 NO ND 519 Ucp4A VDRC 102571 NO ND 521 Ucp4C VDRC 100064 NO ND 522 VAChT BDSC 27684 NO ND 523 VGlut BDSC 40845 NO ND 524 VhaAC39-2 VDRC 34303 NO ND 525 VhaM9.7-c BDSC 26004 NO ND 526 VhaM9.7-d VDRC 106115 NO ND 529 w BDSC 33623 NO ND 530 wun2 BDSC 32423 NO ND 531 yin BDSC 31971 NO ND 532 ZIP1 VDRC 107309 NO ND 533 Zip3 VDRC 37358 NO ND 534 ZnT35C BDSC 39049 NO ND 535 ZnT63C VDRC 105145 NO ND 536 .beta.'COP BDSC 36113 NO ND 187 CG32054 VDRC 107214 Lethal ND 353 CG9717 VDRC 42669 Lethal ND 412 JhI-21 BDSC 41706 Lethal ND 415 kcc BDSC 34584 Lethal ND 488 Sec24CD VDRC 106929 Lethal ND 537 .zeta.COP BDSC 28960 Lethal ND *(in any animal observed) **(in all animals observed, with no other obvious defects) ND = not determined
Example 2
Study of EcI in D. melanogaster Cells
[0063] Using RNAi, the role of EcI in the transport of 20-hydroxyecdysone (20E) was investigated in D. melanogaster.
[0064] The steroid 20E is the primary steroid hormone in insects, which triggers molting and metamorphosis in the larval stage.
[0065] RNAi of ecdysone receptor (EcR; encoding the nuclear receptor for 20E) or EcI using the SG-specific driver (fkh-Gal4) resulted in the loss of 20E-dependent glue-GFP expression in the SG in wandering stage larvae (FIG. 1B). SG cells remained intact in both cases.
[0066] RNAi of EcR or EcI in the FB prevented ecdysone-dependent FB cell migration into the pupal head (FIG. 1C).
[0067] Knockdown of EcI, but not EcR, in the FB reduced the intracellular level of ecdysteroids (steroids including 20E) in the tissue, whereas it did not affect the hemolymph level of ecdysteroids in the white prepupal stage (FIG. 1D).
[0068] Thus, it was found that a gene knockdown of EcI in the salivary gland and fat body tissues of D. melanogaster caused similar effects to the knockdown of ecdysone receptor (EcR), which encodes the nuclear receptor for 20E.
[0069] Next, Flag-HA-tagged EcI was overexpressed in FB cells (CG-Gal4>UAS-Oatp74D-Flag-HA). FIG. 2A is an image of the FB cells, in which the flag-HA tagged EcI can be seen localized at the cellular membrane. Flag-HA-tagged EcI was also overexpressed in SG cells (fkh-Gal4>UAS-Oatp74D-Flag-HA). As shown in FIG. 2B, the tagged EcI was also localized at the cellular membrane in the SG cells. These findings support the conclusion that EcI encodes a 20E importer. The incorporated amount of ecdysteroids (steroids including 20E) in the fat body was reduced when the expression of EcI was suppressed (FIG. 1D).
Example 3
Study of EcI in Culture Cell Lines
[0070] The expression of EcI was next evaluated in the Drosophila S2 culture cells and human HEK culture cells.
[0071] S2 culture cells treated with EcI RNAi were incubated with varying concentrations of 20E or with chromafenozide (CF) for 24 hours, and their response was monitored using a luciferase reporter assay. The relative response of the treated cells was compared to control cells, which were incubated with 20E or CF but not treated with RNAi. As shown in the left and middle graphs of FIG. 3, RNAi treatment significantly decreased the cells' response to 20E as measured by the luciferase assay, but had less effect on response to CF. CF is a non-steroidal EcR agonist, and is expected to enter the cells through an unknown transporter distinct from EcI. A time-course analysis of luciferase activity in S2 cells treated with 10.sup.-7 M 20E and with or without RNAi was also performed (FIG. 3, right graph), which showed RNAi treatment significantly decreased the cells' response to 20E over time.
[0072] Human HEK cells were transfected with EcI and EcR, and their response to incubation with varying concentrations of 20E for 24 hours was evaluated using the luciferase reporter assay. Control HEK cells that were transfected only with EcR were also evaluated. As shown in FIG. 4, the control HEK cells are not responsive to 20E as measured by luciferase assay (solid bottom line), while HEK cells transfected with both EcI and EcR were responsive (dashed top line). These results demonstrate that EcI incorporates 20E into animal cells.
Sequence CWU 1
1
313699DNADrosophila melanogaster 1ttaccgtgtg cggactgtgc cgacgggcag
cggaactcag gcggtgcaac ttgagtgctt 60gataaatttg ttgtattgaa acccaaccgc
aaatacaatt ttcaattggc tgagtgcgtg 120tcttcaacgc acaaacatac
gaatatagca tttaatattg ttcattatgt gtgcaattaa 180ataaaaaaaa
ttgtgtgtga aataacagaa aaacgagtgc ggcgaacaga ggcagcaaca
240aattaaacgc gtaaattgcg tagcaaattc tgaatgaaaa aattcaacaa
aaccagaaag 300cgaaacagga gcgataagct taccaactaa agaaactcac
gcctaacaga aatacacacc 360actgccatat taaagcaaag aatatagaac
tgttttacag accagatcaa ccaatctacc 420tcgacttctg ggccaacaag
caaaataaaa ctaaactcaa tgccagaacc caatttcccg 480ataggaaaat
ggaacaggca ataagcaaga ccacatagag gagattaact aaacggatcc
540gttgagatta cacagttgag atcaataaaa cccattgaag atcgatcaaa
ccatctagtt 600ttttagagtt ccattagaag ccaaaatgac gaagagcaat
ggcgatgtgg aggcggcagc 660ccaggtgcaa tctctgggcg gaaagcccag
caacggacat ggccagctga atgggaatgg 720ctatcatcag aacggcggac
gcagggattc cagtcaagcc ttcacaccac tgctgtcgca 780gcacaataat
ggcaccacca atggcgaggt gaccactccg cctccgagta cagtgctcta
840cgagagcacc ccgagcaata ataacgagtg gaaggccccg gaggatctgg
gacacctgaa 900gaatggcctg ggcaacatac tgagcagtaa taataatggc
accggcaatg ggcacagtct 960gagcgaaaag tatgcccatg aacaggctcc
cctgaccgga ggttacaagt tgccacctcg 1020ctccagtgag tccgaggaat
ccgatttcga ttccgacctc aatggcggct cctccgcgga 1080atccagctcc
agttgcggcc ttttcggctg tcggcccagg tgggccagaa gattcgcgtc
1140cacgcacgtc ttcatggtgg tcttcctgct ggcctacatc ctgcagggca
tgtacatgac 1200ctacttcgtc tctgtgatca ccaccattga gaagctattc
cagatcaagt cgaagaccac 1260gggaattctg ctgagcgcca gtgagatggg
tcagatatgc acggccatgt tgctgaccta 1320cttcgccggc agaggacatc
gtccgagatg gattgcctgc ggcatggtcc tgttctcaat 1380cgccgccttc
tcctgcgccc tgccgcattt catcttcggc gagcagctga tgcactccag
1440tgtaattctg cagcagacgc aggtctcccc ccccaataat tccttctcat
cacactggct 1500gaatgccagc agtgaacagg ttaatcccaa tttgtgcatt
ttgggtggca accaaaccca 1560ttcgggcagc gagtgcaacg aggagcgcca
gctggaacag gcctcccact ctaagatcac 1620cgtcatcgtg ctgtgcatct
tctttggcag cctgctcagc tcgggcattg gccagaccgc 1680cgtggccaca
ctgggcatac cctacatcga tgacaatgta ggcagcaagc agtcgcccat
1740gtacatggcc gtcaccattg gcatgaggat cctgggaccg gcatccggtt
tcatttttgg 1800cagcttctgc actcgctggt atgtcaactt ctcgaatccc
ggcttcgacg ccacggatcc 1860gcgctggatt ggcgcctggt ggctggggcc
tgtggccatt ggcagcctca tgctgctggc 1920ctccatcgcc atgttctcgt
ttcccaagca gttgagaggc aagcagaagc cgccggggca 1980gacagcaact
ccagcagctc cagttgagcc ggaggagaag cccaagctaa aagattttcc
2040caagacagtc cgtcgccagc tgagcaacga catcctgatg ttccgcaccg
cctcgtgcgt 2100gttccacctg ctgcccatcg ccggtctcta tacgttcctg
cccaagtatc tggagacgca 2160gttccggctg gccacctatg atgccaacat
gatcgccgcc ttctgtggca tcctggtcat 2220gggcataggt attgtcattt
ccgggctctt catcctgaag cgaaagccca ctgccagggg 2280cgtggccgcc
tggatcgcct ttacagccct cgtctactcg gcgggcatga tcatcctgat
2340gttcatcggc tgcagcatga acgactttgc cggctacaag ccaagcgatg
gcaacagtcc 2400cgccttgatc gagcccacgt gcagtgccgc tctcaactgt
acctgtgata aggagaactt 2460cgcgcccatc tgcgccgacg gcaaaatgta
catctcggcc tgccacgccg gatgcagcag 2520ctcctcactg cggcccagcg
acaatcgcac actctactcc gattgtgctt gcattccaga 2580tgctccggag
gcggtcaacg gttactgtga taataactgc aagaacttca tctactttat
2640actgatcttt gccatttgcg tatttatgca ttccacctcc gaggtgggca
gcatgctgct 2700cgtcatgcgc tgcacacacc ccaaagataa ggccatggcc
atgggtgtga tacagtcggc 2760catcggcctg ttcggcaacg ttccctgtcc
catcatctac ggcgcagtgg tggactccgc 2820ctgcctcatc tggaagtcgg
tgtgcggcaa gcacggcgcc tgttcactct acgatgcgga 2880cactttccgg
caatatttcc taggaatcac ggctggcatt atgttcctgg cattcctgat
2940ggatctggtg gtgtggcgca aggcgcatcg cattgacatc gcgcccgagg
atccgcagga 3000gggcgggccc gcttccaacg gaaggacctt ggaggtgtcc
gagtccaagc agcccatcac 3060cccggcgccg gacacgacgg tctaggagga
gcgggtcggc gacgagcctt gcacaagctg 3120caggatttcc aataaacgtt
taccttaatt gttaattagt tatcatttgt ctagtttgtt 3180agcctagtgc
aagagttatg tatttagtta agtggcatct tcgagcgtcg ggagacctca
3240ttcaaatcca cattagaacg cgctcgtcca gctcccgctc ccgatcctgc
tccaaatccc 3300agcaccatat tctacatgaa gcccatggat tgcgatttga
atccttgtaa atctcaacgc 3360gaaacaatga aatgaaattc tagacattat
ctaacgtcat gcatgagcgt agttaatcga 3420cgagctaata ctacaaactg
atcgacaatt gtgcaaagtg caagaaaatt tatgaatcct 3480tatgtgtaaa
ctatatgtaa cagttatatc gcgatgtatg taatctataa ttaatatatt
3540agacatacac ttacatgtat gtatgtaatt gcaacctatc tgtggtagtt
aaattttagt 3600taaaattcaa attaattgtc taagtttgtc atacaacaaa
tatatacgaa aagaaattaa 3660atggaaaact atatacaagt aaaaaaaaaa
aaaaaaaaa 369922460DNADrosophila melanogaster 2atgacgaaga
gcaatggcga tgtggaggcg gcagcccagg tgcaatctct gggcggaaag 60cccagcaacg
gacatggcca gctgaatggg aatggctatc atcagaacgg cggacgcagg
120gattccagtc aagccttcac accactgctg tcgcagcaca ataatggcac
caccaatggc 180gaggtgacca ctccgcctcc gagtacagtg ctctacgaga
gcaccccgag caataataac 240gagtggaagg ccccggagga tctgggacac
ctgaagaatg gcctgggcaa catactgagc 300agtaataata atggcaccgg
caatgggcac agtctgagcg aaaagtatgc ccatgaacag 360gctcccctga
ccggaggtta caagttgcca cctcgctcca gtgagtccga ggaatccgat
420ttcgattccg acctcaatgg cggctcctcc gcggaatcca gctccagttg
cggccttttc 480ggctgtcggc ccaggtgggc cagaagattc gcgtccacgc
acgtcttcat ggtggtcttc 540ctgctggcct acatcctgca gggcatgtac
atgacctact tcgtctctgt gatcaccacc 600attgagaagc tattccagat
caagtcgaag accacgggaa ttctgctgag cgccagtgag 660atgggtcaga
tatgcacggc catgttgctg acctacttcg ccggcagagg acatcgtccg
720agatggattg cctgcggcat ggtcctgttc tcaatcgccg ccttctcctg
cgccctgccg 780catttcatct tcggcgagca gctgatgcac tccagtgtaa
ttctgcagca gacgcaggtc 840tcccccccca ataattcctt ctcatcacac
tggctgaatg ccagcagtga acaggttaat 900cccaatttgt gcattttggg
tggcaaccaa acccattcgg gcagcgagtg caacgaggag 960cgccagctgg
aacaggcctc ccactctaag atcaccgtca tcgtgctgtg catcttcttt
1020ggcagcctgc tcagctcggg cattggccag accgccgtgg ccacactggg
cataccctac 1080atcgatgaca atgtaggcag caagcagtcg cccatgtaca
tggccgtcac cattggcatg 1140aggatcctgg gaccggcatc cggtttcatt
tttggcagct tctgcactcg ctggtatgtc 1200aacttctcga atcccggctt
cgacgccacg gatccgcgct ggattggcgc ctggtggctg 1260gggcctgtgg
ccattggcag cctcatgctg ctggcctcca tcgccatgtt ctcgtttccc
1320aagcagttga gaggcaagca gaagccgccg gggcagacag caactccagc
agctccagtt 1380gagccggagg agaagcccaa gctaaaagat tttcccaaga
cagtccgtcg ccagctgagc 1440aacgacatcc tgatgttccg caccgcctcg
tgcgtgttcc acctgctgcc catcgccggt 1500ctctatacgt tcctgcccaa
gtatctggag acgcagttcc ggctggccac ctatgatgcc 1560aacatgatcg
ccgccttctg tggcatcctg gtcatgggca taggtattgt catttccggg
1620ctcttcatcc tgaagcgaaa gcccactgcc aggggcgtgg ccgcctggat
cgcctttaca 1680gccctcgtct actcggcggg catgatcatc ctgatgttca
tcggctgcag catgaacgac 1740tttgccggct acaagccaag cgatggcaac
agtcccgcct tgatcgagcc cacgtgcagt 1800gccgctctca actgtacctg
tgataaggag aacttcgcgc ccatctgcgc cgacggcaaa 1860atgtacatct
cggcctgcca cgccggatgc agcagctcct cactgcggcc cagcgacaat
1920cgcacactct actccgattg tgcttgcatt ccagatgctc cggaggcggt
caacggttac 1980tgtgataata actgcaagaa cttcatctac tttatactga
tctttgccat ttgcgtattt 2040atgcattcca cctccgaggt gggcagcatg
ctgctcgtca tgcgctgcac acaccccaaa 2100gataaggcca tggccatggg
tgtgatacag tcggccatcg gcctgttcgg caacgttccc 2160tgtcccatca
tctacggcgc agtggtggac tccgcctgcc tcatctggaa gtcggtgtgc
2220ggcaagcacg gcgcctgttc actctacgat gcggacactt tccggcaata
tttcctagga 2280atcacggctg gcattatgtt cctggcattc ctgatggatc
tggtggtgtg gcgcaaggcg 2340catcgcattg acatcgcgcc cgaggatccg
caggagggcg ggcccgcttc caacggaagg 2400accttggagg tgtccgagtc
caagcagccc atcaccccgg cgccggacac gacggtctag 24603819PRTDrosophila
melanogaster 3Met Thr Lys Ser Asn Gly Asp Val Glu Ala Ala Ala Gln
Val Gln Ser 1 5 10 15 Leu Gly Gly Lys Pro Ser Asn Gly His Gly Gln
Leu Asn Gly Asn Gly 20 25 30 Tyr His Gln Asn Gly Gly Arg Arg Asp
Ser Ser Gln Ala Phe Thr Pro 35 40 45 Leu Leu Ser Gln His Asn Asn
Gly Thr Thr Asn Gly Glu Val Thr Thr 50 55 60 Pro Pro Pro Ser Thr
Val Leu Tyr Glu Ser Thr Pro Ser Asn Asn Asn65 70 75 80 Glu Trp Lys
Ala Pro Glu Asp Leu Gly His Leu Lys Asn Gly Leu Gly 85 90 95 Asn
Ile Leu Ser Ser Asn Asn Asn Gly Thr Gly Asn Gly His Ser Leu 100 105
110 Ser Glu Lys Tyr Ala His Glu Gln Ala Pro Leu Thr Gly Gly Tyr Lys
115 120 125 Leu Pro Pro Arg Ser Ser Glu Ser Glu Glu Ser Asp Phe Asp
Ser Asp 130 135 140 Leu Asn Gly Gly Ser Ser Ala Glu Ser Ser Ser Ser
Cys Gly Leu Phe145 150 155 160 Gly Cys Arg Pro Arg Trp Ala Arg Arg
Phe Ala Ser Thr His Val Phe 165 170 175 Met Val Val Phe Leu Leu Ala
Tyr Ile Leu Gln Gly Met Tyr Met Thr 180 185 190 Tyr Phe Val Ser Val
Ile Thr Thr Ile Glu Lys Leu Phe Gln Ile Lys 195 200 205 Ser Lys Thr
Thr Gly Ile Leu Leu Ser Ala Ser Glu Met Gly Gln Ile 210 215 220 Cys
Thr Ala Met Leu Leu Thr Tyr Phe Ala Gly Arg Gly His Arg Pro225 230
235 240 Arg Trp Ile Ala Cys Gly Met Val Leu Phe Ser Ile Ala Ala Phe
Ser 245 250 255 Cys Ala Leu Pro His Phe Ile Phe Gly Glu Gln Leu Met
His Ser Ser 260 265 270 Val Ile Leu Gln Gln Thr Gln Val Ser Pro Pro
Asn Asn Ser Phe Ser 275 280 285 Ser His Trp Leu Asn Ala Ser Ser Glu
Gln Val Asn Pro Asn Leu Cys 290 295 300 Ile Leu Gly Gly Asn Gln Thr
His Ser Gly Ser Glu Cys Asn Glu Glu305 310 315 320 Arg Gln Leu Glu
Gln Ala Ser His Ser Lys Ile Thr Val Ile Val Leu 325 330 335 Cys Ile
Phe Phe Gly Ser Leu Leu Ser Ser Gly Ile Gly Gln Thr Ala 340 345 350
Val Ala Thr Leu Gly Ile Pro Tyr Ile Asp Asp Asn Val Gly Ser Lys 355
360 365 Gln Ser Pro Met Tyr Met Ala Val Thr Ile Gly Met Arg Ile Leu
Gly 370 375 380 Pro Ala Ser Gly Phe Ile Phe Gly Ser Phe Cys Thr Arg
Trp Tyr Val385 390 395 400 Asn Phe Ser Asn Pro Gly Phe Asp Ala Thr
Asp Pro Arg Trp Ile Gly 405 410 415 Ala Trp Trp Leu Gly Pro Val Ala
Ile Gly Ser Leu Met Leu Leu Ala 420 425 430 Ser Ile Ala Met Phe Ser
Phe Pro Lys Gln Leu Arg Gly Lys Gln Lys 435 440 445 Pro Pro Gly Gln
Thr Ala Thr Pro Ala Ala Pro Val Glu Pro Glu Glu 450 455 460 Lys Pro
Lys Leu Lys Asp Phe Pro Lys Thr Val Arg Arg Gln Leu Ser465 470 475
480 Asn Asp Ile Leu Met Phe Arg Thr Ala Ser Cys Val Phe His Leu Leu
485 490 495 Pro Ile Ala Gly Leu Tyr Thr Phe Leu Pro Lys Tyr Leu Glu
Thr Gln 500 505 510 Phe Arg Leu Ala Thr Tyr Asp Ala Asn Met Ile Ala
Ala Phe Cys Gly 515 520 525 Ile Leu Val Met Gly Ile Gly Ile Val Ile
Ser Gly Leu Phe Ile Leu 530 535 540 Lys Arg Lys Pro Thr Ala Arg Gly
Val Ala Ala Trp Ile Ala Phe Thr545 550 555 560 Ala Leu Val Tyr Ser
Ala Gly Met Ile Ile Leu Met Phe Ile Gly Cys 565 570 575 Ser Met Asn
Asp Phe Ala Gly Tyr Lys Pro Ser Asp Gly Asn Ser Pro 580 585 590 Ala
Leu Ile Glu Pro Thr Cys Ser Ala Ala Leu Asn Cys Thr Cys Asp 595 600
605 Lys Glu Asn Phe Ala Pro Ile Cys Ala Asp Gly Lys Met Tyr Ile Ser
610 615 620 Ala Cys His Ala Gly Cys Ser Ser Ser Ser Leu Arg Pro Ser
Asp Asn625 630 635 640 Arg Thr Leu Tyr Ser Asp Cys Ala Cys Ile Pro
Asp Ala Pro Glu Ala 645 650 655 Val Asn Gly Tyr Cys Asp Asn Asn Cys
Lys Asn Phe Ile Tyr Phe Ile 660 665 670 Leu Ile Phe Ala Ile Cys Val
Phe Met His Ser Thr Ser Glu Val Gly 675 680 685 Ser Met Leu Leu Val
Met Arg Cys Thr His Pro Lys Asp Lys Ala Met 690 695 700 Ala Met Gly
Val Ile Gln Ser Ala Ile Gly Leu Phe Gly Asn Val Pro705 710 715 720
Cys Pro Ile Ile Tyr Gly Ala Val Val Asp Ser Ala Cys Leu Ile Trp 725
730 735 Lys Ser Val Cys Gly Lys His Gly Ala Cys Ser Leu Tyr Asp Ala
Asp 740 745 750 Thr Phe Arg Gln Tyr Phe Leu Gly Ile Thr Ala Gly Ile
Met Phe Leu 755 760 765 Ala Phe Leu Met Asp Leu Val Val Trp Arg Lys
Ala His Arg Ile Asp 770 775 780 Ile Ala Pro Glu Asp Pro Gln Glu Gly
Gly Pro Ala Ser Asn Gly Arg785 790 795 800 Thr Leu Glu Val Ser Glu
Ser Lys Gln Pro Ile Thr Pro Ala Pro Asp 805 810 815 Thr Thr Val
D00001

D00002

D00003

D00004

D00005

D00006

D00007

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.