Lna Antagonists Targeting The Androgen Receptor
WORM; Jesper
U.S. patent application number 13/149155 was filed with the patent office on 2011-12-29 for lna antagonists targeting the androgen receptor. This patent application is currently assigned to SANTARIS PHARMA A/S. Invention is credited to Jesper WORM.
| Application Number | 20110319471 13/149155 |
| Document ID | / |
| Family ID | 40379703 |
| Filed Date | 2011-12-29 |









View All Diagrams
| United States Patent Application | 20110319471 |
| Kind Code | A1 |
| WORM; Jesper | December 29, 2011 |
LNA ANTAGONISTS TARGETING THE ANDROGEN RECEPTOR
Abstract
The invention relates to oligonucleotide compounds (oligomers), which target androgen receptor mRNA in a cell, leading to reduced expression of the androgen receptor. Reduction of the androgen receptor expression is beneficial for the treatment of certain disorders, such as a hyperproliferative disorders (e.g., cancer). The invention provides therapeutic compositions comprising oligomers and methods for modulating the expression of androgen receptor using said oligomers, including methods of treatment.
| Inventors: | WORM; Jesper; (Copenhagen, DK) |
| Assignee: | SANTARIS PHARMA A/S Hoersholm NJ ENZON PHARMACEUTICALS, INC. Bridgewater |
| Family ID: | 40379703 |
| Appl. No.: | 13/149155 |
| Filed: | May 31, 2011 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 12726554 | Mar 18, 2010 | 7989429 | ||
| 13149155 | ||||
| 12324033 | Nov 26, 2008 | 7737125 | ||
| 12726554 | ||||
| 60990125 | Nov 26, 2007 | |||
| Current U.S. Class: | 514/44A ; 536/26.3 |
| Current CPC Class: | A61P 43/00 20180101; A61P 5/28 20180101; A61P 5/26 20180101; C12N 15/1137 20130101; C12N 2310/11 20130101; A61P 17/00 20180101; A61P 3/00 20180101; C12N 2310/341 20130101; A61P 35/00 20180101; C12N 15/1138 20130101; C12N 2310/3231 20130101; A61P 17/14 20180101; A61P 13/08 20180101; C12N 2310/315 20130101; C12N 2310/351 20130101; C12N 2310/3341 20130101 |
| Class at Publication: | 514/44.A ; 536/26.3 |
| International Class: | A61K 31/712 20060101 A61K031/712; A61P 5/26 20060101 A61P005/26; A61P 17/00 20060101 A61P017/00; A61P 3/00 20060101 A61P003/00; C07H 21/00 20060101 C07H021/00; A61P 35/00 20060101 A61P035/00 |
Claims
1. An oligomer consisting of 10 to 30 contiguous monomers wherein adjacent monomers are covalently linked by a phosphate group or a phosphorothioate group, wherein said oligomer comprises a first region of at least 10 contiguous monomers; wherein at least one monomer of said first region is a nucleoside analogue; wherein the sequence of said first region is at least 80% identical to the reverse complement of the best-aligned target region of a mammalian androgen receptor gene or a mammalian androgen receptor mRNA.
2. The oligomer according to claim 1, wherein the sequence of the first region is at least 80% identical to the sequence of a region of at least 10 contiguous monomers present in SEQ ID NOs: 2-22, 44-80 or 86-106.
3. The oligomer according to claim 1, wherein the sequence of the first region is at least 80% identical to the sequence of a region of at least 10 contiguous monomers present in SEQ ID NO 94, or SEQ ID NO 105.
4. The oligomer according to claim 1 wherein the sequence of the first region comprises 0 to 2 mismatches when compared to the sequence of the best-aligned region of the reverse complement of a mammalian androgen receptor gene or a mammalian androgen receptor mRNA.
5. The oligomer according to claim 1, wherein the first region of said oligomer consists of 10 to 18 contiguous monomers.
6. The oligomer according to claim 1, wherein each nucleoside analogue is independently selected from the group consisting of an LNA monomer, a monomer containing a 2'-O-alkyl-ribose sugar, a monomer containing a 2'-O-methyl-ribose sugar, a monomer containing a 2'-amino-deoxyribose sugar, and a monomer containing a 2'fluoro-deoxyribose sugar.
7. The oligomer according to claim 1, wherein the oligomer is a gapmer, and wherein said gapmer comprises from the 5' end to the 3' end: i. a region A consisting of 1-6 contiguous monomers, wherein at least one monomer is a nucleoside analogue, ii. a region B, the 5' end of which is covalently linked to the 3' end of region A and consisting of 5-12 contiguous monomers, wherein at least one monomer is a nucleoside; and iii. a region C, the 5' end of which is covalently linked to the 3' end of region B and consisting of 1-6 contiguous monomers, wherein at least one monomer s a nucleoside analogue.
8. The oligomer according to claim 7, wherein the oligomer is a gapmer, and wherein said gapmer comprises from the 5'-end to the 3'-end: i. a region A consisting of 2-5 contiguous monomers, wherein all monomers are nucleoside analogues ii. a region B, the 5' end of which is covalently linked to the 3' end of region A and consisting of 6-10 contiguous monomers, wherein all monomers are nucleosides; and iii. a region C, the 5' end of which is covalently linked to the 3' end of region B and consisting of 2-5 contiguous monomers, wherein all monomers are nucleoside analogues.
9. The oligomer according to claim 7, wherein all nucleoside analogues are LNA monomers.
10. The oligomer according to claim 1, which inhibits the expression of a human androgen receptor gene or mRNA in a cell that expresses androgen receptor.
11. A conjugate comprising an oligomer according to claim 1 covalently attached to at least one moiety that is not a nucleic acid or a monomer.
12. A pharmaceutical composition comprising the oligomer according to claim 1 or the conjugate comprising the oligomer of claim 1, and a pharmaceutically acceptable diluent, carrier, salt or adjuvant.
13. A method of (a) inhibiting the expression of androgen receptor or androgen receptor target gene in a cell or a tissue of a mammal; (b) treating cancer in a mammal; or (c) treating a disorder wherein the disorder is selected from the group consisting of alopecia, benign prostatic hyperplasia, spinal and muscular atrophy and Kennedy disease and polyglutamate disease, comprising contacting said cell or tissue of a mammal with; or administering to said mammal, an effective amount of an oligomer consisting of 10 to 30 contiguous monomers wherein adjacent monomers are covalently linked by a phosphate group or a phosphorothioate group or a conjugate comprising the oligomer, wherein said oligomer comprises a first region of at least 10 contiguous monomers; wherein at least one monomer of said first region is a nucleoside analogue; wherein the sequence of said first region is at least 80% identical to the reverse complement of the best-aligned target region of a mammalian androgen receptor gene or a mammalian androgen receptor mRNA.
14. The method of claim 13, wherein the androgen receptor target gene is selected from the group consisting of a protein kinase C delta gene, a glutathione S-transferase theta 2 gene, a transient receptor potential cation channel subfamily V member 3 gene, a pyrroline-5-carboxylate reductase 1 gene and an ornithine aminotransferase gene in a tissue of a mammal comprising contacting said tissue with an effective amount of a conjugate according to claim 11.
15. The method of claim 13, wherein the cancer is breast cancer or prostate cancer.
Description
[0001] This application is a continuation application of U.S. application Ser. No. 12/726,554 filed on Mar. 18, 2010, which is a continuation application of U.S. application Ser. No. 12/324,033 filed on Nov. 26, 2008, now U.S. Pat. No. 7,737,125 issued on Jun. 15, 2010, which claims the benefit under 35 U.S.C. .sctn.119(e) of U.S. Provisional Application Ser. No. 60/990,125 filed Nov. 26, 2007, the disclosure of each of which is incorporated herein by reference in its entirety.
FIELD OF INVENTION
[0002] The invention provides compounds, compositions and methods for modulating the expression of the androgen receptor. In particular, this invention relates to oligomeric compounds (oligomers), which target the androgen receptor mRNA in a cell, leading to reduced expression of the androgen receptor. Reduction of androgen receptor expression is beneficial for a range of medical disorders, such as cancer, particularly prostate cancer or breast cancer.
BACKGROUND
[0003] The androgen receptor ("AR") is a type of nuclear receptor which is activated by binding of either of the androgenic hormones testosterone or dihydrotestosterone. The main function of the androgen receptor is as a DNA binding transcription factor which regulates gene expression. However the androgen receptor also has additional functions independent of DNA binding. The androgen receptor is most closely related to the progesterone receptor, and progestins in higher dosages can block the androgen receptor.
[0004] Whilst in humans the AR gene is single copy and found on the X chromosome at position Xq11-12, the receptor itself exists in two iso-forms (A and B). AR-A is an 87 kDa protein which lacks the first 187 amino acids (N-terminal truncation). Isoform AR-B is the full length 110 kDa version.
[0005] The binding of an androgen to the androgen receptor induces a conformational change in the receptor, resulting in a dissociation of heat shock proteins, dimerization and transport from the cytosol to the cell nucleus where the androgen receptor dimer binds to specific DNA sequences--referred to as hormone response elements. Depending on the interaction with other nuclear proteins, the AR controls gene expression, either increasing or decreasing transcription of specific genes, such as insulin-like growth factor I (IGF-1).
[0006] Androgen receptors can also have cytoplasmic activities through interaction with signal transduction proteins in the cytoplasm. Androgen binding to cytoplasmic androgen receptors can cause rapid changes in cell function independent of gene transcription, for example ion transport, as well as indirect influence of gene transcription, for example via mediating other signal transduction pathways, thereby influencing the activity of other transcription factors.
[0007] The over-expression of androgen receptor, or expression of mutated androgen receptor genes, has been indicated in several diseases, such as cancer, including prostate cancer and breast cancer, as well as other disorders such as polyglutamate disease (Monks et al., PNAS Nov. 2 2007, published on line) alopecia, benign prostatic hyperplasia, spinal and muscular atrophy and Kennedy disease.
[0008] WO97/11170 describes a method of treating a patient diagnosed as having benign prostatic hyperplasia or a prostate cancer comprising administering an anti sense oligonucleotide which selectively hybridises to the androgen receptor mRNA. Three antisense oligonucleotide sequences of between 27-29 nucleotides are disclosed.
[0009] U.S. Pat. No. 6,733,776 and EP 0 692 972 describe a method for treating androgenic alopecia by applying liposomes comprising an antisense nucleic acid that hybridises to an androgen receptor gene. No antisense molecules having specific sequences and targeting the androgen receptor are provided.
[0010] US 2005/0164970 describes a method of treating prostate cancer using siRNA complexes targeting the androgen receptor mRNA.
[0011] WO 2005/027833 describes a method of treating prostate cancer comprising administering to a patient an oligonucleotide comprising between 12-40 morpholino sub-units.
[0012] WO 2001/083740 describes an antisense compound having an uncharged morpholino backbone of between 18 to 20 contiguous units which targets the human androgen receptor.
[0013] Morpholino antisense compounds work via binding to the nucleic acid target to block access to the mRNA by other molecules, such as molecules involved in mRNA splicing or translation initiation.
[0014] U.S. Pat. No. 7,067,256 describes a ribozyme which apparently mediates inactivation of the androgen receptor. A 19-nucleotide RNA antisense molecule targeted to a region of the androgen receptor mRNA is provided.
[0015] However, despite the application of siRNA, morpholino-containing antisense oligonucleotides and ribozymes, none of the above androgen receptor inhibitors have been successful in efficiently down-regulating the androgen-receptor in vivo and at pharmacologically acceptable dosages.
[0016] The invention provides a new class of androgen receptor antagonists which contain locked nucleic acid ("LNA") monomers, and are targeted to particularly effective target sites on the androgen receptor mRNA.
SUMMARY OF INVENTION
[0017] The invention provides an oligomer of from 10-50 monomers, such as 10-30 monomers which comprises a first region of 10-50 monomers, such as 10-30 monomers, wherein the sequence of the first region is at least 80% (e.g., 85%, 90%, 95%, 98%, or 99%) identical to the reverse complement of a target region of a nucleic acid which encodes a mammalian androgen receptor, such as a mammalian androgen receptor gene or mRNA, such as a nucleic acid having the sequence set forth in SEQ ID NO: 1, or naturally occurring variants thereof. Thus, for example, the oligomer hybridizes to a region of a single-stranded nucleic acid molecule having the sequence shown in SEQ ID NO: 1.
[0018] The invention provides for a conjugate comprising the oligomer according to the invention, and at least one non-nucleotide or non-polynucleotide moiety covalently attached to the oligomer.
[0019] The invention provides for a pharmaceutical composition comprising the oligomer or the conjugate according to the invention, and a pharmaceutically acceptable diluent, carrier, salt or adjuvant.
[0020] The invention provides for the oligomer or the conjugate according to the invention, for use as a medicament, such as for the treatment of a disease or a medical disorder as disclosed herein, such as a hyperproliferative disorder, such as cancer or other hyperproliferative disorder. The invention provides for the use of an oligomer or the conjugate according to the invention, for the manufacture of a medicament for the treatment of a disease or disorder as disclosed herein, such as a hyperproliferative disorder, such as cancer.
[0021] The invention provides for a method of treating a disease or disorder as disclosed herein, such as a hyperproliferative disorder, such as cancer, the method comprising administering an oligomer, a conjugate or a pharmaceutical composition according to the invention to a patient suffering from or susceptible to the disease or disorder.
[0022] The invention provides for a method for the inhibition of androgen receptor in a Dell which is expressing androgen receptor, the method comprising administering an oligomer, or a conjugate according to the invention to the cell so as to effect the inhibition of androgen receptor expression in said cell.
[0023] The invention provides an oligomer of from 10-50 monomers, which comprises a first region of 10-50 contiguous monomers, wherein the base sequence is at least 80% identical to the reverse complement of a target region of a nucleic acid which encodes a mammalian androgen receptor.
[0024] The invention further provides a conjugate comprising the oligomer according to the invention, which comprises at least one non-nucleotide or non-polynucleotide moiety ("conjugated moiety") covalently attached to the oligomer of the invention.
[0025] The invention provides for pharmaceutical compositions comprising an oligomer or conjugate of the invention, and a pharmaceutically acceptable diluent, carrier, salt or adjuvant.
[0026] The invention further provides for an oligomer according to the invention, for use in medicine.
[0027] The invention further provides for the use of the oligomer of the invention for the manufacture of a medicament for the treatment of one or more of the diseases referred to herein, such as a disease selected from the group consisting of cancer, such as breast cancer or prostate cancer, alopecia, benign prostatic hyperplasia, spinal and muscular atrophy, Kennedy disease and polyglutamate disease.
[0028] The invention further provides for an oligomer according to the invention, for use for the treatment of one or more of the diseases referred to herein, such as a disease selected from the group consisting of cancer, such as breast cancer or prostate cancer, alopecia, benign prostatic hyperplasia, spinal and muscular atrophy, Kennedy disease and polyglutamate disease.
[0029] Pharmaceutical and other compositions comprising an oligomer of the invention are also provided. Further provided are methods of down-regulating the expression of AR in cells or tissues comprising contacting said cells or tissues, in vitro or in vivo, with one or more of the oligomers, conjugates or compositions of the invention.
[0030] Also disclosed are methods of treating a non-human animal or a human suspected of having, or susceptible to, a disease or condition, associated with expression, or over-expression of AR by administering to the animal or human a therapeutically or prophylactically effective amount of one or more of the oligomers, conjugates or pharmaceutical compositions of the invention. Further, methods of using oligomers for the inhibition of expression of AR, and for treatment of diseases associated with activity of AR are provided.
[0031] The invention provides for a method for treating a disease selected from the group consisting of: cancer, such as breast cancer or prostate cancer, alopecia, benign prostatic hyperplasia, spinal and muscular atrophy, Kennedy disease and polyglutamate disease, the method comprising administering an effective amount of one or more oligomers, conjugates, or pharmaceutical compositions thereof to a patient in need thereof.
[0032] The invention provides for methods of inhibiting (e.g., by down-regulating) the expression of AR in a cell or a tissue, the method comprising the step of contacting the cell or tissue with an effective amount of one or more oligomers, conjugates, or pharmaceutical compositions thereof, to effect down-regulation of expression of AR.
BRIEF DESCRIPTION OF FIGURES
[0033] FIG. 1. Oligonucleotides presented in Table 3 were evaluated for their potential to knockdown the androgen receptor mRNA at concentrations of 1, 4 and 16 nM in MCF7 cells 24 hours after transfection using Real-time PCR. All results were normalised to GAPDH and inhibition of AR mRNA is shown as percent of untreated control. Results shown are an average of three independent experiments.
[0034] FIG. 2. Oligonucleotides presented in Table 3 were evaluated for their potential to knockdown the androgen receptor mRNA at concentrations of 1, 4 and 16 nM in A549 cells 24 hours after transfection using Real-time PCR. All results were normalised to GAPDH and inhibition of AR mRNA is shown as percent of untreated control. Results shown are an average of three independent experiments.
[0035] FIG. 3. Sequence alignment of the human Androgen receptor mRNA sequence (GenBank Accession No.: NM.sub.--000044) and the mouse Androgen receptor mRNA sequence (GenBank Accession No.: NM.sub.--013476).
[0036] FIG. 4. Location of presently preferred target regions of the human AR mRNA (cDNA) targeted by oligomers according to the invention. Although 16mer target sites have been shown, in some embodiments these target regions comprise an additional 4 monomers 5' or 3' to the target regions shown--i.e. are target regions comprising up to 24 contiguous monomers.
[0037] FIG. 5. SEQ ID NO: 1 Homo sapiens androgen receptor (dihydrotestosterone receptor; testicular feminization; spinal and bulbar muscular atrophy; Kennedy disease) (AR), transcript variant 1, mRNA. (GenBank Accession number: NM.sub.--000044).
[0038] FIG. 6. SEQ ID NO 81: Mouse androgen receptor mRNA sequence.
[0039] FIG. 7. SEQ ID NO 82: Rhesus monkey androgen receptor mRNA sequence.
[0040] FIG. 8. SEQ ID NO 83: Homo sapiens androgen receptor protein amino acid sequence.
[0041] FIG. 9. SEQ ID NO 84: Mouse androgen receptor protein amino acid sequence.
[0042] FIG. 10. SEQ ID NO 85: Rhesus monkey androgen receptor protein amino acid sequence.
[0043] FIG. 11: AR mRNA in LNCaP, 24 h post-transfection
[0044] FIG. 12: AR mRNA in A549, 24 h post-transfection
[0045] FIG. 13: Cell proliferation assay--A549, time course post-transfection
[0046] FIG. 14: Cell proliferation assay--time course post-transfection
[0047] FIG. 15: Caspase 3/7 activity in LNCaP cells, 24, 48 or 72 hours post-transfection.
[0048] FIG. 16: Caspase 3/7 activity in A549 cells, 24, 48 or 72 hours post-transfection.
[0049] FIG. 17: Average PSA in plasma after in vivo oligomer treatment.
[0050] FIG. 18: In viva inhibition of tumor growth
DETAILED DESCRIPTION OF INVENTION
The Oligomer
[0051] The invention employs oligomeric compounds (referred herein as oligomers), for use in modulating the function of nucleic acid molecules encoding mammalian androgen receptor, such as the androgen receptor nucleic acid shown in SEQ ID NO:1, and naturally occurring variants of such nucleic acid molecules encoding mammalian androgen receptor. The term "oligomer" in the context of the invention, refers to a molecule formed by covalent linkage of two or more monomers (i.e. an oligonucleotide). In some embodiments, the oligomer comprises or consists of from 10-30 covalently linked monomers.
[0052] The term "monomer" includes both nucleosides and deoxynucleosides (collectively, "nucleosides") that occur naturally in nucleic acids and that do not contain either modified sugars or modified nucleobases, i.e., compounds in which a ribose sugar or deoxyribose sugar is covalently bonded to a naturally-occurring, unmodified nucleobase (base) moiety (i.e., the purine and pyrimidine heterocycles adenine, guanine, cytosine, thymine or uracil) and "nucleoside analogues," which are nucleosides that either do occur naturally in nucleic acids or do not occur naturally in nucleic acids, wherein either the sugar moiety is other than a ribose or a deoxyribose sugar (such as bicyclic sugars or 2' modified sugars, such as 2' substituted sugars), or the base moiety is modified (e.g., 5-methylcytosine), or both.
[0053] An "RNA monomer" is a nucleoside containing a ribose sugar and an unmodified nucleobase;
[0054] A "DNA monomer" is a nucleoside containing a deoxyribose sugar and an unmodified nucleobase.
[0055] A "Locked Nucleic Acid monomer," "locked monomer," or "LNA monomer" is a nucleoside analogue having a bicyclic sugar, as further described herein below.
[0056] The terms "corresponding nucleoside analogue" and "corresponding nucleoside" indicate that the base moiety in the nucleoside analogue and the base moiety in the nucleoside are identical. For example, when the "nucleoside" contains a 2-deoxyribose sugar linked to an adenine, the "corresponding nucleoside analogue" contains, for example, a modified sugar linked to an adenine base moiety.
[0057] The terms "oligomer," "oligomer is compound," and "oligonucleotide" are used interchangeably in the context of the invention, and refer to a molecule formed by covalent linkage of two or more contiguous monomers by, for example, a phosphate group (forming a phosphodiester linkage between nucleosides) or a phosphorothioate group (forming a phosphorothioate linkage between nucleosides). The oligomer consists of, or comprises, 10-50 monomers, such as 10-30 monomers.
[0058] In some embodiments, an oligomer comprises nucleosides, or nucleoside analogues, or mixtures thereof as referred to herein. An "LNA oligomer" or "LNA oligonucleotide" refers to an oligonucleotide containing one or more LNA monomers.
[0059] Nucleoside analogues that are optionally included within oligomers may function similarly to corresponding nucleosides, or may have specific improved functions. Oligomers wherein some or all of the monomers are nucleoside analogues are often preferred over native forms because of several desirable properties of such oligomers, such as the ability to penetrate a cell membrane, good resistance to extra- and/or intracellular nucleases and high affinity and specificity for the nucleic acid target, LNA monomers are particularly preferred, for example, for conferring several of the above-mentioned properties.
[0060] In various embodiments, one or more nucleoside analogues present within the oligomer are "silent" or "equivalent" in function to the corresponding natural nucleoside, i.e., have no functional effect on the way the oligomer functions to inhibit target gene expression. Such "equivalent" nucleoside analogues are nevertheless useful if, for example, they are easier or cheaper to manufacture, or are more stable under storage or manufacturing conditions, or can incorporate a tag or label. Typically, however, the analogues will have a functional effect on the way in which the oligomer functions to inhibit expression; for example, by producing increased binding affinity to the target region of the target nucleic acid and/or increased resistance to intracellular nucleases and/or increased ease of transport into the cell.
[0061] Thus, in various embodiments, oligomers according to the invention comprise nucleoside monomers and at least one nucleoside analogue monomer, such as an LNA monomer, or other nucleoside analogue monomers.
[0062] The term "at least one" comprises the integers larger than or equal to 1, such as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 and so forth. In various embodiments, such as when referring to the nucleic acid or protein targets of the compounds of the invention, the term "at least one" includes the terms "at least two" and "at least three" and "at least four." Likewise, in some embodiments, the term "at least two" comprises the terms "at least three" and "at least four."
[0063] In some embodiments, the oligomer comprises or consists of 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29 or 30 contiguous monomers.
[0064] In some embodiments, the oligomer comprises or consists of 10-22 contiguous monomers, such as 12-18 contiguous monomers, such as 13-17 or 12-16 contiguous monomers, such as 13, 14, 15, 16 contiguous monomers.
[0065] In certain embodiments, the oligomer comprises or consists of 10, 11, 12, 13, or 14 contiguous monomers.
[0066] In various embodiments, the oligomer according to the invention consists of no more than 22 monomers, such as no more than 20 monomers, such as no more than 18 monomers, such as 15, 16 or 17 monomers. In some embodiments, the oligomer of the invention comprises less than 20 monomers.
[0067] In various embodiments, the compounds of the invention do not comprise RNA monomers.
[0068] In various embodiments, the compounds according to the invention are linear molecules or are linear as synthesised. The oligomer, in such embodiments, is a single stranded molecule, and typically does not comprise short regions of, for example, at least 3, 4 or 5 contiguous monomers, which are complementary to another region within the same oligomer such that the oligomer forms an internal duplex. In some embodiments, the oligomer is essentially not double stranded, i.e., is not a siRNA.
[0069] In some embodiments, the oligomer of the invention consists of a contiguous stretch of monomers, the sequence of which is identified by a SEQ ID NO disclosed herein (see, e.g., Tables 1-3). In other embodiments, the oligomer comprises a first region, the region consisting of a contiguous stretch of monomers, and one or more additional regions which consist of at least one additional monomer. In some embodiments, the sequence of the first region is identified by a SEQ ID NO disclosed herein.
Gapmer Design
[0070] Typically, the oligomer of the invention is a gapmer.
[0071] A "gapmer" is an oligomer which comprises a contiguous stretch of monomers capable of recruiting an RNAse (e.g., such as RNAseH) as further described herein below, such as a region of at least 6 or 7 DNA monomers, referred to herein as region B, wherein region B is flanked both on its 5' and 3' ends by regions respectively referred to as regions A and C, each of regions A and C comprising or consisting of nucleoside analogues, such as affinity-enhancing nucleoside analogues, such as 1-6 nucleoside analogues.
[0072] Typically, the gapmer comprises regions, from 5' to 3', A-B-C, or optionally A-B-C-D or D-A-B-C, wherein: region A consists of or comprises at least one nucleoside analogue, such as at least one LNA monomer, such as 1-6 nucleoside analogues, such as LNA monomers, and region B consists of or comprises at least five contiguous monomers which are capable of recruiting RNAse (when formed in a duplex with a complementary target region of the target RNA molecule, such as the mRNA target), such as DNA monomers; region C consists of or comprises at least one nucleoside analogue, such as at least one LNA monomer, such as 1-6 nucleoside analogues, such as LNA monomers; and region D, when present, consists of or comprises 1, 2 or 3 monomers, such as DNA monomers.
[0073] In various embodiments, region A consists of 1, 2, 3, 4, 5 or 6 nucleoside analogues, such as LNA monomers, such as 2-5 nucleoside analogues, such as 2-5 LNA monomers, such as 3 or 4 nucleoside analogues, such as 3 or 4 LNA monomers; and/or region C consists of 1, 2, 3, 4, 5 or 6 nucleoside analogues, such as LNA monomers, such as 2-5 nucleoside analogues, such as 2-5 LNA monomers, such as 3 or 4 nucleoside analogues, such as 3 or 4 LNA monomers.
[0074] In certain embodiments, region B consists of or comprises 5, 6, 7, 8, 9, 10, 11 or 12 contiguous monomers which are capable of recruiting RNAse, or 6-10, or 7-9, such as 8 contiguous monomers which are capable of recruiting RNAse. In certain embodiments, region B consists of or comprises at least one DNA monomer, such as 1-12 DNA monomers, preferably 4-12 DNA monomers, more preferably 6-10 DNA monomers, such as 7-10 DNA monomers, most preferably 8, 9 or 10 DNA monomers.
[0075] In various embodiments, region A consists of 3 or 4 nucleoside analogues, such as LNA monomers, region B consists of 7, 8, 9 or 10 DNA monomers, and region C consists of 3 or 4 nucleoside analogues, such as LNA monomers. Such designs include (A-B-C) 3-10-3, 3-10-4, 4-10-3, 3-9-3, 3-9-4, 4-9-3, 3-8-3, 3-8-4, 4-8-3, 3-7-3, 3-7-4, 4-7-3, and may further include region D, which may have one or 2 monomers, such as DNA monomers.
[0076] Further gapmer designs are disclosed in WO2004/046160, which is hereby incorporated by reference.
[0077] U.S. provisional application, 60/977,409, hereby incorporated by reference, refers to `shortmer` gapmer oligomers. In some embodiments, oligomers presented here may be such shortmer gapmers.
[0078] In certain embodiments, the oligomer consists of 10, 11, 12, 13 or 14 contiguous monomers, wherein the regions of the oligomer have the pattern (5'-3'), A-B-C, or optionally A-B-C-D or D-A-B-C, wherein: region A consists of 1, 2 or 3 nucleoside analogue monomers, such as LNA monomers; region B consists of 7, 8 or 9 contiguous monomers which are capable of recruiting RNAse when formed in a duplex with a complementary RNA molecule (such as a mRNA target); and region C consists of 1, 2 or 3 nucleoside analogue monomers, such as LNA monomers. When present, region D consists of a single DNA monomer.
[0079] In certain embodiments, region A consists of 1 LNA monomer. In certain embodiments, region A consists of 2 LNA monomers. In certain embodiments, region A consists of 3 LNA monomers. In certain embodiments, region C consists of 1 LNA monomer. In certain embodiments, region C consists of 2 LNA monomers. In certain embodiments, region C consists of 3 LNA monomers. In certain embodiments, region B consists of 7 nucleoside monomers. In certain embodiments, region B consists of 8 nucleoside monomers. In certain embodiments, region B consists of 9 nucleoside monomers. In certain embodiments, region B comprises 1-9 DNA monomers, such as 2, 3, 4, 5, 6, 7 or 8 DNA monomers. In certain embodiments, region B consists of DNA monomers. In certain embodiments, region B comprises at least one LNA monomer which is in the alpha-L configuration, such as 2, 3, 4, 5, 6, 7, 8 or 9 LNA monomers in the alpha-L-configuration. In certain embodiments, region B comprises at least one alpha-L-oxy LNA monomer. In certain embodiments, all the LNA monomers in region B that are in the alpha-L-configuration are alpha-L-oxy LNA units. In certain embodiments, the number of monomers present in the A-B-C regions are selected from the group consisting of (nucleoside analogue monomers--region B--nucleoside analogue monomers): 1-8-1, 1-8-2, 2-8-1, 2-8-2, 3-8-3, 2-8-3, 3-8-2, 4-8-1, 4-8-2, 1-8-4, 2-8-4, or; 1-9-1, 1-9-2, 2-9-1, 2-9-2, 2-9-3, 3-9-2, 1-9-3, 3-9-1, 4-9-1, 1-9-4, or; 1-10-1, 1-10-2, 2-10-1, 2-10-2, 1-10-3, 3-10-1. In certain embodiments, the number of monomers present in the A-B-C regions of the oligomer of the invention is selected from the group consisting of: 2-7-1, 1-7-2, 2-7-2, 3-7-3, 2-7-3, 3-7-2, 3-7-4, and 4-7-3. In certain embodiments, each of regions A and C consists of two LNA monomers, and region B consists of 8 or 9 nucleoside monomers, preferably DNA monomers.
[0080] In various embodiments, other gapmer designs include those where regions A and/or C consists of 3, 4, 5 or 6 nucleoside analogued, such as monomers containing a 2'-O-methoxyethyl-ribose sugar (2'-MOE) or monomers containing a 2'-fluoro-deoxyribose sugar, and region B consists of 8, 9, 10, 11 or 12 nucleosides, such as DNA monomers, where regions A-B-C have 5-10-5 or 4-12-4 monomers. Further gapmer designs are disclosed in WO 2007/146511A2, hereby incorporated by reference.
Internucleoside Linkages
[0081] The monomers of the oligomers described herein are coupled together via linkage groups. Suitably, each monomer is linked to the 3' adjacent monomer via a linkage group.
[0082] The terms "linkage group" or "internucleoside linkage" means a group capable of covalently coupling together two contiguous monomers. Specific and preferred examples include phosphate groups (forming a phosphodiester between adjacent nucleoside monomers) and phosphorothioate groups (forming a phosphorothioate linkage between adjacent nucleoside monomers).
[0083] Suitable linkage groups include those listed in PCT/DK2006/000512, for example in the first paragraph of page 34 of PCT/DK2006/000512 (hereby incorporated by reference).
[0084] It is, in various embodiments, preferred to modify the linkage group from its normal phosphodiester to one that is more resistant to nuclease attack, such as phosphorothioate or boranophosphate--these two being cleavable by RNase H, thereby permitting RNase-mediated antisense inhibition of expression of the target gene.
[0085] In some embodiments, suitable sulphur (S) containing linkage groups as provided herein are preferred. In various embodiments, phosphorothioate linkage groups are preferred, particularly for the gap region (B) of gapmers. In certain embodiments, phosphorothioate linkages are used to link together monomers in the flanking regions (A and C). In various embodiments, phosphorothioate linkages are used for linking regions A or C to region D, and for linking together monomers within region D.
[0086] In various embodiments, regions A, B and C, comprise linkage groups other than phosphorothioate, such as phosphodiester linkages, particularly, for instance when the use of nucleoside analogues protects the linkage groups within regions A and C from endo-nuclease degradation--such as when regions A and C comprise LNA monomers.
[0087] In various embodiments, adjacent monomers of the oligomer are linked to each other by means of phosphorothioate groups.
[0088] It is recognised that the inclusion of phosphodiester linkages, such as one or two linkages, into an oligomer with a phosphorothioate backbone, particularly with phosphorothioate linkage groups between or adjacent to nucleoside analogue monomers (typically in region A and/or C), can modify the bioavailability and/or bio-distribution of an oligomer--see WO2008/053314, hereby incorporated by reference.
[0089] In some embodiments, such as the embodiments referred to above, where suitable and not specifically indicated, all remaining linkage groups are either phosphodiester or phosphorothioate, or a mixture thereof.
[0090] In some embodiments all the internucleoside linkage groups are phosphorothioate.
[0091] When referring to specific gapmer oligonucleotide sequences, such as those provided herein, it will be understood that, in various embodiments, when the linkages are phosphorothioate linkages, alternative linkages, such as those disclosed herein may be used, for example phosphate (phosphodiester) linkages may be used, particularly for linkages between nucleoside analogues, such as LNA monomers. Likewise, in various embodiments, when referring to specific gapmer oligonucleotide sequences, such as those provided herein, when one or more monomers in region C comprises a 5-methylcytosine base, other monomers in that region may contain unmodified cytosine bases.
Target Nucleic Acid
[0092] The terms "nucleic acid" and "polynucleotide" are used interchangeably herein, and are defined as a molecule formed by covalent linkage of two or more monomers, as above-described. Including 2 or more monomers, "nucleic acids" may be of any length, and the term is generic to "oligomers", which have the lengths described herein. The terms "nucleic acid" and "polynucleotide" include single-stranded, double-stranded, partially double-stranded, and circular molecules.
[0093] The term "target nucleic acid", as used herein, refers to DNA or RNA (e.g., mRNA or pre-mRNA) encoding a mammalian androgen receptor polypeptide, such as human androgen receptor, such as the nucleic acid having the sequence shown in SEQ ID NO: 1, and naturally occurring allelic variants of such nucleic acids. In certain embodiments, the mammalian androgen receptor is a mouse androgen receptor. In some embodiments, for example when used in research or diagnostics, the "target nucleic acid" is a cDNA or a synthetic oligonucleotide derived from the above DNA or RNA nucleic acid targets. The oligomers according to the invention are typically capable of hybridising to the target nucleic acid.
[0094] Exemplary target nucleic acids include mammalian androgen receptor-encoding nucleic acids having the GenBank Accession numbers shown in the table below, along with their corresponding protein sequences;
TABLE-US-00001 GenBank Accession Number Nucleic acid (mRNA/cDNA GenBank Accession Number sequence) Polypeptide (deduced) Human NM_000044 NP_000035 Mouse NM_013476 NP_038504 Rhesus NM_001032911 NP_001028083 monkey
[0095] It is recognised that the above-disclosed GenBank Accession numbers for nucleic acids refer to cDNA sequences and not to mRNA sequences per se. The sequence of a mature mRNA can be derived directly from the corresponding cDNA sequence with thymine bases (T) being replaced by uracil bases (U).
[0096] The term "naturally occurring variant thereof" refers to variants of the androgen receptor polypeptide or nucleic acid sequence which exist naturally within the defined taxonomic group, such as mammalian, such as mouse, monkey, and preferably human AR. Typically, when referring to "naturally occurring variants" of a polynucleotide the term also encompasses any allelic variant of the androgen receptor encoding genomic DNA which is found at the Chromosome X: 66.68-66.87 Mb by chromosomal translocation or duplication, and the RNA, such as mRNA derived therefrom. "Naturally occurring variants" may also include variants derived from alternative splicing of the androgen receptor mRNA. When referenced to a specific polypeptide sequence, e.g., the term also includes naturally occurring forms of the protein which may therefore be processed, e.g. by co- or post-translational modifications, such as signal peptide cleavage, proteolytic cleavage, glycosylation, etc.
[0097] It is recognised that the human androgen receptor gene exhibits allelic variations that are associated with disease phenotypes (Mooney et al, NAR 15; 31(8) 2003). For example, a (CAG).sub.n repeat expansion is associated with polyglutamine expansion disorder. Other characterised allelic variants include a (GGC).sub.n trinucleotide repeat and single nucleotide polymorphisms R726L, T887A and L710H, of which the latter two single nucleotide polymorphisms have been shown to be correlated to enhanced promiscuity of the AR receptor for other steroid ligands. In one embodiment "n" ranges from 5-31. CAG repeats of less than 22 have been associated with an enhanced risk of prostate cancer in African American males.
[0098] In various embodiments, the target nucleic acid is an AR allelic variant which comprises a (CAG).sub.n trinucleotide repeat, or (GGC).sub.n trinucleotide repeat. In other embodiments, the target nucleic acid is an AR allelic variant which comprises one or more single nucleotide polymorphisms, including R726L, T887A and L710H.
[0099] In certain embodiments, oligomers described herein bind to a region of the target nucleic acid (the "target region") by either Watson-Crick base pairing, Hoogsteen hydrogen bonding, or reversed Hoogsteen hydrogen bonding, between the monomers of the oligomer and monomers of the target nucleic acid. Such binding is also referred to as "hybridisation." Unless otherwise indicated, binding is by Watson-Crick pairing of complementary bases (i.e., adenine with thymine (DNA) or uracil (RNA), and guanine with cytosine), and the oligomer binds to the target region because the sequence of the oligomer is identical to, or partially-identical to, the sequence of the reverse complement of the target region; for purposes herein, the oligomer is said to be "complementary" or "partially complementary" to the target region, and the percentage of "complementarity" of the oligomer sequence to that of the target region is the percentage "identity" to the reverse complement of the sequence of the target region.
[0100] Unless otherwise made clear by context, the "target region" herein will be the region of the target nucleic acid having the sequence that best aligns with the reverse complement of the sequence of the specified oligomer (or region thereof), using the alignment program and parameters described herein below.
[0101] In determining the degree of "complementarily" between oligomers of the invention (or regions thereof) and the target region of the nucleic acid which encodes mammalian androgen receptor, such as those disclosed herein, the degree of "complementarity" (also, "homology") is expressed as the percentage identity between the sequence of the oligomer (or region thereof) and the reverse complement of the sequence of the target region that best aligns therewith. The percentage is calculated by counting the number of aligned bases that are identical as between the 2 sequences, dividing by the total number of contiguous monomers in the oligomer, and multiplying by 100. In such a comparison, if gaps exist, it is preferable that such gaps are merely mismatches rather than areas where the number of monomers within the gap differs between the oligomer of the invention and the target region.
[0102] Amino acid and polynucleotide alignments, percentage sequence identity, and degree of complementarity may be determined for purposes of the invention using the ClustalW algorithm using standard settings: see http://www.ebi.ac.uk/emboss/align/index.html, Method: EMBOSS::water (local): Gap Open=10.0, Gap extend=0.5, using Blosum 62 (protein), or DNAfull for nucleotide/nucleobase sequences.
[0103] As will be understood, depending on context, "mismatch" refers to a non-identity in sequence (as, for example, between the nucleobase sequence of an oligomer and the reverse complement of the target region to which it binds; as for example, between the base sequence of two aligned AR encoding nucleic acids), or to noncomplementarity in sequence (as, for example, between an oligomer and the target region to which it binds).
[0104] The androgen receptor is known to regulate the expression of several genes, such as a gene selected from the group consisting of Protein kinase C delta (PRKCD), Glutathione S-transferase theta 2 (GSTT2), transient receptor potential cation channel subfamily V member 3 (TRPV3), Pyrroline-5-carboxylate reductase 1 (PYCR1) and ornithine aminotransferase (OAT). Such genes regulated by AR are referred to herein as "androgen receptor (AR) target genes". In various embodiments, the oligomers according to the invention are capable of inhibiting (such as, by down-regulating) the expression of one or more AR target genes in a cell which is expressing, or is capable of expressing (i.e. by alleviating AR repression of the AR target gene in a cell) an AR target gene.
[0105] The oligomers which target the androgen receptor mRNA, may hybridize to any site along the target mRNA nucleic acid, such as the 5' untranslated leader, exons, introns and 3'untranslated tail. However, it is preferred that the oligomers which target the androgen receptor mRNA hybridise to the mature mRNA form of the target nucleic acid.
[0106] Suitably, the oligomer of the invention or conjugate thereof is capable of down-regulating expression of the androgen receptor gene. In various embodiments, the oligomer (or conjugate) of the invention can effect the inhibition of androgen receptor, typically in a mammalian cell, such as a human cell. In certain embodiments, the oligomers of the invention, or conjugates thereof, bind to the target nucleic acid and effect inhibition of AR mRNA expression of at least 10% or 20% compared to the expression level immediately prior to dosing of the oligomer, more preferably of at least 30%, 40%, 50%, 60%, 70%, 80%, 90% or 95% as compared to the AR expression level immediately prior to dosing of the oligomer. In some embodiments, such inhibition is seen when using from about 0.04 nM to about 25 nM, such as from about 0.8 nM to about 20 nM of the oligomer or conjugate.
[0107] In various embodiments, the inhibition of mRNA expression is less than 100% (i.e., less than complete inhibition of expression), such as less than 98% inhibition, less than 95% inhibition, less than 90% inhibition, less than 80% inhibition, such as less than 70% inhibition. In various embodiments, modulation of gene expression can be determined by measuring protein levels, e.g. by the methods such as SDS-PAGE followed by western blotting using suitable antibodies raised against the target protein. Alternatively, modulation of expression levels can be determined by measuring levels of mRNA, e.g. by northern blotting or quantitative RT-PCR. When measuring via mRNA levels, the level of down-regulation when using an appropriate dosage, such as from about 0.04 nM to about 25 nM, such as from about 0.8 nM to about 20 nM, is, in various embodiments, typically to a level of 10-20% of the normal levels in the absence of the compound or conjugate of the invention.
[0108] The invention therefore provides a method of down-regulating or inhibiting the expression of the androgen receptor protein and/or mRNA in a cell which is expressing the androgen receptor protein and/or mRNA, the method comprising contacting the cell with an effective amount of the oligomer or conjugate according to the invention to down-regulate or inhibit the expression of the androgen receptor protein and/or mRNA in the cell. Suitably the cell is a mammalian cell, such as a human cell. The contacting may occur, in some embodiments, in vitro. The contacting may occur, in some embodiments, in vivo.
Oligomer Sequences
[0109] In some embodiments, the oligomers of the invention have sequences that are identical to a sequence selected from the group consisting of SEQ ID NOS: 2-22. Target regions in human AR mRNA (cDNA) that bind to the oligomers having sequences as set forth in SEQ ID NOs: 2-22 are shown in FIG. 4 (bold and underlined, with the corresponding oligomer SEQ ID NOs indicated above).
[0110] Further provided are target nucleic acids (e.g., DNA or mRNA encoding AR) that contain target regions that are complementary or partially-complementary to one or more of the oligomers of the invention. In certain embodiments, the oligomers bind to variants of AR target regions, such as allelic variants (such as an AR gene present at gene locus Xq11-12). In some embodiments, a variant of an AR target region has at least 60%, more preferably at least 70%, more preferably at least 80%, more preferably at least 85%, more preferably at least 90%, more preferably at least 91%, at least 92%, at least 93%, at least 94%, at least 95% sequence identity to the target region in wild-type AR. Thus, in other embodiments, the oligomers of the invention have sequences that differ in 1, 2 or 3 bases when compared to a sequence selected from the group consisting of SEQ ID NOs: 2-22. Typically, an oligomer of the invention that binds to a variant of an AR target region is capable of inhibiting (e.g., by down-regulating) AR.
[0111] In other embodiments, oligomers of the invention are LNA oligomers, for example, those oligomers having the sequences shown in SEQ ID NOs: 44-80. In various embodiments, the oligomers of the invention are potent inhibitors of androgen receptor mRNA and protein expression. In various embodiments, oligomers of the invention are LNA oligomers having the sequences of SEQ ID NO: 58 or SEQ ID NO: 77.
[0112] In various embodiments, the oligomer comprises or consists of a region having a base sequence sequence which is identical or partially identical to the sequence of the reverse complement of a target region in SEQ ID NO: 1. In various embodiments, the oligomer comprises or consists of a region having a sequence selected from the group consisting of SEQ ID NOS: 2-22 and 86-106.
[0113] In certain embodiments, the oligomer comprises or consists of a region having a base sequence which is fully complementary (perfectly complementary) to a target region of a nucleic acid which encodes a mammalian androgen receptor.
[0114] However, in some embodiments, the oligomer includes 1, 2, 3, or 4 (or more) mismatches as compared to the best-aligned target region of an AR target nucleic acid, and still sufficiently binds to the target region to effect inhibition of AR mRNA or protein expression. The destabilizing effect of mismatches on Watson-Crick hydrogen-bonded duplex may, for example, be compensated by increased length of the oligomer and/or an increased number of nucleoside analogues, such as LNA monomers, present within the oligomer.
[0115] In various embodiments, the oligomer base sequence comprises no more than 3, such as no more than 2 mismatches compared to the base sequence of the best-aligned target region of, for example, a target nucleic acid which encodes a mammalian androgen receptor.
[0116] In some embodiments, the oligomer base sequence comprises no more than a single mismatch when compared to the base sequence of the best-aligned target region of a nucleic acid which encodes a mammalian androgen receptor.
[0117] In various embodiments, the base sequence of the oligomer of the invention, or of a first region thereof, is preferably at least 80% identical to a base sequence selected from the group consisting of SEQ ID NOS: 2-22 and 86-106, such as at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96% identical, such as 100% identical.
[0118] In certain embodiments, the base sequence of the oligomer of the invention or of a first region thereof is at least 80% identical to the base sequence of the reverse complement of a target region present in SEQ ID NO: 1, such as at least 85%, at least 90%, at least 91%, at least 92% at least 93%, at least 94%, at least 95%, at least 96% identical, at least 97% identical, at least 98% identical, at least 99% identical, such as 100% identical.
[0119] In various embodiments, the base sequence of the oligomer of the invention, or of a first region thereof, is preferably at least 80% complementary to a target region of SEQ ID NO: 1, such as at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96% complementary, at least 97% complementary, at least 98% complementary, at least 99% complementary, such as 100% complementary (perfectly complementary).
[0120] In some embodiments the oligomer (or a first region thereof) has a base sequence selected from the group consisting of SEQ ID NOs: 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, and 22, or is selected from the group consisting of at least 10 contiguous monomers of SEQ ID NOs: 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, and 22. In other embodiments, the sequence of the oligomer of the invention or a first region thereof comprises one, two, or three base moieties that differ from those in oligomers having sequences of SEQ ID NOs: 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, or 22, or the sequences of at least 10 contiguous monomers thereof, when optimally aligned with the selected sequence or region thereof.
[0121] In some embodiments the oligomer (or a first region thereof) has a base sequence selected from the group consisting of SEQ ID NOs: 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103, 104, 105 and 106, or the sequences of at least 10 contiguous monomers thereof. In other embodiments, the sequence of the oligomer (or a first region thereof) comprises one, two, or three base moieties that differ from those in oligomers having sequences of SEQ ID NOs: 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103, 104, 105 or 106, or the sequences of at least 10 contiguous monomers thereof, when optimally aligned with the selected sequence or region thereof.
[0122] In various embodiments, the oligomers comprise a region of 12, 13, 14, 15 or 16 contiguous monomers having a base sequence identically present in a sequence selected from the group consisting of SEQ ID No 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, and 22. In other embodiments, the oligomers include a region which comprises one, two, or three base moieties that differ from those in oligomers having sequences of SEQ ID NOs: 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, or 22.
[0123] In some embodiments the region consists of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, or 29 contiguous monomers, such as 12-22, such as 12-18 monomers. Suitably, in some embodiments, the region is of the same length as the oligomer of the invention.
[0124] In some embodiments the oligomer comprises additional monomers at the 5' or 3' ends, such as, independently, 1, 2, 3, 4 or 5 additional monomers at the 5' end and/or the 3' end of the oligomer, which are non-complementary to the target region. In various embodiments, the oligomer of the invention comprises a region that is complementary to the target, which is flanked 5' and/or 3' by additional monomers. In some embodiments the additional 5' or 3' monomers are nucleosides, such as DNA or RNA monomers. In various embodiments, the 5' or 3' monomers represent region D as referred to in the context of gapmer oligomers herein.
[0125] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO:2, such as SEQ NO: 44, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0126] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID No: 3, such as SEQ ID NO: 45, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0127] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 4, such as SEQ ID NO: 46, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0128] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 5, such as SEQ ID NO: 47, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0129] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 6, such as SEQ ID NOs: 48, 49 or 50, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0130] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 7, such as SEQ ID NOs: 51, 52, or 53, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0131] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 8, such as SEQ ID NOs: 54, 55 or 56, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0132] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 9, such as SEQ ID NO: 57, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0133] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 10, such as SEQ ID NOs: 58, 59, or 60, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0134] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 11, such as SEQ ID NO: 61, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0135] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 12, such as SEQ ID NO: 62, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0136] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 13, such as SEQ ID NOs: 63, 64 or 65, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0137] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 14, such as SEQ ID NO: 66, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0138] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 15, such as SEQ ID NO: 67, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0139] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 16, such as SEQ ID NO: 68, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0140] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ NO: 17, such as SEQ ID NOs: 69, 70 or 71, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0141] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 18, such as SEQ ID NO: 72, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0142] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 19, such as SEQ ID NOs: 73, 74 or 75, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0143] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 20, such as SEQ ID NO: 76, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0144] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 21, such as SEQ ID NOs: 77, 78 or 79, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
[0145] In certain embodiments, the oligomer according to the invention consists of or comprises contiguous monomers having a nucleobase sequence according to SEQ ID NO: 22, such as SEQ ID NO: 80, or according to a region of at least 10 contiguous monomers thereof, such as 11, 12, 13, 14, 15 or 16 contiguous monomers thereof.
Nucleosides and Nucleoside Analogues
[0146] In various embodiments, at least one of the monomers present in the oligomer is a nucleoside analogue that contains a modified base, such as a base selected from 5-methylcytosine, isocytosine, pseudoisocytosine, 5-bromouracil, 5-propynyluracil, 6-aminopurine, 2-aminopurine, inosine, diaminopurine, 2-chloro-6-aminopurine, xanthine and hypoxanthine.
[0147] In various embodiments, at least one of the monomers present in the oligomer is a nucleoside analogue that contains a modified sugar.
[0148] In some embodiments, the linkage between at least 2 contiguous monomers of the oligomer is other than a phosphodiester linkage.
[0149] In certain embodiments, the oligomer includes at least one monomer that has a modified base, at least one monomer (which may be the same monomer) that has a modified sugar, and at least one inter-monomer linkage that is non-naturally occurring.
[0150] Specific examples of nucleoside analogues are described by e.g. Freier & Altmann; Nucl. Acid Res., 1997, 25, 4429-4443 and Uhlmann; Curr. Opinion in Drug Development, 2000, 3(2), 293-213, and in Scheme 1 (in which some nucleoside analogues are shown as nucleotides):
##STR00001## ##STR00002##
[0151] The oligomer may thus comprise or consist of a simple sequence of naturally occurring nucleosides--preferably DNA monomers, but also possibly RNA monomers, or a combination of nucleosides and one or more nucleoside analogues. In some embodiments, such nucleoside analogues suitably enhance the affinity of the oligomer for the target region of the target nucleic acid.
[0152] Examples of suitable and preferred nucleoside analogues are described in PCT/DK2006/000512, or are referenced therein.
[0153] In some embodiments, the nucleoside analogue comprises a sugar moiety modified to provide a 2'-substituent group, such as 2'-O-alkyl-ribose sugars, 2'-amino-deoxyribose sugars, and 2'-fluoro-deoxyribose sugars.
[0154] In some embodiments, the nucleoside analogue comprises a sugar in which a bridged structure, creating a bicyclic sugar (LNA), which enhances binding affinity and may also provide some increased nuclease resistance. In various embodiments, the LNA monomer is selected from oxy-LNA (such as beta-D-oxy-LNA, and alpha-L-oxy-LNA), and/or amino-LNA (such as beta-D-amino-LNA and alpha-L-amino-LNA) and/or thio-LNA (such as beta-D-thio-LNA and alpha-L-thio-LNA) and/or ENA (such as beta-D-ENA and alpha-L-ENA). In certain embodiments, the LNA monomers are beta-D-oxy-LNA. LNA monomers are further described below.
[0155] In various embodiments, incorporation of affinity-enhancing nucleoside analogues in the oligomer, such as LNA monomers or monomers containing 2'-substituted sugars, or incorporation of modified linkage groups provides increased nuclease resistance. In various embodiments, incorporation of affinity-enhancing nucleoside analogues allows the size of the oligomer to be reduced, and also reduces the size of the oligomer that binds specifically to a target region of a target sequence.
[0156] In some embodiments, the oligomer comprises at least 2 nucleoside analogues. In some embodiments, the oligomer comprises from 3-8 nucleoside analogues, e.g. 6 or 7 nucleoside analogues. In various embodiments, at least one of the nucleoside analogues is a locked nucleic acid (LNA) monomer; for example at least 3 or at least 4, or at least 5, or at least 6, or at least 7, or 8, nucleoside analogues are LNA monomers. In some embodiments, all the nucleoside analogues are LNA monomers.
[0157] It will be recognised that when referring to a preferred oligomer base sequence, in certain, embodiments, the oligomers comprise a corresponding nucleoside analogue, such as a corresponding LNA monomer or other corresponding nucleoside analogue, which raise the duplex stability (T.sub.m) of the oligomer/target region duplex (i.e. affinity enhancing nucleoside analogues).
[0158] In various embodiments, any mismatches (i.e., non-complementarities) between the base sequence of the oligomer and the base sequence of the target region, if present, are preferably located other than in the regions of the oligomer that contain affinity-enhancing nucleoside analogues (e.g., regions A or C), such as within region B as referred to herein, and/or within region D as referred to herein, and/or in regions consisting of DNA monomers, and/or in regions which are 5' or 3' to the region of the oligomer that is complementary to the target region.
[0159] In some embodiments the nucleoside analogues present within the oligomer of the invention (such as in regions A and C mentioned herein) are independently selected from, for example: monomers containing 2'-O-alkyl-ribose sugars, monomers containing 2'-amino-deoxyribose sugars, monomers containing 2'-fluoro-deoxyribose sugars, LNA monomers, monomers containing arabinose sugars ("ANA monomers"), monomers containing 2'-fluoro-arabinose sugars, monomers containing d-arabino-hexitol sugars ("RNA monomers"), intercalating monomers as defined in Christensen (2002) Nucl. Acids. Res. 30: 4918-4925, hereby incorporated by reference, and 2'-O-methoxyethyl-ribose (2'MOE) sugars. In some embodiments, there is only one of the above types of nucleoside analogues present in the oligomer of the invention, or region thereof.
[0160] In certain embodiments, the nucleoside analogues contain 2'MOE sugars, 2'-fluoro-deoxyribose sugars, or LNA sugars, and as such the oligonucleotide of the invention may comprise nucleoside analogues which are independent) selected from these three types. In certain oligomer embodiments containing nucleoside analogues, at least one of said nucleoside analogues contains a 2'-MOE-ribose sugar, such as 2, 3, 4, 5, 6, 7, 8, 9 or 10 nucleoside analogues containing 2'-MOE-ribose sugars. In some embodiments, at least one nucleoside analogue contains a 2'-fluoro-deoxyribose sugar, such as 2, 3, 4, 5, 6, 7, 8, 9 or 10 nucleoside analogues containing 2'-fluoro-DNA nucleotide sugars.
[0161] In various embodiments, the oligomer according to the invention comprises at least one Locked Nucleic Acid (LNA) monomer, such as 1, 2, 3, 4, 5, 6, 7, or 8 LNA monomers, such as 3-7 or 4 to 8 LNA monomers, or 3, 4, 5, 6 or 7 LNA monomers. In various embodiments, all the nucleoside analogues are LNA monomers. In certain embodiments, the oligomer comprises both beta-D-oxy-LNA monomers, and one or more of the following LNA monomers: thio-LNA monomers, amino-LNA monomers, oxy-LNA monomers, and/or ENA monomers in either the beta-D or alpha-L configurations, or combinations thereof. In certain embodiments, the cytosine base moieties of all LNA monomers in the oligomer are 5-methylcytosines. In certain embodiments of the invention, the oligomer comprises both LNA and DNA monomers. Typically, the combined total of LNA and DNA monomers is 10-25, preferably 10-20, even more preferably 12-16. In some embodiments of the invention, the oligomer or region thereof consists of at least one LNA monomer, and the remaining monomers are DNA monomers. In certain embodiments, the oligomer comprises only LNA monomers and nucleosides (such as RNA or DNA monomers, most preferably DNA monomers) optionally with modified linkage groups such as phosphorothioate.
[0162] In various embodiments, at least one of the nucleoside analogues present in the oligomer has a modified base selected from the group consisting of 5-methylcytosine, isocytosine, pseudoisocytosine, 5-bromouracil, 5-propynyluracil, 6-aminopurine, 2-aminopurine, inosine, diaminopurine, and 2-chloro-6-aminopurine.
LNA
[0163] The term "LNA monomer" refers to a nucleoside analogue containing a bicyclic sugar (an "LNA sugar"). The terms "LNA oligonucleotide" and "LNA oligomer" refer to an oligomer containing one or more LNA monomers.
[0164] The LNA used in the oligonucleotide compounds of the invention preferably has the structure of the general formula I:
##STR00003##
[0165] wherein X is selected from --O--, --S--, --N(R.sup.N*)--, --C(R.sup.6R.sup.6*)--;
[0166] B is selected from hydrogen, optionally substituted C.sub.1-4-alkoxy, optionally substituted C.sub.1-4alkyl, optionally substituted C.sub.1-4-acyloxy, nucleobases, DNA intercalators, photochemically active groups, thermochemically active groups, chelating groups, reporter groups, and ligands;
[0167] P designates the radical position for an internucleoside linkage to a succeeding monomer, or a 5'-terminal group, such internucleoside linkage or 5'-terminal group optionally including the substituent R.sup.5 or equally applicable the substituent R.sup.5*;
[0168] P* designates an internucleoside linkage to a preceding monomer, or a 3'-terminal group;
[0169] R.sup.4* and R.sup.2* together designate a biradical consisting of 1-4 groups/atoms selected from --C(R.sup.aR.sup.b)--, --C(R.sup.a).dbd.C(R.sup.b)--, --C(R.sup.a).dbd.N--, --O--, --Si(R.sup.b).sub.2, --S--, --SO.sub.2--, --N(R.sup.a)--, and >C.dbd.Z,
[0170] wherein Z is selected from --O--, --S--, and --N(R.sup.a)--, and R.sup.a and R.sup.b each is independently selected from hydrogen, optionally substituted C.sub.1-12-alkyl, optionally substituted C.sub.2-12-alkenyl, optionally substituted C.sub.2-12-alkynyl, hydroxy, C.sub.1-12-alkoxy, C.sub.2-12-alkoxyalkyl, C.sub.2-12-alkenyloxy, carboxy, C.sub.1-12-alkoxycarbonyl, C.sub.1-12-alkylcarbonyl, formyl, aryl, aryloxy-carbonyl, aryloxy, arylcarbonyl, heteroaryl, heteroaryloxy-carbonyl, heteroaryloxy, heteroarylcarbonyl, amino, mono- and di(C.sub.1-6-alkyl)amino, carbamoyl, mono- and di(C.sub.1-6alkyl)-amino-carbonyl, amino-C.sub.1-6-alkyl-aminocarbonyl, mono- and di(C.sub.1-6-alkyl)amino-C.sub.1-6-alkyl-aminocarbonyl, C.sub.1-6-alkyl-carbonylamino, carbamido, C.sub.1-6-alkanoyloxy, sulphono, C.sub.1-6-alkylsulphonyloxy, nitro, azido, sulphanyl, C.sub.1-6-alkylthio, halogen, DNA intercalators, photochemically active groups, thermochemically active groups, chelating groups, reporter groups, and ligands, where aryl and heteroaryl may be optionally substituted and where two geminal substituents R.sup.a and R.sup.b together may designate optionally substituted methylene (.dbd.CH.sub.2), and
[0171] each of the substituents R.sup.1*, R.sup.2, R.sup.3, R.sup.5, R.sup.5*, R.sup.6 and R.sup.6*, which are present is independently selected from hydrogen, optionally substituted C.sub.1-12-alkyl, optionally substituted C.sub.2-12-alkenyl, optionally substituted C.sub.2-12-alkynyl, hydroxy, C.sub.1-12-alkoxy, C.sub.2-12-alkoxyalkyl, C.sub.2-12-alkenyloxy, carboxy, C.sub.1-12-alkoxycarbonyl, C.sub.1-12-alkylcarbonyl, formyl, aryl, aryloxy-carbonyl, aryloxy, arylcarbonyl, heteroaryl, heteroaryloxy-carbonyl, heteroaryloxy, heteroarylcarbonyl, amino, mono- and di(C.sub.1-6alkyl)amino, carbamoyl, mono- and di(C.sub.1-6-alkyl)-amino-carbonyl, amino-C.sub.1-6alkyl-aminocarbonyl, mono- and di(C.sub.1-6-alkyl)amino-C.sub.1-6-alkyl-aminocarbonyl, C.sub.1-6-alkyl-carbonylamino, carbamido, C.sub.1-6-alkanoyloxy, sulphono, C.sub.1-6-alkylsulphonyloxy, nitro, azido, sulphanyl, C.sub.1-6-alkylthio, halogen, DNA intercalators, photochemically active groups, thermochemically active groups, chelating groups, reporter groups, and ligands, where aryl and heteroaryl may be optionally substituted, and where two geminal substituents together may designate oxo, thioxo, imino, or optionally substituted methylene, or together may form a spiro biradical consisting of a 1-5 carbon atom(s) alkylene chain which is optionally interrupted and/or terminated by one or more heteroatoms/groups selected from --O--, --S', and --(NR.sup.N)-- where R.sup.N is selected from hydrogen and C.sub.1-4-alkyl, and where two adjacent (non-geminal) substituents may designate an additional bond resulting in a doable bond; and R.sup.N*, when present and not involved in a biradical, is selected from hydrogen and C.sub.1-4 alkyl; and basic salts and acid addition salts thereof;
[0172] In some embodiments, R.sup.5* is selected from H, --CH.sub.3, --CH.sub.2--CH.sub.3, --CH.sub.2--O--CH.sub.3, and --CH.dbd.CH.sub.2.
[0173] In various embodiments, R.sup.4* and R.sup.2* together designate a biradical selected from --C(R.sup.aR.sup.b)--O--, --C(R.sup.aR.sup.b)--C(R.sup.cR.sup.d)--O--, --C(R.sup.aR.sup.b)--C(R.sup.cR.sup.d)--C(R.sup.eR.sup.f)--O--, --C(R.sup.aR.sup.b)--O--C(R.sup.cR.sup.d)--, --C(R.sup.aR.sup.b)--O--C(R.sup.cR.sup.d)--O--, --C(R.sup.aR.sup.b)--C(R.sup.cR.sup.d)--, --C(R.sup.aR.sup.b)--C(R.sup.cR.sup.d)--C(R.sup.eR.sup.f)--, --C(R.sup.a).dbd.C(R.sup.b)--C(R.sup.cR.sup.d)--, --C(R.sup.aR.sup.b)--N(R.sup.c)--, --C(R.sup.aR.sup.b)--C(R.sup.cR.sup.d)--N(R.sup.e)--, --C(R.sup.aR.sup.b)--N(R.sup.c)--O--, and --C(R.sup.aR.sup.b)--S--, --C(R.sup.aR.sup.b)--C(R.sup.cR.sup.d)--S--, wherein R.sup.a, R.sup.b, R.sup.c, R.sup.d, R.sup.e, and R.sup.f each is independently selected from hydrogen, optionally substituted C.sub.1-12-alkyl, optionally substituted C.sub.2-12-alkenyl, optionally substituted C.sub.2-12-alkynyl, hydroxy, C.sub.1-12-alkoxy, C.sub.2-12-alkoxyalkyl, C.sub.2-12-alkenyloxy, carboxy, C.sub.1-12-alkoxycarbonyl, formyl, aryl, aryloxy-carbonyl, aryloxy, arylcarbonyl, heteroaryl, heteroaryloxy-carbonyl, heteroaryloxy, heteroarylcarbonyl, amino, mono- and di(C.sub.1-6-alkyl)amino, carbamoyl, mono- and di(C.sub.1-6-alkyl)-amino-carbonyl, amino-C.sub.1-6-alkyl-aminocarbonyl, mono- and di(C.sub.1-6alkyl)amino-C.sub.1-6-alkyl-aminocarbonyl, C.sub.1-6-alkyl-carbanylamino, carbamido, C.sub.1-6alkanoyloxy, sulphono, C.sub.1-6-alkylsulphonyloxy, nitro, azido, sulphanyl, C.sub.1-6-alkylthio, halogen, DNA intercalators, photochemically active groups, thermochemically active groups, chelating groups, reporter groups, and ligands, where aryl and heteroaryl may be optionally substituted and where two geminal substituents R.sup.a and R.sup.b together may designate optionally substituted methylene (.dbd.CH.sub.2),
[0174] In a further embodiment R.sup.4* and R.sup.2* together designate a biradical (bivalent group) selected from --CH.sub.2--O--, --CH.sub.2--S--, --CH.sub.2--NH--, --CH.sub.2--N(CH.sub.2)--, --CH.sub.2--CH.sub.2--O--, --CH.sub.2--CH(CH.sub.2)--, --CH.sub.2--CH.sub.2--S--, --CH.sub.2--CH.sub.2--NH--, --CH.sub.2--CH.sub.2--CH.sub.2--, --CH.sub.2--CH.sub.2--CH.sub.2--O--, --CH.sub.2--CH.sub.2--CH(CH.sub.2)--, --CH.dbd.CH--CH.sub.2, --CH.sub.2--O--CH.sub.2--O--, --CH.sub.2--NH--O--, --CH.sub.2--N(CH.sub.2)--O--, --CH.sub.2--O--CH.sub.2--, --CH(CH.sub.2)--O--, --CH(CH.sub.2--O--CH.sub.2)--O--.
[0175] For all chiral centers, asymmetric groups may be found in either R or S orientation.
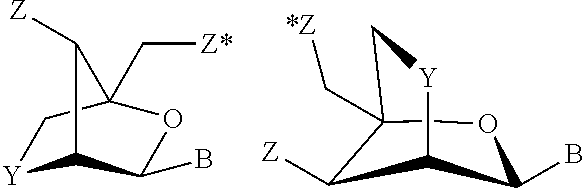
[0176] Preferably, the LNA monomer used in the oligomer of the invention comprises at least one LNA monomer according to any of the formulas
##STR00004##
[0177] wherein Y is --O--, --O--CH2-, --S--, --NH--, or N(RH); Z and Z* are independently selected among an internucleotide linkage, a terminal group or a protecting group; B constitutes a natural or non-natural nucleotide base moiety, and R.sup.H is selected from hydrogen and C.sub.1-4-alkyl.
[0178] Specifically preferred LNA monomers are shown in Scheme 2:
##STR00005##
[0179] The term "thio-LNA" refers to an LNA monomer in which Y in the general formula above is selected from S or --CH.sub.2--S--. Thio-LNA can be in either the beta-D or alpha-L-configuration.
[0180] The term "amino-LNA" refers to an LNA monomer in which Y in the general formula above is selected from --N(H)--, N(R)--, CH.sub.2--N(H)--, and --CH.sub.2--N(R)-- where R is selected from hydrogen and C.sub.1-4-alkyl. Amino-LNA can be in either the beta-D or alpha-L-configuration.
[0181] The term "oxy-LNA" refers to an LNA monomer in which Y in the general formula above represents --O-- or --CH.sub.2--O--. Oxy-LNA can be in either the beta-D or alpha-L-configuration.
[0182] The term "ENA" refers to an LNA monomer in which Y in the general formula above is --CH.sub.2--O-- (where the oxygen atom of --CH.sub.2--O-- is attached to the 2'-position relative to the base B).
[0183] In various embodiments, the LNA monomer is selected from a beta-D-oxy-LNA monomer, and alpha-L-oxy-LNA monomer, a beta-D-amino-LNA monomer, and beta-D-thio-LNA monomer, in particular a beta-D-oxy-LNA monomer.
[0184] In the present context, the term "C.sub.1-4alkyl" means a linear or branched saturated hydrocarbon chain wherein the chain has from one to four carbon atoms, such as methyl, ethyl, n-propyl, isopropyl, n-butyl, isobutyl, sec-butyl and tert-butyl.
RNAse H Recruitment
[0185] In some embodiments, an oligomer functions via non-RNase-mediated degradation of a target mRNA, such as by steric hindrance of translation, or other mechanisms; however, in various embodiments, oligomers of the invention are capable of recruiting an endo-ribonuclease (RNase), such as RNase H.
[0186] Typically, the oligomer, comprises a region of at least 6, such as at least 7 contiguous monomers, such as at least 8 or at least 9 contiguous monomers, including 7, 8, 9, 10, 11, 12, 13, 14, 15 or 16 contiguous monomers, which, when forming a duplex with the target region of the target RNA, is capable a recruiting RNase. The region of the oligomer which is capable of recruiting RNAse may be region B, as referred to in the context of a gapmer as described herein. In some embodiments, the region of the oligomer which is capable of recruiting RNAse, such as region B, consists of 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 monomers.
[0187] EP 1 222 309 provides in vitro methods for determining RNaseH activity, which may be used to determine the ability of the oligomers of the invention to recruit RNaseH. An oligomer is deemed capable of recruiting RNaseH if, when contacted with the complementary region of the RNA target, it has an initial rate, as measured in pmol/l/min, of at least 1%, such as at least 5%, such as at least 10% or less than 20% of an oligonucleotide having the same base sequence but containing only DNA monomers, with no 2' substitutions, with phosphorothioate linkage groups between all monomers in the oligonucleotide, using the methodology provided by Examples 91-95 of EP 1 222 309, incorporated herein by reference.
[0188] In some embodiments, an oligomer is deemed essentially incapable of recruiting RNaseH if, when contacted with the complementary target region of the RNA target, and RNaseH, the RNaseH initial rate, as measured in pmol/l/min, is less than 1%, such as less than 5%, such as less than 10% or less than 20% of the initial rate determined using an oligonucleotide having the same base sequence, but containing only DNA monomers, with no 2' substitutions, with phosphorothioate linkage groups between all monomers in the oligonucleotide, using the methodology provided by Examples 91-95 of EP 1 222 309.
[0189] In other embodiments, an oligomer is deemed capable of recruiting RNaseH if, when contacted with the complementary target region of the RNA target, and RNaseH, the RNaseH initial rate, as measured in pmol/l/min, is at least 20%, such as at least 40%, such as at least 60%, such as at least 80% of the initial rate determined using an oligonucleotide having the same base sequence, but containing only DNA monomers, with no 2' substitutions, with phosphorothioate linkage groups between all monomers in the oligonucleotide, using the methodology provided by Examples 91-95 of EP 1 222 309.
[0190] Typically, the region of the oligomer which forms the duplex with the complementary target region of the target RNA and is capable of recruiting RNase contains DNA monomers and LNA monomers and forms a DNA/RNA-like duplex with the target region. The LNA monomers are preferably in the alpha-L configuration, particularly preferred being alpha-L-oxy LNA.
[0191] In various embodiments, the oligomer of the invention comprises both nucleosides and nucleoside analogues, and is in the form of a gapmer, a headmer or a mixmer.
[0192] A "headmer" is defined as an oligomer that comprises a first region and a second region that is contiguous thereto, with the 5'-most monomer of the second region linked to the 3'-most monomer of the first region. The first region comprises a contiguous stretch of non-RNase recruiting nucleoside analogues and the second region comprises a contiguous stretch (such as at least 7 contiguous monomers) of DNA monomers or nucleoside analogue monomers recognizable and cleavable by the RNase
[0193] A "tailmer" is defined as an oligomer that comprises a first region and a second region that is contiguous thereto, with the 5'-most monomer of the second region linked to the 3'-most monomer of the first region. The first region comprises a contiguous stretch (such as at least 7 contiguous monomers) of DNA monomers or nucleoside analogue monomers recognizable and cleavable by the RNase, and the second region comprises a contiguous stretch of non-RNase recruiting nucleoside analogues.
[0194] Other "chimeric" oligomers, called "mixmers", consist of an alternating composition of (i) DNA monomers or nucleoside analogue monomers recognizable and cleavable by RNase, and (ii) non-RNase recruiting nucleoside analogue monomers.
[0195] In some embodiments, in addition to enhancing affinity of the oligomer for the target region, some nucleoside analogues also mediate RNase (e.g., RNaseH) binding and cleavage. Since -L-LNA monomers recruit RNaseH activity to a certain extent, in some embodiments, gap regions (e.g., region B as referred to herein) of oligomers containing -L-LNA monomers consist of fewer monomers recognizable and cleavable by the RNaseH, and more flexibility in the mixmer construction is introduced.
Conjugates
[0196] In the context of this disclosure, the term "conjugate" indicates a compound formed by the covalent attachment ("conjugation") of an oligomer as described herein, to one or more moieties that are not themselves nucleic acids or monomers ("conjugated moieties"). Examples of such conjugated moieties include macromolecular compounds such as proteins, fatty acid chains, sugar residues, glycoproteins, polymers, or combinations thereof. Typically proteins may be antibodies for a target protein. Typical polymers may be polyethylene glycol.
[0197] Accordingly, provided herein are conjugates comprising an oligomer as herein described, and at least one conjugated moiety that is not a nucleic acid or monomer, covalently attached to said oligomer. Therefore, in certain embodiments where the oligomer of the invention consists of contiguous monomers having a specified sequence of bases, as herein disclosed, the conjugate may also comprise at least one conjugated moiety that is covalently attached to the oligomer.
[0198] In various embodiments of the invention, the oligomer is conjugated to a moiety that increases the cellular uptake of oligomeric compounds. WO2007/031091 provides suitable ligands and conjugates, which are hereby incorporated by reference.
[0199] In various embodiments, conjugation (to a conjugated moiety) may enhance the activity, cellular distribution or cellular uptake of the oligomer of the invention. Such moieties include, but are not limited to, antibodies, polypeptides, lipid moieties such as a cholesterol moiety, cholic acid, a thioether, e.g. Hexyl-s-tritylthiol, a thiocholesterol, an aliphatic chain, e.g., dodecandiol or undecyl residues, a phospholipids, e.g., di-hexadecyl-rac-glycerol or triethylammonium 1,2-di-o-hexadecyl-rac-glycero-3-h-phosphonate, a polyamine or a polyethylene glycol chain, an adamantane acetic acid, a palmityl moiety, an octadecylamine or hexylamino-carbonyl-oxycholesterol moiety.
[0200] In certain embodiments, the oligomers of the invention are conjugated to active drug substances, for example, aspirin, ibuprofen, a sulfa drug, an antidiabetic, an antibacterial or an antibiotic.
[0201] In certain embodiments the conjugated moiety is a sterol, such as cholesterol.
[0202] In various embodiments, the conjugated moiety comprises or consists of a positively charged polymer, such as a positively charged peptides of, for example 1-50, such as 2-20 such as 3-10 amino acid residues in length, and/or polyalkylene oxide such as polyethylene glycol (PEG) or polypropylene glycol--see WO 2008/034123, hereby incorporated by reference. Suitably the positively charged polymer, such as a polyalkylene oxide may be attached to the oligomer of the invention via a linker such as the releasable linker described in WO 2008/034123.
[0203] By way of example, the following moieties may be used in the conjugates of the invention:
##STR00006##
Activated Oligomers
[0204] The term "activated oligomer," as used herein, refers to an oligomer of the invention that is covalently linked (i.e., functionalized) to at least one functional moiety that permits covalent linkage of the oligomer to one or more conjugated moieties, i.e., moieties that are not themselves nucleic acids or monomers, to form the conjugates herein described. Typically, a functional moiety will comprise a chemical group that is capable of covalently bonding to the oligomer via, e.g., a 3'-hydroxyl group or the exocyclic NH.sub.2 group of the adenine base, a spacer that is preferably hydrophilic and a terminal group that is capable of binding to a conjugated moiety (e.g., an amino, sulfhydryl or hydroxyl group). In some embodiments, this terminal group is not protected, e.g., is an NH.sub.2 group. In other embodiments, the terminal group is protected, for example, by any suitable protecting group such as those described in "Protective Groups in Organic Synthesis" by Theodora W. Greene and Peter G. M. Wuts, 3rd edition (John Wiley & Sons, 1999). Examples of suitable hydroxyl protecting groups include esters such as acetate ester, aralkyl groups such as benzyl, diphenylmethyl, or triphenylmethyl, and tetrahydropyranyl. Examples of suitable amino protecting groups include benzyl, alpha-methylbenzyl, diphenylmethyl, triphenylmethyl, benzyloxycarbonyl, tert-butoxycarbonyl, and acyl groups such as trichloroacetyl or trifluoroacetyl.
[0205] In some embodiments, the functional moiety is self-cleaving. In other embodiments, the functional moiety is biodegradable. See e.g., U.S. Pat. No. 7,087,229, which is incorporated by reference herein in its entirety.
[0206] In some embodiments, oligomers of the invention are functionalized at the 5' end in order to allow covalent attachment of the conjugated moiety to the 5' end of the oligomer. In other embodiments, oligomers of the invention can be functionalized at the 3' end. In still other embodiments, oligomers of the invention can be functionalized along the backbone or on the heterocyclic base moiety. In yet other embodiments, oligomers of the invention can be functionalized at more than one position independently selected from the 5' end, the 3' end, the backbone and the base.
[0207] In some embodiments, activated oligomers of the invention are synthesized by incorporating during the synthesis one or more monomers that is covalently attached to a functional moiety. In other embodiments, activated oligomers of the invention are synthesized with, monomers that have not been functionalized, and the oligomer is functionalized upon completion of synthesis.
[0208] In some embodiments, the oligomers are functionalized with a hindered ester containing an aminoalkyl linker, wherein the alkyl portion has the formula (CH.sub.2).sub.n, wherein w is an integer ranging from 1 to 10, preferably about 6, wherein the alkyl portion of the alkylamino group can be straight chain or branched chain, and wherein the functional group is attached to the oligomer via an ester group (--O--C(O)--(CH.sub.2).sub.nNH).
[0209] In other embodiments, the oligomers are functionalized with a hindered ester containing a (CH.sub.2).sub.w-sulfhydryl (SH) linker, wherein w is an integer ranging from 1 to 10, preferably about 6, wherein the alkyl portion of the alkylamino group can be straight chain or branched chain, and wherein the functional group attached to the oligomer via an ester group (--O--C(O)--(CH.sub.2).sub.wSH) In some embodiments, sulfhydryl-activated oligonucleotides are conjugated with polymer moieties such as polyethylene glycol or peptides (via formation of a disulfide bond).
[0210] Activated oligomers containing hindered esters as described above can be synthesized by any method known in the art, and in particular, by methods disclosed in PCT Publication No. WO 2008/034122 and the examples therein, which is incorporated herein by reference in its entirety.
[0211] Activated oligomers covalently linked to at least one functional moiety can be synthesized by any method known in the art, and in particular, by methods disclosed in U.S. Patent Publication No. 2004/0235773, which is incorporated herein by reference in its entirety, and in Zhao et al. (2007) J. Controlled Release 119:143-152; and Zhao et al. (2005) Bioconjugate Chem. 16:758-766.
[0212] In still other embodiments, the oligomers of the invention are functionalized by introducing sulfhydryl, amino or hydroxyl groups into the oligomer by means of a functionalizing reagent substantially as described in U.S. Pat. Nos. 4,962,029 and 4,914,210, i.e., a substantially linear reagent having a phosphoramidite at one end linked through a hydrophilic spacer chain to the opposing end which comprises a protected or unprotected sulfhydryl, amino or hydroxyl group. Such reagents primarily react with hydroxyl groups of the oligomer. In some embodiments, such activated oligomers have a functionalizing reagent coupled to a 5'-hydroxyl group of the oligomer. In other embodiments, the activated oligomers have a functionalizing reagent coupled to a 3'-hydroxyl group. In still other embodiments, the activated oligomers of the invention have a functionalizing reagent coupled to a hydroxyl group on the backbone of the oligomer. In yet further embodiments, the oligomer of the invention is functionalized with more than one of the functionalizing reagents as described in U.S. Pat. Nos. 4,962,029 and 4,914,210, incorporated herein by reference in their entirety. Methods of synthesizing such functionalizing reagents and incorporating them into monomers or oligomers are disclosed in U.S. Pat. Nos. 4,962,029 and 4,914,210.
[0213] In some embodiments, the 5'-terminus of a solid-phase bound oligomer is functionalized with a dienyl phosphoramidite derivative, followed by conjugation of the deprotected oligomer with, e.g., an amino acid or peptide via a Diels-Alder cycloaddition reaction.
[0214] In various embodiments, the incorporation of monomers containing 2'-sugar modifications, such as a 2'-carbamate substituted sugar or a 2'-(O-pentyl-N-phthalimido)-deoxyribose sugar into the oligomer facilitates covalent attachment of conjugated moieties to the sugars of the oligomer. In other embodiments, an oligomer with an amino-containing linker at the 2'-position of one or more monomers is prepared using a reagent such as, for example, 5'-dimethoxytrityl-2'-O-(e-phthalimidylaminopentyl)-2'-deoxyadenosine-3'-- N,N-diisopropyl-cyanoethoxy phosphoramidite. See, e.g., Manoharan, et al., Tetrahedron Letters, 1991, 34, 7171.
[0215] In still further embodiments, the oligomers of the invention have amine-containing functional moieties on the nucleobase, including on the N6 purine amino groups, on the exocyclic N2 of guanine, or on the N4 or 5 positions of cytosine. In various embodiments, such functionalization may be achieved by using a commercial reagent that is already functionalized in the oligomer synthesis.
[0216] Some functional moieties are commercially available, for example, heterobifunctional and homobifunctional linking moieties are available from the Pierce Co. (Rockford, Ill.). Other commercially available linking groups are 5'-Amino-Modifier C6 and 3'-Amino-Modifier reagents, both available from Glen Research Corporation (Sterling, Va.). 5'-Amino-Modifier C6 is also available from ABI (Applied Biosystems Inc., Foster City, Calif.) as Aminolink-2, and 3'-Amino-Modifier is also available from Clontech Laboratories Inc. (Palo Alto, Calif.).
Compositions
[0217] In various embodiments, the oligomer of the invention is used in pharmaceutical formulations and compositions. Suitably, such compositions comprise a pharmaceutically acceptable diluent, carrier, salt or adjuvant. PCT/DK2006/000512 provides suitable and preferred pharmaceutically acceptable diluents, carriers and adjuvants--which are hereby incorporated by reference. Suitable dosages, formulations, administration routes, compositions, dosage forms, combinations with other therapeutic agents, pro-drug formulations are also provided in PCT/DK2006/000512--which are also hereby incorporated by reference. Details on techniques for formulation and administration also may be found in the latest edition of "REMINGTON'S PHARMACEUTICAL SCIENCES" (Maack Publishing Ca, Easton Pa.),
[0218] In some embodiments, an oligomer of the invention is covalently linked to a conjugated moiety to aid in delivery of the oligomer across cell membranes. An example of a conjugated moiety that aids in delivery of the oligomer across cell membranes is a lipophilic moiety, such as cholesterol. In various embodiments, an oligomer of the invention is formulated with lipid formulations that form liposomes, such as Lipofectamine 2000 or Lipofectamine RNAiMAX, both of which are commercially available from Invitrogen. In some embodiments, the oligomers of the invention are formulated with a mixture of one or more lipid-like non-naturally occurring small molecules ("lipidoids"). Libraries of lipidoids can be synthesized by conventional synthetic chemistry methods and various amounts and combinations of lipidoids can be assayed in order to develop a vehicle for effective delivery of an oligomer of a particular size to the targeted tissue by the chosen route of administration. Suitable lipidoid libraries and compositions can be found, for example in Akinc et at. (2008) Nature Biotechnol., available at http://www.nature.com/nbt/journal/vaop/incurrent/abs/nbt1402.html, which is incorporated by reference herein.
[0219] As used herein, the term "pharmaceutically acceptable salts" refers to salts that retain the desired biological activity of the herein identified compounds and exhibit acceptable levels of undesired toxic effects. Non-limiting examples of such salts can be formed with organic amino acid and base addition salts formed with metal cations such as zinc, calcium, bismuth, barium, magnesium, aluminum, copper, cobalt, nickel, cadmium, sodium, potassium, and the like, or with a cation formed from ammonia, N,N'-dibenzylethylene-diamine, D-glucosamine, tetraethylammonium, or ethylenediamine; or (c) combinations of (a) and (b); e.g., a zinc tannate salt or the like.
[0220] In certain embodiments, the pharmaceutical compositions according to the invention comprise other active ingredients in addition to an oligomer or conjugate of the invention, including active agents useful for the treatment of cancer, such as prostate cancer or breast cancer, particularly agents used in conventional antiandrogen therapy.
[0221] In some embodiments, additional active agents are non-steroidal antiandrogens (NSAAs), which block the binding of androgens at the receptor site, or luteinizing hormone-releasing hormone analogues (LHRH-As) that suppress testicular production of androgens to castrate levels.
[0222] NSAAs such as CASODEX, when used with an LHRH-A as part of Combined Androgen Blockade therapy, help to inhibit the growth of prostate cancer cells. In one embodiment, the invention provides for a combined androgen blockade therapy, characterised in that the therapy comprises administering the pharmaceutical composition according to the invention, and an NSAA and/or LHRH-A agent, which in certain embodiments are administered prior to, during or subsequent to the administration of the pharmaceutical compositions of the invention.
[0223] The invention also provides a kit of parts wherein a first part comprises at least one oligomer, conjugate and/or the pharmaceutical composition according to the invention and a further part comprises a non-steroidal antiandrogen and/or a luteinizing hormone-releasing hormone analogue. It is therefore envisaged that the kit of parts may be used in a method of treatment, as referred to herein, where the method comprises administering both the first part and the further part, either simultaneously or one after the other.
Applications
[0224] The term "treatment" as used herein refers to both treatment of an existing disease (e.g., a disease or disorder as referred to herein below), or prevention of a disease, i.e., prophylaxis. It will therefore be recognised that, in certain embodiments, "treatment" includes prophylaxis.
[0225] In various embodiments, the oligomers of the invention may be utilized as research reagents for, for example, diagnostics, therapeutics and prophylaxis.
[0226] In some embodiments, such oligomers may be used for research purposes to specifically inhibit the expression of androgen receptor protein (typically by degrading or inhibiting the AR mRNA and thereby preventing protein formation) in cells and experimental animals, thereby facilitating functional analysis of the target or an appraisal of its usefulness as a target for therapeutic intervention.
[0227] In certain embodiments, the oligomers may be used in diagnostics to detect and quantitate androgen receptor expression in cells and tissues by Northern blotting, in-situ hybridisation or similar techniques.
[0228] In various therapeutic embodiments, a non-human animal or a human suspected of having a disease or disorder which can be treated by modulating the expression of androgen receptor is treated by administering an effective amount of an oligomer in accordance with this invention. Further provided are methods of treating a mammal, such as treating a human, suspected of having or being prone to a disease or condition, associated with expression of androgen receptor by administering a therapeutically or prophylactically effective amount of one or more of the oligomers, conjugates or compositions of the invention.
[0229] In certain embodiments, the invention also provides for the use of the compounds or conjugates of the invention as described for the manufacture of a medicament for the treatment of a disorder as referred to herein, or for a method of the treatment of a disorder as referred to herein.
[0230] In various embodiments, the invention also provides for a method for treating a disorder as referred to herein, said method comprising administering a compound according to the invention as herein described, and/or a conjugate according to the invention, and/or a pharmaceutical composition according to the invention to a patient in need thereof.
Medical Indications
[0231] In certain therapeutic embodiments, the disorder to be treated is cancer, such as prostate cancer or breast cancer. In various embodiments, the treatment of such a disease or condition according to the invention may be combined with one or more other anti-cancer treatments, such as radiotherapy, chemotherapy or immunotherapy.
[0232] In certain other embodiments, the disorder to be treated is selected from alopecia, benign prostatic hyperplasia, spinal and muscular atrophy and Kennedy disease and polyglutamate disease.
[0233] In various embodiments, the disease or disorder is associated with a mutation of the AR gene or a gene whose protein product is associated with or interacts with AR. Therefore, in various embodiments, the target mRNA is a mutated form of the AR sequence, for example, it comprises one or more single point mutations or triplet repeats.
[0234] In other embodiments, the disease or disorder is associated with abnormal levels of a mutated form of androgen receptor. In various embodiments, the disease or disorder is associated with abnormal levels of a wild-type form of AR.
[0235] In various embodiments, the invention relates to methods of modulating the expression of the gene product of an androgen receptor target gene, i.e., a gene that is regulated by AR. Such AR receptor target gene products are selected form the group consisting of Protein kinase C delta (PPKCD), Glutathione S-transferase theta 2 (GSTT2), transient receptor potential cation channel subfamily V member 3 (TRPV3), Pyrroline-5-carboxylate reductase 1 (PYCR1) and ornithine aminotransferase (OAT). In some embodiments, modulation of an AR target gene results in increased expression or activity of the target gene. In other embodiments, modulation of an AR target gene results in decreased expression or activity of the target gene.
[0236] The invention further provides use of a compound of the invention in the manufacture of a medicament for the treatment of any and all conditions disclosed herein.
[0237] In various embodiments, the invention is directed to a method of treating a mammal suffering from or susceptible to a condition associated with abnormal levels of androgen receptor mRNA or protein, comprising administering to the mammal a therapeutically effective amount of an oligomer of the invention, or a conjugate thereof, that comprises one or more LNA monomers.
[0238] An interesting aspect of the invention is directed to the use of an oligomer (compound) as defined herein or a conjugate as defined herein for the preparation of a medicament for the treatment of a condition as disclosed herein above.
[0239] In various embodiments, the invention encompasses a method of preventing or treating a disease comprising administering a therapeutically effective amount of an oligomer according to the invention, or a conjugate thereof, to a human in need of such therapy.
[0240] In certain embodiments, the LNA oligomers of the invention, or conjugates thereof, are administered for a short period time rather than continuously.
[0241] In certain embodiments of the invention, the oligomer (compound) is linked to a conjugated moiety, for example, in order to increase the cellular uptake of the oligomer. In one embodiment the conjugated moiety is a sterol, such as cholesterol.
[0242] In various embodiments, the invention is directed to a method for treating abnormal levels of androgen receptor, the method comprising administering an oligomer of the invention, or a conjugate or a pharmaceutical composition thereof, to a patient in need of such treatment, and further comprising the administration of a further chemotherapeutic agent. In some embodiments, the chemotherapeutic agent is conjugated to the oligomer, is present in the pharmaceutical composition, or is administered in a separate formulation.
[0243] The invention also relates to an oligomer, a composition or a conjugate as defined herein for use as a medicament.
[0244] The invention further relates to use of a compound, composition, or a conjugate as defined herein for the manufacture of a medicament for the treatment of abnormal levels of androgen receptor or expression of mutant forms of AR (such as allelic variants, such as those associated with one of the diseases referred to herein).
[0245] Moreover, in various embodiments, the invention relates to a method of treating a subject suffering from a disease or condition selected from cancer, such as breast cancer or prostate cancer, alopecia, benign prostatic hyperplasia, spinal and muscular atrophy, Kennedy disease and polyglutamate disease, the method comprising the step of administering a pharmaceutical composition as defined herein to the subject in need thereof.
[0246] Suitable dosages, formulations, administration routes, compositions, dosage forms, combinations with other therapeutic agents, pro-drug formulations are also provided in PCT/DK2006/000512--which is hereby incorporated by reference.
[0247] The invention also provides for a pharmaceutical composition comprising a compound or a conjugate as herein described or a conjugate, and a pharmaceutically acceptable diluent, carrier or adjuvant. PCT/DK2006/000512 provides suitable and preferred pharmaceutically acceptable diluents, carriers and adjuvants--which are hereby incorporated by reference.
Embodiments
[0248] The following embodiments of the invention may be used in combination with the other embodiments described herein.
[0249] 1. An oligomer of between 10-50 nucleobases in length which comprises a contiguous nucleobase sequence of a total of between 10-50 nucleobases, wherein said contiguous nucleobase sequence is at least 80% homologous to a corresponding region of a nucleic acid which encodes a mammalian androgen receptor.
[0250] 2. The oligomer according to embodiment 1, wherein said oligomer comprises at least one LNA unit.
[0251] 3. The oligomer according to embodiment 1 or 2, wherein the contiguous nucleobase sequence comprises no more than 3, such as no more than 2 mismatches to the corresponding region of a nucleic acid which encodes a mammalian androgen receptor.
[0252] 4. The oligomer according to embodiment 3, wherein said contiguous nucleobase sequence comprises no more than a single mismatch to the corresponding region of a nucleic acid which encodes a mammalian androgen receptor.
[0253] 5. The oligomer according to embodiment 4, wherein said contiguous nucleobase sequence comprises no mismatches, (i.e. is complementary to) the corresponding region of a nucleic acid which encodes a mammalian androgen receptor.
[0254] 6. The oligomer according to any one of embodiments 1-5, wherein the nucleobase sequence of the oligomer consists of the contiguous nucleobase sequence.
[0255] 7. The oligomer according to any one of embodiments 1-6, wherein the nucleic acid which encodes a mammalian androgen receptor is the human androgen receptor nucleotide sequence such as SEQ ID No 1, or a naturally occurring allelic variant thereof.
[0256] 8. The oligomer according to any one of embodiments 1-7, wherein the contiguous nucleobase sequence is complementary to a corresponding region of both the human androgen receptor nucleic acid sequence and a non-human mammalian androgen receptor nucleic acid sequence, such as the mouse androgen receptor nucleic acid sequence.
[0257] 9. The oligomer according to any one of embodiments 1 to 8, wherein the contiguous nucleobase sequence comprises a contiguous subsequence of at least 7, nucleobase residues which, when formed in a duplex with the complementary androgen receptor target RNA is capable of recruiting RNaseH.
[0258] 10. The oligomer according to embodiment 9, wherein the contiguous nucleobase sequence comprises of a contiguous subsequence of at least 8, at least 9 or at least 10 nucleobase residues which, when formed in a duplex with the complementary androgen receptor target RNA is capable of recruiting RNaseH.
[0259] 11. The oligomer according to any one of embodiments 9 or 10 wherein said contiguous subsequence is at least 9 or at least 10 nucleobases in length, such as at least 12 nucleobases or at least 14 nucleobases in length, such as 14, 15 or 16 nucleobases residues which, when formed in a duplex with the complementary androgen receptor target RNA is capable of recruiting RNaseH.
[0260] 12. The oligomer according to embodiment any one of embodiments 1-11 wherein said oligomer is conjugated with one or more non-nucleobase compounds.
[0261] 13. The oligomer according to any one of embodiments 1-12, wherein said oligomer has a length of between 10-22 nucleobases.
[0262] 14. The oligomer according to any one of embodiments 1-13, wherein said oligomer has a length of between 12-18 nucleobases.
[0263] 15. The oligomer according to any one of embodiments 1-14, wherein said oligomer has a length of 14, 15 or 16 nucleobases.
[0264] 16. The oligomer according to any one of embodiments 1-15, wherein said continuous nucleobase sequence corresponds to a contiguous nucleotide sequence present in a nucleic acid sequence selected from the group consisting of SEQ ID NO 86-106.
[0265] 17. The oligomer according to any one of embodiments 1-16, wherein the oligomer or contiguous nucleobase sequence comprises, or is selected from a corresponding nucleobase sequence present in a nucleotide sequence selected from the group consisting of SEQ ID NO 2-22.
[0266] 18. The oligomer according to any one of embodiments 1-17, wherein said contiguous nucleobase sequence comprises at least one affinity enhancing nucleotide analogue.
[0267] 19. The oligomer according to embodiment 18, wherein said contiguous nucleobase sequence comprises a total of 2, 3, 4, 5, 6, 7, 8, 9 or 10 affinity enhancing nucleotide analogues, such as between 5 and 8 affinity enhancing nucleotide analogues.
[0268] 20. The oligomer according to any one of embodiments 1-19 which comprises at least one affinity enhancing nucleotide analogue, wherein the remaining nucleobases are selected from the group consisting of DNA nucleotides and RNA nucleotides, preferably DNA nucleotides.
[0269] 21. The oligomer according to any one of embodiments 1-20, wherein the oligomer comprises of a sequence of nucleobases of formula, in 5' to 3' direction, A-B-C, and optionally of formula A-B-C-D, wherein:
[0270] (a) consists or comprises of at least one nucleotide analogue, such as 1, 2, 3, 4, 5 or 6 nucleotide analogues, preferably between 2-5 nucleotide analogues, preferably 2, 3 or 4 nucleotide analogues, most preferably 2, 3 or 4 consecutive nucleotide analogues and;
[0271] (b) consists or comprises at least five consecutive nucleobases which are capable of recruiting RNAseH (when formed in a duplex with a complementary RNA molecule, such as the AR mRNA target), such as DNA nucleobases, such as 5, 6, 7, 8, 9, 10, 11 or 12 consecutive nucleobases which are capable of recruiting RNAseH, or between 6-10, or between 7-9, such as 8 consecutive nucleobases which are capable of recruiting RNAseH, and;
[0272] (c) consists or comprises of at least one nucleotide analogue, such as 1, 2, 3, 4, 5, or 6 nucleotide analogues, preferably between 2-5 nucleotide analogues, such as 2, 3 or 4 nucleotide analogues, most preferably 2, 3 or 4 consecutive nucleotide analogues, and;
[0273] (d) when present, consists or comprises, preferably consists, of one or more DNA nucleotide, such as between 1-3 or 1-2 DNA nucleotides.
[0274] 22. The oligomer according to embodiment 21, wherein region A consists or comprises of 2, 3 or 4 consecutive nucleotide analogues.
[0275] 23. The oligomer according to any one of embodiments 21-22, wherein region B consists or comprises of 7, 8, 9 or 10 consecutive DNA nucleotides or equivalent nucleobases which are capable of recruiting RNAseH when formed in a duplex with a complementary RNA, such as the androgen receptor mRNA target.
[0276] 24. The oligomer according to any one of embodiments 21-23, wherein region C consists or comprises of 2, 3 or 4 consecutive nucleotide analogues.
[0277] 25. The oligomer according to any one of embodiments 21-24, wherein region D consists, where present, of one or two DNA nucleotides.
[0278] 26. The oligomer according to any one of embodiments 21-25, wherein:
[0279] (a) Consists or comprises of 3 contiguous nucleotide analogues;
[0280] (b) Consists or comprises of 7, 8, 9 or 10 contiguous DNA nucleotides or equivalent nucleobases which are capable of recruiting RNAseH when formed in a duplex with a complementary RNA, such as the androgen receptor mRNA target;
[0281] (c) Consists or comprises of 3 contiguous nucleotide analogues;
[0282] (d) Consists, where present, of one or two DNA nucleotides.
[0283] 27. The oligomer according to embodiment 26, wherein the contiguous nucleobase sequence consists of 10, 11, 12, 13 or 14 nucleobases, and wherein;
[0284] (a) Consists of 1, 2 or 3 contiguous nucleotide analogues;
[0285] (b) Consists of 7, 8, or 9 consecutive DNA nucleotides or equivalent nucleobases which are capable of recruiting RNAseH when formed in a duplex with a complementary RNA, such as the androgen receptor mRNA target;
[0286] (c) Consists of 1, 2 or 3 contiguous nucleotide analogues;
[0287] (d) Consists, where present, of one DNA nucleotide.
[0288] 28. The oligomer according to anyone of embodiments 21-27, wherein B comprises at least one LNA nucleobase which is in the alpha-L configuration, such as alpha-L-oxy LNA.
[0289] 29. The oligomer according to any one of embodiments 1-28, wherein the nucleotide analogue(s) are independently or collectively selected from the group consisting of: Locked Nucleic Acid (LNA) units; 2'-O-alkyl-RNA units, 2'-OMe-RNA units, 2'-amino-DNA units, 2'-fluoro-DNA units, PNA units, HNA units, and INA units.
[0290] 30. The oligomer according to embodiment 29 wherein all the nucleotide analogues(s) are LNA units.
[0291] 31. The oligomer according to any one of embodiments 1-30, which comprises 1, 2, 3, 4, 5, 6, 7. 8. 9 or 10 LNA units such as between 2 and 8 nucleotide LNA units.
[0292] 32. The oligomer according to any one of the embodiments 29-31, wherein the LNAs are independently selected from oxy-LNA, thio-LNA, and amino-LNA, in either of the beta-D and alpha-L configurations or combinations thereof.
[0293] 33. The oligomer according to embodiment 32, wherein the LNAs are all beta-D-oxy-LNA.
[0294] 34. The oligomer according to any one of embodiments 21-33, wherein the nucleotide analogues or nucleobases of regions A and C are beta-D-oxy-LNA.
[0295] 35. The oligomer according to any one of embodiments 1-34, wherein at least one of the nucleobases present in the oligomer is a modified nucleobase selected from the group consisting of 5-methylcytosine, isocrosine, pseudoisocytosine, 5-bromouracil, 5-propynyluracil, 6-aminopurine, 2-aminopurine, inosine, diaminopurine, and 2-chloro-6-aminopurine.
[0296] 36. The oligomer according to any one of embodiments 1-35, wherein said oligomer hybridises with a corresponding mammalian androgen receptor mRNA with a T.sub.m of at least 50.degree. C.
[0297] 37. The oligomer according to any one of embodiments 1-36, wherein said oligomer hybridises with a corresponding mammalian androgen receptor mRNA with a T.sub.m of no greater than 80.degree. C.
[0298] 38. The oligomer according to any one of embodiments 1-37, wherein the internucleoside linkages are independently selected from the group consisting of: phosphodiester, phosphorothioate and boranophosphate.
[0299] 39. The oligomer according to embodiment 38, wherein the oligomer comprises at least one phosphorothioate internucleoside linkage.
[0300] 40. The oligomer according to embodiment 39, wherein the internucleoside linkages adjacent to or between DNA or RNA units, or within region B are phosphorothioate linkages.
[0301] 41. The oligomer according to embodiment 39 or 40, wherein the linkages between at least one pair of consecutive nucleotide analogues is a phosphodiester linkage.
[0302] 42. The oligomer according to embodiment 39 or 40, wherein all the linkages between consecutive nucleotide analogues are phosphodiester linkages.
[0303] 43. The oligomer according to embodiment 42 wherein all the internucleoside linkages are phosphorothioate linkages.
[0304] 44. A conjugate comprising the oligomer according to any one of the embodiments 1-43 and at least one non-nucleotide or non-polynucleotide moiety covalently attached to said compound.
[0305] 45. A pharmaceutical composition comprising an oligomer as defined in any of embodiments 1-43 or a conjugate as defined in embodiment 44, and a pharmaceutically acceptable diluent, carrier, salt or adjuvant.
[0306] 46. A pharmaceutical composition according to 45, wherein the oligomer is constituted as a pro-drug.
[0307] 47. A pharmaceutical composition according to embodiment 45 or 46, which further comprises a further therapeutic agent selected from the group consisting of: Non-steroidal Antiandrogens and Luteinizing hormone-releasing hormone analogues.
[0308] 48. Use of an oligomer as defined in any one of the embodiments 1-43, or a conjugate as defined in embodiment 44, for the manufacture of a medicament for the treatment of a disease or disorder selected from the group consisting of: Cancer such as breast cancer or prostate cancer, alopecia, benign prostatic hyperplasia, spinal and muscular atrophy, Kennedy disease and polyglutamate disease.
[0309] 49. An oligomer as defined in any one of the embodiments 1-43, or a conjugate as defined in embodiment 44, for use in the treatment of a disease or disorder selected from the group consisting of: Cancer such as breast cancer or prostate cancer, alopecia, benign prostatic hyperplasia, spinal and muscular atrophy, Kennedy disease and polyglutamate disease.
[0310] 50. A method for treating a disease or disorder selected from the group consisting of: Cancer such as breast cancer or prostate cancer, alopecia, benign prostatic hyperplasia, spinal and muscular atrophy, Kennedy disease and polyglutamate disease, said method comprising administering an oligomer as defined in one of the embodiments 1-43, or a conjugate as defined in embodiment 44, or a pharmaceutical composition as defined in any one of the embodiments 45-47, to a patient in need thereof.
[0311] 51. A method for treating an cancer such as prostate cancer or breast cancer, said method comprising administering an oligomer as defined in one of the embodiments 1-43, or a conjugate as defined in embodiment 44, or a pharmaceutical composition as defined in any one of the embodiments 45-47, to a patient in need thereof.
[0312] 52. A method of reducing or inhibiting the expression of androgen receptor in a cell or a tissue, the method comprising the step of contacting said cell or tissue with a compound as defined in one of the embodiments 1-43, or a conjugate as defined in embodiment 44, or a pharmaceutical composition as defined in any one of the embodiments 45-47, so that expression of androgen receptor is reduce or inhibited.
[0313] A method for modulating the expression of a gene which is regulated by the androgen receptor (i.e. an androgen receptor target) in a cell which is expressing said gene, said method comprising the step of contacting said cell or tissue with a compound as defined in one of the embodiments 1-43, or a conjugate as defined in embodiment 44, or a pharmaceutical composition as defined in any one of the embodiments 45-47, so that expression of androgen receptor target is modulated.
EXAMPLES
Example 1
Monomer Synthesis
[0314] The LNA monomer building blocks and derivatives were prepared following published procedures and references cited therein--see WO07/031081 and the references cited therein.
Example 2
Oligonucleotide Synthesis
[0315] Oligonucleotides were synthesized according to the method described in WO07/031081. Table 1 shows examples of sequences of antisense oligonucleotides of the invention. Tables 2 and 3 show examples of antisense oligonucleotides (oligomers) of the invention.
Example 3
Design of the Oligonucleotides
[0316] In accordance with the invention, a series of oligomers were designed to target different regions of human androgen receptor mRNA (GenBank Accession number NM.sub.--000044; SEQ ID NO: 1).
[0317] SEQ ID NOS: 2-22, shown in Table 1, below, are sequences of oligomers designed to target human androgen receptor mRNA. The target region of the target nucleic acid is indicated in the table.
TABLE-US-00002 TABLE 1 Antisense Oligonucleotide Sequences Length Target site SEQ ID NO Sequence (5'-3') (bases) NM_000044 SEQ ID NO: 2 GAGAACCATCCTCACC 16 1389-1404 SEQ ID NO: 3 GGACCAGGTAGCCTGT 16 1428-1443 SEQ ID NO: 4 CCCCTGGACTCAGATG 16 1881-1896 SEQ ID NO: 5 GCACAAGGAGTGGGAC 16 1954-1969 SEQ ID NO: 6 GCTGTGAAGAGAGTGT 16 2422-2437 SEQ ID NO: 7 TTTGACACAAGTGGGA 16 2663-2678 SEQ ID NO: 8 GTGACACCCAGAAGCT 16 2813-2828 SEQ ID NO: 9 CATCCCTGCTTCATAA 16 2975-2990 SEQ ID NO: 10 ACCAAGTTTCTTCAGC 16 3008-3023 SEQ ID NO: 11 CTTGGCCCACTTGACC 16 3263-3278 SEQ ID NO: 12 TCCTGGAGTTGACATT 16 3384-3399 SEQ ID NO: 13 CACTGGCTGTACATCC 16 3454-3469 SEQ ID NO: 14 CATCCAAACTCTTGAG 16 3490-3505 SEQ ID NO: 15 GCTTTCATGCACAGGA 16 3529-3544 SEQ ID NO: 16 GAAGTTCATCAAAGAA 16 3594-3609 SEQ ID NO: 17 AGTTCCTTGATGTAGT 16 3616-3631 SEQ ID NO: 18 TTGCACAGAGATGATC 16 3809-3824 SEQ ID NO: 19 GATGGGCTTGACTTTC 16 3845-3860 SEQ ID NO: 20 CAGGCAGAAGACATCT 16 3924-3939 SEQ ID NO: 21 CCCAAGGCACTGCAGA 16 3960-3975 SEQ ID NO: 86 TGGGGAGAACCATCCTCACCCTGC 24 1385-1408 SEQ ID NO: 87 TCCAGGACCAGGTAGCCTGTGGGG 24 1424-1447 SEQ ID NO: 88 TGTTCCCCTGGACTCAGATGCTCC 24 1877-1990 SEQ ID NO: 89 TGGGGCACAAGGAGTGGGACGCAC 24 1950-1973 SEQ ID NO: 90 TTCGGCTGTGAAGAGAGTGTGCCA 24 2418-2441 SEQ ID NO: 91 CGCTTTTGACACAAGTGGGACTGG 24 2659-2682 SEQ ID NO: 92 CATAGTGACACCCAGAAGCTTCAT 24 2809-2832 SEQ ID NO: 93 GAGTCATCCCTGCTTCATAACATT 24 2971-2994 SEQ ID NO: 94 GATTACCAAGTTTCTTCAGCTTCC 24 3004-3027 SEQ ID NO: 95 AGGCCTTGGCCCACTTGACCACGT 24 3259-3282 SEQ ID NO: 96 AGCATCCTGGAGTTGACATTGGTG 24 3380-3403 SEQ ID NO: 97 GACACACTGGCTGTACATCCGGGA 24 3450-3473 SEQ ID NO: 98 GAGCCATCCAAACTCTTGAGAGAG 24 3486-3509 SEQ ID NO: 99 CAGTGCTTTCATGCACAGGAATTC 24 3525-3548 SEQ ID NO: 100 ATTCGAAGTTCATCAAAGAATTTT 24 3590-3613 SEQ ID NO: 101 ATCGAGTTCCTTGATGTAGTTCAT 24 3612-3635 SEQ ID NO: 102 GCACTTGCACAGAGATGATCTCTG 24 3805-3828 SEQ ID NO: 103 AATAGATGGGCTTGACTTTCCCAG 24 3841-3864 SEQ ID NO: 104 ATAACAGGCAGAAGACATCTGAAA 24 3920-3943 SEQ ID NO: 105 ATTCCCCAAGGCACTGCAGAGGAG 24 3956-3979 SEQ ID NO: 106 ATGGGCTGACATTCATAGCCTTCA 24 3110-3133
[0318] In SEQ ID NOs: 23-43, shown below in Table 2, upper case, boldface letters indicate nucleotide analogue monomers (e.g., -D-oxy LNA monomers) and subscript "s" represents phosphorothiote linkage groups between the monomers. The absence of a subscript "s" (if any) indicates a phosphodiester linkage group. Lower case letters represent DNA monomers.
TABLE-US-00003 TABLE 2 Oligonucleotide designs SEQ ID NO Sequence (5'-3') SEQ ID NO: 23 SEQ ID NO: 24 SEQ ID NO: 25 SEQ ID NO: 26 SEQ ID NO: 27 SEQ ID NO: 28 SEQ ID NO: 29 SEQ ID NO: 30 SEQ ID NO: 31 SEQ ID NO: 32 SEQ ID NO: 33 SEQ ID NO: 34 SEQ ID NO: 35 SEQ ID NO: 36 SEQ ID NO: 37 SEQ ID NO: 38 SEQ ID NO: 39 SEQ ID NO: 40 SEQ ID NO: 41 SEQ ID NO: 42 SEQ ID NO: 43
Example 4
In Vitro Model: Cell Culture
[0319] The effect of antisense oligonucleotides on target nucleic acid expression can be tested in any of a variety of cell types provided that the target nucleic acid is present at measurable levels. The target can be expressed endogenously or by transient or stable transfection of a nucleic acid encoding said target. The expression level of target nucleic acid can be routinely determined using, for example, Northern blot analysis, Real-Time PCR, Ribonuclease protection assays. The following cell types are provided for illustrative purposes, but other cell types can be routinely used, provided that the target is expressed in the cell type chosen.
[0320] Cells were cultured in the appropriate medium as described below and maintained at 37.degree. C. at 95-98% humidity and 5% CO.sub.2. Cells were routinely passaged 2-3 times weekly.
[0321] A549 The human lung cancer cell line A5439 was cultured in DMEM (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+gentamicin (25 .mu.g/ml).
[0322] MCF7 The human breast cancer cell line MCF7 was cultured in EagleMEM (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+1.times. NEAA+gentamicin (25 .mu.g/ml).
Example 5
In Vitro Model: Treatment with Antisense Oligonucleotide
[0323] The cell lines listed in Example 4 were treated with an oligomer using the cationic liposome formulation LipofectAMINE 2000 (Gibco) as transfection vehicle. Cells were seeded in 6-well cell culture plates (NUNC) and treated when 80-90% confluent. Oligomer concentrations used ranged from 1 nM to 16 nM final concentration. Formulation of oligomer-lipid complexes were carried out essentially as described by the manufacturer using serum-free OptiMEM (Gibco) and a final lipid concentration of 5 .mu.g/mL LipofectAMINE 2000. Cells were incubated at 37.degree. C. for 4 hours and treatment was stopped by removal of oligomer-containing culture medium. Cells were washed and serum-containing media was added. After oligomer treatment, cells were allowed to recover for 20 hours before they were harvested for RNA analysis.
Example 6
In Vitro Model: Extraction of RNA and cDNA Synthesis
Total RNA Isolation and First Strand Synthesis
[0324] Total RNA was extracted from cells transfected as described above and using the Qiagen RNeasy kit (Qiagen cat. no. 74104) according to the manufacturer's instructions. First strand synthesis was performed using Reverse Transcriptase reagents from Ambion according to the manufacturer's instructions.
[0325] For each sample, the volume of 0.5g total RNA was adjusted to 10.8l with RNase free H.sub.2O and mixed with 2l random decamers (50M) and 4l dNTP mix (2.5 mM each dNTP) and heated to 70.degree. C. for 3 min, after which the samples were rapidly cooled on ice. After cooling the samples on ice, 2l 10.times. Buffer RT, 1l MMLV Reverse Transcriptase (100 U/l) and 0.25l RNase inhibitor (10 U/l) were added to each sample, followed by incubation at 42.degree. C. for 60 min, heat inactivation of the enzyme at 95.degree. C. for 10 min and then cooling of the sample to 4.degree. C.
Example 7
In Vitro Model: Analysis of Oligonucleotide Inhibition of Androgen Receptor Expression by Real-Time PCR
[0326] Antisense modulation of androgen receptor expression can be assayed in a variety of ways known in the art. For example, androgen receptor mRNA levels can be quantitated by, e.g., Northern blot analysis, competitive polymerase chain reaction (PCR), or real-time PCR. Real-time quantitative PCR is presently preferred. RNA analysis can be performed on total cellular RNA or mRNA.
[0327] Methods of RNA isolation and RNA analysis such as Northern blot analysis are routine in the art and are taught in, for example, Current Protocols in Molecular Biology, John Wiley and Sons.
[0328] Real-time quantitative (PCR) can be conveniently accomplished using the commercially available Multi-Color Real Time PCR Detection System, available from Applied Biosystems.
[0329] Real-time Quantitative PCR Analysis of Androgen Receptor mRNA Levels
[0330] The amount of human androgen receptor mRNA in the samples was quantified using the human androgen receptor ABI Prism Pre-Developed TaqMan Assay Reagents (Applied Biosystems cat, no. Hs00171172_ml) according to the manufacturer's instructions.
[0331] Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) mRNA quantity was used as an endogenous control for normalizing any variance in sample preparation.
[0332] The amount of human GAPDH mRNA in the samples was quantified using the human GAPDH ABI Prism Pre-Developed TaqMan Assay Reagent (Applied Biosystems cat. no. 4310884E) according to the manufacturer's instructions.
[0333] Real-time Quantitative PCR is a technique well known in the art and is taught in for example Heid et al. Real time quantitative PCR, Genome Research (1996), 6: 986-994.
Real Time PCR
[0334] The cDNA from the first strand synthesis performed as described in Example 6 was diluted 2-20 times, and analyzed by real time quantitative PCR using Taqman 7500 FAST or 7900 FAST from Applied Biosystems. The primers and probe were mixed with 2.times. Taqman Fast Universal PCR master mix (2.times.) (Applied Biosystems Cat. #4364103) and added to 4 .mu.l cDNA to a final volume of 10 .mu.l. Each sample was analysed in duplicate. Standard curves were generated by assaying 2-fold dilutions of a cDNA that had been prepared on material purified from a cell line expressing the RNA of interest. Sterile H.sub.2O was used instead of cDNA for the no-template control. PCR program: 95.degree. C. for 30 seconds, followed by 40 cycles of 95.degree. C., 3 seconds, 60.degree. C., 20-30 seconds. Relative quantities of target mRNA were determined from the calculated Threshold cycle using the Applied Biosystems Fast System SDS Software Version 1.3.1.21. or SDS Software Version 2.3.
Example 8
In Vitro Analysis: Antisense Inhibition of Human Androgen Receptor mRNA Expression by Oligonucleotide Compounds
[0335] Oligonucleotides presented in Table 3 were evaluated for their potential to knock down androgen receptor mRNA expression at concentrations of 1, 4 and 16 nM (see FIGS. 1 and 2).
[0336] The data in Table 3 are presented as percentage down-regulation relative to mock transfected cells at 16 nM. Lower case letters represent DNA monomers, bold, upper case letters represent .beta.-D-oxy-LNA monomers. All cytosine bases in the LNA monomers are 5-methylcytosines. Subscript `a` represents a phosphorothioate linkage.
TABLE-US-00004 TABLE 3 Inhibition of human androgen receptor mRNA expression by oligonucleotides Percent Percent inhibition inhibition of Test of Androgen Androgen substance Sequence (5'-3') receptor-MCF7 receptor-A549 SEQ ID NO: 44 80.1 63.8 SEQ ID NO: 45 89.0 88.2 SEQ ID NO: 46 89.4 82.8 SEQ ID NO: 47 83.1 77.7 SEQ ID NO: 48 93.8 96.7 SEQ ID NO: 49 n.d. n.d. SEQ ID NO: 50 n.d. n.d. SEQ ID NO: 51 96.9 95.5 SEQ ID NO: 52 n.d. n.d. SEQ ID NO: 53 n.d. n.d. SEQ ID NO: 54 95.4 98.3 SEQ ID NO: 55 n.d. n.d. SEQ ID NO: 56 n.d. n.d. SEQ ID NO: 57 89.5 88.9 SEQ ID NO: 58 95.6 98.9 SEQ ID NO: 59 n.d. n.d. SEQ ID NO: 60 n.d. n.d. SEQ ID NO: 61 86.7 93.3 SEQ ID NO: 62 81.3 93.0 SEQ ID NO: 63 90.9 98.4 SEQ ID NO: 64 n.d. n.d. SEQ ID NO: 65 n.d. n.d. SEQ ID NO: 66 79.8 95.3 SEQ ID NO: 67 83.5 97.0 SEQ ID NO: 68 88.2 85.6 SEQ ID NO: 69 92.7 94.0 SEQ ID NO: 70 n.d. n.d. SEQ ID NO: 71 n.d. n.d. SEQ ID NO: 72 79.2 90.4 SEQ ID NO: 73 91.1 97.3 SEQ ID NO: 74 n.d. n.d. SEQ ID NO: 75 n.d. n.d. SEQ ID NO: 76 85.9 94.3 SEQ ID NO: 77 93.0 98.5 SEQ ID NO: 78 n.d. n.d. SEQ ID NO: 79 n.d. n.d. SEQ ID NO: 80 n.d. n.d.
[0337] As shown in Table 3, oligonucleotides having the sequences set forth in SEQ ID NOs: 48, 51, 54, 58, 63, 69, 73 and 77 at 16 nM demonstrated greater than 90% inhibition of androgen receptor mRNA expression in A549 and MCF7 cells in these experiments.
[0338] In certain embodiments, oligomers based on the tested antisense oligomer sequences and designs, but having, for example, different lengths (shorter or longer) and/or monomer content (e.g. the type and/or number of nucleoside analogues) than those shown, e.g., in Table 3, could also provide suitable inhibition of androgen receptor expression.
Example 9
In Vivo Analysis: Antisense Inhibition of Mouse Androgen Receptor mRNA Liver Expression by Oligonucleotide Compounds
[0339] Nude mice were dosed i.v. q3dx4 with 100 mg/kg oligonucleotide (group size of 5 mice). The antisense oligonucleotides (SEQ ID:48, SEQ ID:51, SEQ ID:58, SEQ ID:63, SEQ ID:77) were dissolved in phosphate buffered saline. Animals were sacrificed 24 h after last dosing and liver tissue was sampled and stored in RNA later until RNA extraction and QPCR analysis. Total RNA was extracted and AR mRNA expression in liver samples was measured by QPCR as described in Example 7 using a mouse AR QPCR assay (cat. Mm01238475_ml, Applied Biosystems). Results were normalised to mouse GAPDH (cat. no. 4352339E, Applied Biosystems) and knock-down was quantitated relative to saline treated controls. The data in Table 4 are presented as percentage down-regulation relative to saline treated animals.
TABLE-US-00005 TABLE 4 In vivo knock-down of AR mRNA expression Compound Liver (% KD) Saline 0 SEQ ID: 51 100 mg/kg 65.0 +/- 12.6 SEQ ID: 58 100 mg/kg 95.2 +/- 1.0 SEQ ID: 77 100 mg/kg 91.9 +/- 3.9
[0340] As shown in Table 4, oligonucleotides of SEQ ID NOs: 58 and 77 at 100 mg/kg demonstrated greater than 90% inhibition of androgen receptor mRNA expression in mouse liver cells in these experiments.
Example 10
In Vitro Analysis: Antisense Inhibition of Human Androgen Receptor mRNA
Measurement of Proliferating Viable Cells (MTS Assay)
[0341] LNCaP prostate cancer and A549 lung cancer cells were seeded to a density of 150,000 cells per well in a 6-well plate the day prior to transfection. A549 cells were cultured in DMEM (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+gentamicin (25 .mu.g/ml) whereas LNCaP cells were cultured in RPMI 1640 Medium (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+gentamicin (25 .mu.g/ml). On the following day, medium was removed followed by addition of 1.2 ml OptiMEM containing 5 .mu.g/ml Lipofectamine2000 (Invitrogen). Cells were incubated for 7 min before adding 0.3 ml oligonucleotides diluted in OptiMEM. The final oligonucleotide concentrations were 4 nM and 16 nM. After 4 hours of treatment, media was removed and cells were trypsinized and seeded to a density of 5000 cells per well in a clear 96 well plate (Scientific Orange no. 1472030100) in 100 .mu.l media. Viable cells were measured at the times indicated by adding 10 .mu.l the tetrazolium compound [3-(4,5-dimethyl-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-te- trazolium, inner salt; MTS] and an electron coupling reagent (phenazine ethosulfate; PES) (CellTiter 96.RTM. AQueous One Solution Cell Proliferation Assay, Promega). Viable cells were measured at 490 nm in a Powerwave (Biotek Instruments). The OD490 nm measurements were plotted against time/h. (See FIG. 13 and FIG. 14). As shown in FIG. 13 and FIG. 14, oligonucleotides of SEQ ID NOs: 58 and 77 inhibit growth of both LNCaP prostate and A549 lung cancer cells.
Example 11
In Vitro Analysis: Caspase 3/7 Activity by Antisense Inhibition of Human Androgen Receptor mRNA
[0342] LNCaP prostate cancer cells and A549 lung cancer cells were seeded to a density of 150,000 cells per well in a 6-well plate the day prior to transfection. A549 cells were cultured in DMEM (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+gentamicin (25 .mu.g/ml) whereas LNCaP cells were cultured in RPMI 1640 Medium (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+gentamicin (25 .mu.g/ml). The next day medium was removed followed by addition of 1.2 ml OptiMEM containing 5 .mu.g/ml Lipofectamine2000 (Invitrogen). Cells were incubated for 7 min before adding 0.3 ml oligonucleotides diluted in OptiMEM. The final oligonucleotide concentrations were 4 nM and 16 nM. After 4 hours of treatment, media was removed and cells were trypsinized and seeded to a density of 5000 cells per well in a white 96 well plate (Nunc) in 100 .mu.l media. Caspase 3/7 activity was measured at the times indicated by adding 100 .mu.l Caspase-Glo 3/7 assay (Promega). Caspase 3/7 activity was measured using a luminometer. The caspase 3/7 activities were measured at three different time points 14 h, 48 h and 72 h (See FIG. 15 and FIG. 16). As shown in FIG. 15 and FIG. 16, oligonucleotides of SEQ ID NOs: 58 and 77 induce caspase 3/7 activity in both LNCaP prostate and A549 lung cancer cells.
Example 12
In Vitro Analysis: Antisense Inhibition of Human Androgen Receptor mRNA Expression by Oligonucleotide Compounds in Prostate Cancer Cell Line LNCaP and Lung Cancer Cell Line A549
[0343] Oligonucleotides were evaluated for their potential to knock down androgen receptor mRNA expression at concentrations of 0.5, 1, 2, 4, 8 and 16 nM (see FIG. 11). LNCaP prostate cancer cells and A549 lung cancer cells were seeded to a density of 150,000 cells per well in a 6-well plate the day prior to transfection. A549 cells were cultured DMEM (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+gentamicin (25 .mu.g/ml). LNCaP cells were cultured in RPMI 1640 Medium (Sigma)+10% fetal bovine serum (FBS)+2 mM Glutamax I+gentamicin (25 .mu.g/ml). On the following day, medium was removed followed by addition of 1.2 ml OptiMEM containing 5 .mu.g/ml Lipofectamine2000 (Invitrogen). Cells were incubated for 7 min before adding 0.3 ml oligonucleotides diluted in OptiMEM. The final oligonucleotide concentrations were 0.5, 1, 2, 4, 8 and 16 nM. Cells were washed and serum-containing media was added. After oligomer treatment cells were allowed to recover for 20 hours before they were harvested for RNA analysis. The procedure for RNA isolation, cDNA synthesis and qPCR were as described in Examples 5, 6 and 7. As shown in FIGS. 11 and 12 oligonucleotides of SEQ ID NOs: 58 and 77 were potent in knocking down AR mRNA expression in both the lung cancer cell line A549 and in the androgen receptor-dependent LNCaP prostate cancer cell line.
Example 13
In Vivo Analysis: Effect of Antisense Oligonucleotides on PSA Levels and Androgen-Dependent Prostate Tumor Growth in Mice
[0344] Six to seven week old male athymic mu/mu mice (Harlan Sprague Dawley) weighing an average of 27.3.+-.2.4 g were used in the study. Ten million cells of 22RV1 (androgen-independent prostate cancer line) were suspended in PBS (Gibco#14190) and Matrigel (BD#356234) with a ratio of 1:1 were injected subcutaneously into each mouse. When tumors reached an average volume of 150-200 mm.sup.3, the mice were divided into nine experimental groups. Two hundred .mu.l of oligomer were injected intravenously when the average tumor size reached 152.66.+-.27.97 mm.sup.3. Oligomers were given every 3 days for a total of 5 dosings. The control vehicles were given using the same dosing regimen as the oligomers. On day 16, mice were sacrificed and blood collected in EDTA laced tubes and spun for 5 min. 50 .mu.l of the supernatants were then subjected to PSA assay using the ELISA kit from ALPCO Diagnostics in Salem (PSAHU-L01). Results of the experiment are shown in FIG. 17.
[0345] Six to seven week old male athymic mu/mu mice (Harlan Sprague Dawley) weighing an average of 27.3.+-.2.4 g were used in the study. Ten million cells of 22RV1 (androgen-independent prostate cancer line) were suspended in PBS (Gibco#14190) and Matrigel (BD#356234) with a ratio of 1:1 were injected subcutaneously into each mouse. When tumors reached an average volume of 150-200 mm.sup.3, the mice were divided into nine experimental groups. Two hundred .mu.l of oligomer was injected intravenously when the average tumor size reached 152.66.+-.27.97 mm.sup.3. Oligomers were given every 3 days for a total of 5 dosings. The control vehicles were given using the same dosing regimen as the oligomers. The tumor volumes for each mouse were determined by measuring two dimensions with calipers and calculated using the formula: tumor volume=(length.times.width.sup.2)/2). Results of the experiment are shown in FIG. 18.
Example 14
Preparation of Conjugates of Oligomers with Polyethylene Glycol
[0346] The oligomers having sequences shown as SEQ ID NO: 48 or SEQ ID NO: 63 are functionalized on the 5' terminus by attaching an aminoalkyl group, such as hexan-1-amine blocked with a blocking group such as Fmoc to the 5' phosphate groups of the oligomers using routine phosphoramidite chemistry, oxidizing the resultant compounds, deprotecting them and purifying them to achieve the functionalized oligomers, respectively, having the formulas (IA) and (IB):
##STR00007##
wherein the bold uppercase letters represent nucleoside analogue monomers, lowercase letters represent DNA monomers, the subscript "s" represents a phosphorothioate linkage, and .sup.MeC represents 5-methylcytosine.
[0347] A solution of activated PEG, such as the one shown in formula (II):
##STR00008##
wherein the PEG moiety has an average molecular weight of 12,000, and each of the compounds of formulas (IA) and (IB) in PBS buffer are stirred in separate vessels at room temperature for 12 hours. The reaction solutions are extracted three times with methylene chloride and the combined organic layers are dried over magnesium sulphate and filtered and the solvent is evaporated under reduced pressure. The resulting residues are dissolved in double distilled water and loaded onto an anion exchange column. Unreacted PEG linker is eluted with water and the products are eluted with NH.sub.4HCO.sub.3 solution. Fractions containing pure products are pooled and lypophilized to yield the conjugates SEQ ID NOs: 48 and 63, respectively as show in formulas (IIIA) and (IIIB):
##STR00009##
wherein each of the oligomers of SEQ ID NOs: 48 and 63 is attached to a PEG polymer having average molecular weight of 12,000 via a releasable linker.
[0348] Chemical structures of PEG polymer conjugates that can be made with oligomers having sequences shown in SEQ ID NOs: 51, 58 and 77 using the process described above are respectively shown in formulas (IVA), (IVB) and (IVC):
##STR00010##
wherein bold uppercase letters represent beta-D-oxy-LNA monomers, lowercase letters represent DNA monomers, the subscript "s" represents a phosphorothioate linkage and .sup.MeC represent 5-methylcytosine.
[0349] Activated oligomers that can be used in this process to respectively make the conjugates shown in formulas (IVA), (IVB) and (IVC) have the chemical structures shown in formulas (VA), (VB) and (VC):
##STR00011##
Sequence CWU 1
1
10614314DNAhomo sapiens 1cgagatcccg gggagccagc ttgctgggag
agcgggacgg tccggagcaa gcccagaggc 60agaggaggcg acagagggaa aaagggccga
gctagccgct ccagtgctgt acaggagccg 120aagggacgca ccacgccagc
cccagcccgg ctccagcgac agccaacgcc tcttgcagcg 180cggcggcttc
gaagccgccg cccggagctg ccctttcctc ttcggtgaag tttttaaaag
240ctgctaaaga ctcggaggaa gcaaggaaag tgcctggtag gactgacggc
tgcctttgtc 300ctcctcctct ccaccccgcc tccccccacc ctgccttccc
cccctccccc gtcttctctc 360ccgcagctgc ctcagtcggc tactctcagc
caacccccct caccaccctt ctccccaccc 420gcccccccgc ccccgtcggc
ccagcgctgc cagcccgagt ttgcagagag gtaactccct 480ttggctgcga
gcgggcgagc tagctgcaca ttgcaaagaa ggctcttagg agccaggcga
540ctggggagcg gcttcagcac tgcagccacg acccgcctgg ttaggctgca
cgcggagaga 600accctctgtt ttcccccact ctctctccac ctcctcctgc
cttccccacc ccgagtgcgg 660agccagagat caaaagatga aaaggcagtc
aggtcttcag tagccaaaaa acaaaacaaa 720caaaaacaaa aaagccgaaa
taaaagaaaa agataataac tcagttctta tttgcaccta 780cttcagtgga
cactgaattt ggaaggtgga ggattttgtt tttttctttt aagatctggg
840catcttttga atctaccctt caagtattaa gagacagact gtgagcctag
cagggcagat 900cttgtccacc gtgtgtcttc ttctgcacga gactttgagg
ctgtcagagc gctttttgcg 960tggttgctcc cgcaagtttc cttctctgga
gcttcccgca ggtgggcagc tagctgcagc 1020gactaccgca tcatcacagc
ctgttgaact cttctgagca agagaagggg aggcggggta 1080agggaagtag
gtggaagatt cagccaagct caaggatgga agtgcagtta gggctgggaa
1140gggtctaccc tcggccgccg tccaagacct accgaggagc tttccagaat
ctgttccaga 1200gcgtgcgcga agtgatccag aacccgggcc ccaggcaccc
agaggccgcg agcgcagcac 1260ctcccggcgc cagtttgctg ctgctgcagc
agcagcagca gcagcagcag cagcagcagc 1320agcagcagca gcagcagcag
cagcagcagc agcaagagac tagccccagg cagcagcagc 1380agcagcaggg
tgaggatggt tctccccaag cccatcgtag aggccccaca ggctacctgg
1440tcctggatga ggaacagcaa ccttcacagc cgcagtcggc cctggagtgc
caccccgaga 1500gaggttgcgt cccagagcct ggagccgccg tggccgccag
caaggggctg ccgcagcagc 1560tgccagcacc tccggacgag gatgactcag
ctgccccatc cacgttgtcc ctgctgggcc 1620ccactttccc cggcttaagc
agctgctccg ctgaccttaa agacatcctg agcgaggcca 1680gcaccatgca
actccttcag caacagcagc aggaagcagt atccgaaggc agcagcagcg
1740ggagagcgag ggaggcctcg ggggctccca cttcctccaa ggacaattac
ttagggggca 1800cttcgaccat ttctgacaac gccaaggagt tgtgtaaggc
agtgtcggtg tccatgggcc 1860tgggtgtgga ggcgttggag catctgagtc
caggggaaca gcttcggggg gattgcatgt 1920acgccccact tttgggagtt
ccacccgctg tgcgtcccac tccttgtgcc ccattggccg 1980aatgcaaagg
ttctctgcta gacgacagcg caggcaagag cactgaagat actgctgagt
2040attccccttt caagggaggt tacaccaaag ggctagaagg cgagagccta
ggctgctctg 2100gcagcgctgc agcagggagc tccgggacac ttgaactgcc
gtctaccctg tctctctaca 2160agtccggagc actggacgag gcagctgcgt
accagagtcg cgactactac aactttccac 2220tggctctggc cggaccgccg
ccccctccgc cgcctcccca tccccacgct cgcatcaagc 2280tggagaaccc
gctggactac ggcagcgcct gggcggctgc ggcggcgcag tgccgctatg
2340gggacctggc gagcctgcat ggcgcgggtg cagcgggacc cggttctggg
tcaccctcag 2400ccgccgcttc ctcatcctgg cacactctct tcacagccga
agaaggccag ttgtatggac 2460cgtgtggtgg tggtgggggt ggtggcggcg
gcggcggcgg cggcggcggc ggcggcggcg 2520gcggcggcgg cggcgaggcg
ggagctgtag ccccctacgg ctacactcgg ccccctcagg 2580ggctggcggg
ccaggaaagc gacttcaccg cacctgatgt gtggtaccct ggcggcatgg
2640tgagcagagt gccctatccc agtcccactt gtgtcaaaag cgaaatgggc
ccctggatgg 2700atagctactc cggaccttac ggggacatgc gtttggagac
tgccagggac catgttttgc 2760ccattgacta ttactttcca ccccagaaga
cctgcctgat ctgtggagat gaagcttctg 2820ggtgtcacta tggagctctc
acatgtggaa gctgcaaggt cttcttcaaa agagccgctg 2880aagggaaaca
gaagtacctg tgcgccagca gaaatgattg cactattgat aaattccgaa
2940ggaaaaattg tccatcttgt cgtcttcgga aatgttatga agcagggatg
actctgggag 3000cccggaagct gaagaaactt ggtaatctga aactacagga
ggaaggagag gcttccagca 3060ccaccagccc cactgaggag acaacccaga
agctgacagt gtcacacatt gaaggctatg 3120aatgtcagcc catctttctg
aatgtcctgg aagccattga gccaggtgta gtgtgtgctg 3180gacacgacaa
caaccagccc gactcctttg cagccttgct ctctagcctc aatgaactgg
3240gagagagaca gcttgtacac gtggtcaagt gggccaaggc cttgcctggc
ttccgcaact 3300tacacgtgga cgaccagatg gctgtcattc agtactcctg
gatggggctc atggtgtttg 3360ccatgggctg gcgatccttc accaatgtca
actccaggat gctctacttc gcccctgatc 3420tggttttcaa tgagtaccgc
atgcacaagt cccggatgta cagccagtgt gtccgaatga 3480ggcacctctc
tcaagagttt ggatggctcc aaatcacccc ccaggaattc ctgtgcatga
3540aagcactgct actcttcagc attattccag tggatgggct gaaaaatcaa
aaattctttg 3600atgaacttcg aatgaactac atcaaggaac tcgatcgtat
cattgcatgc aaaagaaaaa 3660atcccacatc ctgctcaaga cgcttctacc
agctcaccaa gctcctggac tccgtgcagc 3720ctattgcgag agagctgcat
cagttcactt ttgacctgct aatcaagtca cacatggtga 3780gcgtggactt
tccggaaatg atggcagaga tcatctctgt gcaagtgccc aagatccttt
3840ctgggaaagt caagcccatc tatttccaca cccagtgaag cattggaaac
cctatttccc 3900caccccagct catgccccct ttcagatgtc ttctgcctgt
tataactctg cactactcct 3960ctgcagtgcc ttggggaatt tcctctattg
atgtacagtc tgtcatgaac atgttcctga 4020attctatttg ctgggctttt
tttttctctt tctctccttt ctttttcttc ttccctccct 4080atctaaccct
cccatggcac cttcagactt tgcttcccat tgtggctcct atctgtgttt
4140tgaatggtgt tgtatgcctt taaatctgtg atgatcctca tatggcccag
tgtcaagttg 4200tgcttgttta cagcactact ctgtgccagc cacacaaacg
tttacttatc ttatgccacg 4260ggaagtttag agagctaaga ttatctgggg
aaatcaaaac aaaaacaagc aaac 4314216DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 2gagaaccatc ctcacc
16316DNAartificialLNA oligomer Sequence/oligomer Sequence motif
3ggaccaggta gcctgt 16416DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 4cccctggact cagatg 16516DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 5gcacaaggag tgggac
16616DNAartificialLNA oligomer Sequence/oligomer Sequence motif
6gctgtgaaga gagtgt 16716DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 7tttgacacaa gtggga 16816DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 8gtgacaccca gaagct
16916DNAartificialLNA oligomer Sequence/oligomer Sequence motif
9catccctgct tcataa 161016DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 10accaagtttc ttcagc
161116DNAartificialLNA oligomer Sequence/oligomer Sequence motif
11cttggcccac ttgacc 161216DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 12tcctggagtt gacatt
161316DNAartificialLNA oligomer Sequence/oligomer Sequence motif
13cactggctgt acatcc 161416DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 14catccaaact cttgag
161516DNAartificialLNA oligomer Sequence/oligomer Sequence motif
15gctttcatgc acagga 161616DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 16gaagttcatc aaagaa
161716DNAartificialLNA oligomer Sequence/oligomer Sequence motif
17agttccttga tgtagt 161816DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 18ttgcacagag atgatc
161916DNAartificialLNA oligomer Sequence/oligomer Sequence motif
19gatgggcttg actttc 162016DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 20caggcagaag acatct
162116DNAartificialLNA oligomer Sequence/oligomer Sequence motif
21cccaaggcac tgcaga 162216DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 22gctgacattc atagcc
162316DNAartificialLNA oligomer Sequence/oligomer Sequence motif
23gagaaccatc ctcacc 162416DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 24ggaccaggta gcctgt
162516DNAartificialLNA oligomer Sequence/oligomer Sequence motif
25cccctggact cagatg 162616DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 26gcacaaggag tgggac
162716DNAartificialLNA oligomer Sequence/oligomer Sequence motif
27gctgtgaaga gagtgt 162816DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 28tttgacacaa gtggga
162916DNAartificialLNA oligomer Sequence/oligomer Sequence motif
29gtgacaccca gaagct 163016DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 30catccctgct tcataa
163116DNAartificialLNA oligomer Sequence/oligomer Sequence motif
31accaagtttc ttcagc 163216DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 32cttggcccac ttgacc
163316DNAartificialLNA oligomer Sequence/oligomer Sequence motif
33tcctggagtt gacatt 163416DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 34cactggctgt acatcc
163516DNAartificialLNA oligomer Sequence/oligomer Sequence motif
35catccaaact cttgag 163616DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 36gctttcatgc acagga
163716DNAartificialLNA oligomer Sequence/oligomer Sequence motif
37gaagttcatc aaagaa 163816DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 38agttccttga tgtagt
163916DNAartificialLNA oligomer Sequence/oligomer Sequence motif
39ttgcacagag atgatc 164016DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 40gatgggcttg actttc
164116DNAartificialLNA oligomer Sequence/oligomer Sequence motif
41caggcagaag acatct 164216DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 42cccaaggcac tgcaga
164316DNAartificialLNA oligomer Sequence/oligomer Sequence motif
43gctgacattc atagcc 164416DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 44gagaaccatc ctcacc
164516DNAartificialLNA oligomer Sequence/oligomer Sequence motif
45ggaccaggta gcctgt 164616DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 46cccctggact cagatg
164716DNAartificialLNA oligomer Sequence/oligomer Sequence motif
47gcacaaggag tgggac 164816DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 48gctgtgaaga gagtgt
164914DNAartificialLNA oligomer Sequence/oligomer Sequence motif
49ctgtgaagag agtg 145012DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 50tgtgaagaga gt 125116DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 51tttgacacaa gtggga
165214DNAartificialLNA oligomer Sequence/oligomer Sequence motif
52ttgacacaag tggg 145312DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 53tgacacaagt gg 125416DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 54gtgacaccca gaagct
165514DNAartificialLNA oligomer Sequence/oligomer Sequence motif
55tgacacccag aagc 145612DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 56gacacccaga ag 125716DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 57catccctgct tcataa
165816DNAartificialLNA oligomer Sequence/oligomer Sequence motif
58accaagtttc ttcagc 165914DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 59ccaagtttct tcag
146012DNAartificialLNA oligomer Sequence/oligomer Sequence motif
60caagtttctt ca 126116DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 61cttggcccac ttgacc 166216DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 62tcctggagtt gacatt
166316DNAartificialLNA oligomer Sequence/oligomer Sequence motif
63cactggctgt acatcc 166414DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 64actggctgta catc
146512DNAartificialLNA oligomer Sequence/oligomer Sequence motif
65ctggctgtac at 126616DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 66catccaaact cttgag 166716DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 67gctttcatgc acagga
166816DNAartificialLNA oligomer Sequence/oligomer Sequence motif
68gaagttcatc aaagaa 166916DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 69agttccttga tgtagt
167014DNAartificialLNA oligomer Sequence/oligomer Sequence motif
70gttccttgat gtag 147112DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 71ttccttgatg ta 127216DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 72ttgcacagag atgatc
167316DNAartificialLNA oligomer Sequence/oligomer Sequence motif
73gatgggcttg actttc 167414DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 74atgggcttga cttt
147512DNAartificialLNA oligomer Sequence/oligomer Sequence motif
75tgggcttgac tt 127616DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 76caggcagaag acatct 167716DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 77cccaaggcac tgcaga
167814DNAartificialLNA oligomer Sequence/oligomer Sequence motif
78ccaaggcact gcag 147912DNAartificialLNA oligomer Sequence/oligomer
Sequence motif 79caaggcactg ca 128016DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 80gctgacattc atagcc 16812999DNAMus
musculus 81gaattcggtg gaagctacag acaagctcaa ggatggaggt gcagttaggg
ctgggaaggg 60tctacccacg gcccccatcc aagacctatc gaggagcgtt ccagaatctg
ttccagagcg 120tgcgcgaagc gatccagaac ccgggcccca ggcaccctga
ggccgctaac atagcacctc 180ccggcgcctg tttacagcag aggcaggaga
ctagcccccg gcggcggcgg cggcagcagc 240acactgagga tggttctcct
caagcccaca tcagaggccc cacaggctac ctggccctgg 300aggaggaaca
gcagccttca cagcagcagg cagcctccga gggccaccct gagagcagct
360gcctccccga gcctggggcg gccaccgctc ctggcaaggg gctgccgcag
cagccaccag 420ctcctccaga tcaggatgac tcagctgccc catccacgtt
gtccctgctg ggccccactt 480tcccaggctt aagcagctgc tccgccgaca
ttaaagacat tttgaacgag gccggcacca 540tgcaacttct tcagcagcag
caacaacagc agcagcacca acagcagcac caacagcacc 600aacagcagca
ggaggtaatc tccgaaggca gcagcgcaag agccagggag gccacggggg
660ctccctcttc ctccaaggat agttacctag ggggcaattc aaccatatct
gacagtgcca 720aggagttgtg taaagcagtg tctgtgtcca tgggattggg
tgtggaagca ttggaacatc 780tgagtccagg ggaacagctt cggggagact
gcatgtacgc gtcgctcctg ggaggtccac 840ccgcggtgcg tcccactcct
tgtgcgccgc tgcccgaatg caaaggtctt cccctggacg 900aaggcccagg
caaaagcact gaagagactg ctgagtattc ctctttcaag ggaggttacg
960ccaaaggatt ggaaggtgag agcttggggt gctctggcag cagtgaagca
ggtagctctg 1020ggacacttga gatcccgtcc tctctgtctc tgtataaatc
tggagcacta gacgaggcag 1080cagcatacca gaatcgcgac tactacaact
ttccgctggc tctgtccggg ccgccgcacc 1140ccccgccccc tacccatcca
cacgcccgta tcaagctgga gaacccattg gactacggca 1200gcgcctgggc
tgcggcggca gcgcaatgcc gctatgggga cttgggtagt ctacatggag
1260ggagtgtagc cgggcccagc actggatcgc ccccagccac cacctcttct
tcctggcata 1320ctctcttcac agctgaagaa ggccaattat atgggccagg
aggcgggggc ggcagcagca 1380gcccaagcga tgccgggcct gtagccccct
atggctacac tcggccccct caggggctga 1440caagccagga gagtgactac
tctgcctccg aagtgtggta tcctggtgga gttgtgaaca 1500gagtacccta
tcccagtccc aattgtgtca aaagtgaaat gggaccttgg atggagaact
1560actccggacc ttatggggac atgcgtttgg acagtaccag ggaccatgtt
ttacccatcg 1620actattactt tccaccccag aagacctgcc tgatctgtgg
agatgaagct tctggctgtc 1680actacggagc tctcacttgt ggcagctgca
aggtcttctt caaaagagcc gctgaaggga 1740aacagaagta tctatgtgcc
agcagaaacg attgtaccat tgataaattt cggaggaaaa 1800attgcccatc
ttgtcgtctc cggaaatgtt atgaagcagg gatgactctg ggagctcgta
1860agctgaagaa acttggaaat ctaaaactac aggaggaagg agaaaactcc
aatgctggca 1920gccccactga ggacccatcc cagaagatga ctgtatcaca
cattgaaggc tatgaatgtc 1980agcctatctt tcttaacgtc ctggaagcca
ttgagccagg agtggtgtgt gccggacatg 2040acaacaacca accagattcc
tttgctgcct tgttatctag cctcaatgag cttggagaga 2100ggcagcttgt
gcatgtggtc aagtgggcca aggccttgcc tggcttccgc aacttgcatg
2160tggatgacca gatggcggtc attcagtatt cctggatggg actgatggta
tttgccatgg 2220gttggcggtc cttcactaat gtcaactcca ggatgctcta
ctttgcacct gacttggttt 2280tcaatgagta ccgcatgcac aagtctcgga
tgtacagcca gtgtgtgagg atgaggcacc 2340tgtctcaaga gtttggatgg
ctccaaataa ccccccagga attcctgtgc atgaaagcac 2400tgctgctctt
cagcattatt ccagtggatg ggctgaaaaa tcaaaaattc tttgatgaac
2460ttcgaatgaa ctacatcaag gaactcgatc gcatcattgc atgcaaaaga
aagaatccca 2520catcctgctc aaggcgcttc taccagctca ccaagctcct
ggattctgtg cagcctattg 2580caagagagct gcatcagttc acttttgacc
tgctaatcaa gtcccatatg gtgagcgtgg 2640actttcctga aatgatggca
gagatcatct ctgtgcaagt gcccaagatc ctttctggga 2700aagtcaagcc
catctatttc cacacacagt gaagatttgg aaaccctaat acccaaaacc
2760caccttgttc cctttccaga tgtcttctgc ctgttatata actctgcact
acttctctgc 2820agtgccttgg gggaaattcc tctactgatg tacagtctgt
cgtgaacagg ttcctcagtt 2880ctatttcctg ggcttctcct tctttttttt
tcttcttccc tccctctttc accctcccat 2940ggcacatttt gaatctgctg
cgtattgtgg ctcctgcctt tgttttgatt tctgttgta 2999823175DNAMacaca
mulatta 82cccaaaaaat aaaaacaaac aaaaacaaaa caaaacaaaa aaaacgaata
aagaaaaagg 60taataactca gttcttattt gcacctactt ccagtggaca ctgaatttgg
aaggtggagg 120attcttgttt tttcttttaa gatcgggcat cttttgaatc
tacccctcaa gtgttaagag 180acagactgtg agcctagcag ggcagatctt
gtccaccgtg tgtcttcttt tgcaggagac 240tttgaggctg tcagagcgct
ttttgcgtgg ttgctcccgc aagtttcctt ctctggagct 300tcccgcaggt
gggcagctag ctgcagcgac taccgcatca tcacagcctg ttgaactctt
360ctgagcaaga gaaggggagg cggggtaagg gaagtaggtg gaagattcag
ccaagctcaa 420ggatggaggt gcagttaggg ctggggaggg tctaccctcg
gccgccgtcc aagacctacc 480gaggagcttt ccagaatctg ttccagagcg
tgcgcgaagt gatccagaac ccgggcccca 540ggcacccaga ggccgcgagc
gcagcacctc ccggcgccag tttgcagcag cagcagcagc 600agcagcaaga
aactagcccc cggcaacagc agcagcagca gcagggtgag gatggttctc
660cccaagccca tcgtagaggc cccacaggct acctggtcct ggatgaggaa
cagcagcctt 720cacagcctca gtcagccccg gagtgccacc ccgagagagg
ttgcgtccca gagcctggag 780ccgccgtggc cgccggcaag gggctgccgc
agcagctgcc agcacctccg gacgaggatg 840actcagctgc cccatccacg
ttgtctctgc tgggccccac tttccccggc ttaagcagct 900gctccgccga
ccttaaagac atcctgagcg aggccagcac catgcaactc cttcagcaac
960agcagcagga agcagtatcc gaaggcagca gcagcgggag agcgagggag
gcctcggggg 1020ctcccacttc ctccaaggac aattacttag agggcacttc
gaccatttct gacagcgcca 1080aggagctgtg taaggcagtg tcggtgtcca
tgggcttggg tgtggaggcg ttggagcatc 1140tgagtccagg ggaacagctt
cggggggatt gcatgtacgc cccagttttg ggagttccac 1200ccgctgtgcg
tcccactccg tgtgccccat tggccgaatg caaaggttct ctgctagacg
1260acagcgcagg caagagcact gaagatactg ctgagtattc ccctttcaag
ggaggttaca 1320ccaaagggct agaaggcgag agcctaggct gctctggcag
cgctgcagca gggagctccg 1380ggacacttga actgccgtcc accctgtctc
tctacaagtc cggagcactg gacgaggcag 1440ctgcgtacca gagtcgcgac
tactacaact ttccactggc tctggccggg ccgccgcccc 1500ctccaccgcc
tccccatccc cacgctcgca tcaagctgga gaacccgctg gactatggca
1560gcgcctgggc ggctgcggcg gcgcagtgcc gctatgggga cctggcgagc
ctgcatggcg 1620cgggtgcagc gggacccggc tctgggtcac cctcagcggc
cgcttcctca tcctggcaca 1680ctctcttcac agccgaagaa ggccagttgt
atggaccgtg tggtggtggg ggcggcggcg 1740gtggcggcgg cggcggcggc
gcaggcgagg cgggagctgt agccccctac ggctacactc 1800ggccacctca
ggggctggcg ggccaggaag gcgacttcac cgcacctgat gtgtggtacc
1860ctggcggcat ggtgagcaga gtgccctatc ccagtcccac ttgtgtcaaa
agcgagatgg 1920gcccctggat ggatagctac tccggacctt acggggacat
gcgtttggag actgccaggg 1980accatgtttt gccaattgac tattactttc
caccccagaa gacctgcctg atctgtggag 2040atgaagcttc tgggtgtcac
tatggagctc tcacatgtgg aagctgcaag gtcttcttca 2100aaagagccgc
tgaagggaaa cagaagtacc tgtgtgccag cagaaatgat tgcactattg
2160ataaattccg aaggaaaaat tgtccatctt gccgtcttcg gaaatgttat
gaagcaggga 2220tgactctggg agcccggaag ctgaagaaac ttggtaatct
gaaactacag gaggaaggag 2280aggcttccag caccaccagc cccactgagg
agacagccca gaagctgaca gtgtcacaca 2340ttgaaggcta tgaatgtcag
cccatctttc tgaatgtcct ggaggccatt gagccaggtg 2400tggtgtgtgc
tggacatgac aacaaccagc ccgactcctt cgcagccttg ctctctagcc
2460tcaatgaact gggagagaga cagcttgtac atgtggtcaa gtgggccaag
gccttgcctg 2520gcttccgcaa cttacacgtg gacgaccaga tggctgtcat
tcagtactcc tggatggggc 2580tcatggtgtt tgccatgggc tggcgatcct
tcaccaatgt caactccagg atgctctact 2640ttgcccctga tctggttttc
aatgagtacc gcatgcacaa atcccggatg tacagccagt 2700gtgtccgaat
gaggcacctc tctcaagagt ttggatggct ccaaatcacc ccccaggaat
2760tcctgtgcat gaaagcgctg ctactcttca gcattattcc agtggatggg
ctgaaaaatc 2820aaaaattctt tgatgaactt cgaatgaact acatcaagga
actcgatcgt atcattgcat 2880gcaaaagaaa aaatcccaca tcctgctcaa
ggcgtttcta ccagctcacc aagctcctgg 2940actccgtgca gcctattgcg
agagagctgc atcagttcac ttttgacctg ctaatcaagt 3000cacacatggt
gagcgtggac tttccggaaa tgatggcaga gatcatctct gtgcaagtgc
3060ccaagatcct ttctgggaaa gtcaagccca tctatttcca cacccagtga
agcattggaa 3120atccctattt cctcacccca gctcatgccc cctttcagat
gtcttctgcc tgtta 317583920PRTHomo sapiens 83Met Glu Val Gln Leu Gly
Leu Gly Arg Val Tyr Pro Arg Pro Pro Ser1 5 10 15Lys Thr Tyr Arg Gly
Ala Phe Gln Asn Leu Phe Gln Ser Val Arg Glu 20 25 30Val Ile Gln Asn
Pro Gly Pro Arg His Pro Glu Ala Ala Ser Ala Ala 35 40 45Pro Pro Gly
Ala Ser Leu Leu Leu Leu Gln Gln Gln Gln Gln Gln Gln 50 55 60Gln Gln
Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln Gln65 70 75
80Glu Thr Ser Pro Arg Gln Gln Gln Gln Gln Gln Gly Glu Asp Gly Ser
85 90 95Pro Gln Ala His Arg Arg Gly Pro Thr Gly Tyr Leu Val Leu Asp
Glu 100 105 110Glu Gln Gln Pro Ser Gln Pro Gln Ser Ala Leu Glu Cys
His Pro Glu 115 120 125Arg Gly Cys Val Pro Glu Pro Gly Ala Ala Val
Ala Ala Ser Lys Gly 130 135 140Leu Pro Gln Gln Leu Pro Ala Pro Pro
Asp Glu Asp Asp Ser Ala Ala145 150 155 160Pro Ser Thr Leu Ser Leu
Leu Gly Pro Thr Phe Pro Gly Leu Ser Ser 165 170 175Cys Ser Ala Asp
Leu Lys Asp Ile Leu Ser Glu Ala Ser Thr Met Gln 180 185 190Leu Leu
Gln Gln Gln Gln Gln Glu Ala Val Ser Glu Gly Ser Ser Ser 195 200
205Gly Arg Ala Arg Glu Ala Ser Gly Ala Pro Thr Ser Ser Lys Asp Asn
210 215 220Tyr Leu Gly Gly Thr Ser Thr Ile Ser Asp Asn Ala Lys Glu
Leu Cys225 230 235 240Lys Ala Val Ser Val Ser Met Gly Leu Gly Val
Glu Ala Leu Glu His 245 250 255Leu Ser Pro Gly Glu Gln Leu Arg Gly
Asp Cys Met Tyr Ala Pro Leu 260 265 270Leu Gly Val Pro Pro Ala Val
Arg Pro Thr Pro Cys Ala Pro Leu Ala 275 280 285Glu Cys Lys Gly Ser
Leu Leu Asp Asp Ser Ala Gly Lys Ser Thr Glu 290 295 300Asp Thr Ala
Glu Tyr Ser Pro Phe Lys Gly Gly Tyr Thr Lys Gly Leu305 310 315
320Glu Gly Glu Ser Leu Gly Cys Ser Gly Ser Ala Ala Ala Gly Ser Ser
325 330 335Gly Thr Leu Glu Leu Pro Ser Thr Leu Ser Leu Tyr Lys Ser
Gly Ala 340 345 350Leu Asp Glu Ala Ala Ala Tyr Gln Ser Arg Asp Tyr
Tyr Asn Phe Pro 355 360 365Leu Ala Leu Ala Gly Pro Pro Pro Pro Pro
Pro Pro Pro His Pro His 370 375 380Ala Arg Ile Lys Leu Glu Asn Pro
Leu Asp Tyr Gly Ser Ala Trp Ala385 390 395 400Ala Ala Ala Ala Gln
Cys Arg Tyr Gly Asp Leu Ala Ser Leu His Gly 405 410 415Ala Gly Ala
Ala Gly Pro Gly Ser Gly Ser Pro Ser Ala Ala Ala Ser 420 425 430Ser
Ser Trp His Thr Leu Phe Thr Ala Glu Glu Gly Gln Leu Tyr Gly 435 440
445Pro Cys Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly Gly
450 455 460Gly Gly Gly Gly Gly Gly Gly Gly Gly Glu Ala Gly Ala Val
Ala Pro465 470 475 480Tyr Gly Tyr Thr Arg Pro Pro Gln Gly Leu Ala
Gly Gln Glu Ser Asp 485 490 495Phe Thr Ala Pro Asp Val Trp Tyr Pro
Gly Gly Met Val Ser Arg Val 500 505 510Pro Tyr Pro Ser Pro Thr Cys
Val Lys Ser Glu Met Gly Pro Trp Met 515 520 525Asp Ser Tyr Ser Gly
Pro Tyr Gly Asp Met Arg Leu Glu Thr Ala Arg 530 535 540Asp His Val
Leu Pro Ile Asp Tyr Tyr Phe Pro Pro Gln Lys Thr Cys545 550 555
560Leu Ile Cys Gly Asp Glu Ala Ser Gly Cys His Tyr Gly Ala Leu Thr
565 570 575Cys Gly Ser Cys Lys Val Phe Phe Lys Arg Ala Ala Glu Gly
Lys Gln 580 585 590Lys Tyr Leu Cys Ala Ser Arg Asn Asp Cys Thr Ile
Asp Lys Phe Arg 595 600 605Arg Lys Asn Cys Pro Ser Cys Arg Leu Arg
Lys Cys Tyr Glu Ala Gly 610 615 620Met Thr Leu Gly Ala Arg Lys Leu
Lys Lys Leu Gly Asn Leu Lys Leu625 630 635 640Gln Glu Glu Gly Glu
Ala Ser Ser Thr Thr Ser Pro Thr Glu Glu Thr 645 650 655Thr Gln Lys
Leu Thr Val Ser His Ile Glu Gly Tyr Glu Cys Gln Pro 660 665 670Ile
Phe Leu Asn Val Leu Glu Ala Ile Glu Pro Gly Val Val Cys Ala 675 680
685Gly His Asp Asn Asn Gln Pro Asp Ser Phe Ala Ala Leu Leu Ser Ser
690 695 700Leu Asn Glu Leu Gly Glu Arg Gln Leu Val His Val Val Lys
Trp Ala705 710 715 720Lys Ala Leu Pro Gly Phe Arg Asn Leu His Val
Asp Asp Gln Met Ala 725 730 735Val Ile Gln Tyr Ser Trp Met Gly Leu
Met Val Phe Ala Met Gly Trp 740 745 750Arg Ser Phe Thr Asn Val Asn
Ser Arg Met Leu Tyr Phe Ala Pro Asp 755 760 765Leu Val Phe Asn Glu
Tyr Arg Met His Lys Ser Arg Met Tyr Ser Gln 770 775 780Cys Val Arg
Met Arg His Leu Ser Gln Glu Phe Gly Trp Leu Gln Ile785 790 795
800Thr Pro Gln Glu Phe Leu Cys Met Lys Ala Leu Leu Leu Phe Ser Ile
805 810 815Ile Pro Val Asp Gly Leu Lys Asn Gln Lys Phe Phe Asp Glu
Leu Arg 820 825 830Met Asn Tyr Ile Lys Glu Leu Asp Arg Ile Ile Ala
Cys Lys Arg Lys 835 840 845Asn Pro Thr Ser Cys Ser Arg Arg Phe Tyr
Gln Leu Thr Lys Leu Leu 850 855 860Asp Ser Val Gln Pro Ile Ala Arg
Glu Leu His Gln Phe Thr Phe Asp865 870 875 880Leu Leu Ile Lys Ser
His Met Val Ser Val Asp Phe Pro Glu Met Met 885 890 895Ala Glu Ile
Ile Ser Val Gln Val Pro Lys Ile Leu Ser Gly Lys Val 900 905 910Lys
Pro Ile Tyr Phe His Thr Gln 915 92084899PRTMus musculus 84Met Glu
Val Gln Leu Gly Leu Gly Arg Val Tyr Pro Arg Pro Pro Ser1 5 10 15Lys
Thr Tyr Arg Gly Ala Phe Gln Asn Leu Phe Gln Ser Val Arg Glu 20 25
30Ala Ile Gln Asn Pro Gly Pro Arg His Pro Glu Ala Ala Asn Ile Ala
35 40 45Pro Pro Gly Ala Cys Leu Gln Gln Arg Gln Glu Thr Ser Pro Arg
Arg 50 55 60Arg Arg Arg Gln Gln His Thr Glu Asp Gly Ser Pro Gln Ala
His Ile65 70 75 80Arg Gly Pro Thr Gly Tyr Leu Ala Leu Glu Glu Glu
Gln Gln Pro Ser 85 90 95Gln Gln Gln Ala Ala Ser Glu Gly His Pro Glu
Ser Ser Cys Leu Pro 100 105 110Glu Pro Gly Ala Ala Thr Ala Pro Gly
Lys Gly Leu Pro Gln Gln Pro 115 120 125Pro Ala Pro Pro Asp Gln Asp
Asp Ser Ala Ala Pro Ser Thr Leu Ser 130 135 140Leu Leu Gly Pro Thr
Phe Pro Gly Leu Ser Ser Cys Ser Ala Asp Ile145 150 155 160Lys Asp
Ile Leu Asn Glu Ala Gly Thr Met Gln Leu Leu Gln Gln Gln 165 170
175Gln Gln Gln Gln Gln His Gln Gln Gln His Gln Gln His Gln Gln Gln
180 185 190Gln Glu Val Ile Ser Glu Gly Ser Ser Ala Arg Ala Arg Glu
Ala Thr 195 200 205Gly Ala Pro Ser Ser Ser Lys Asp Ser Tyr Leu Gly
Gly Asn Ser Thr 210 215 220Ile Ser Asp Ser Ala Lys Glu Leu Cys Lys
Ala Val Ser Val Ser Met225 230 235 240Gly Leu Gly Val Glu Ala Leu
Glu His Leu Ser Pro Gly Glu Gln Leu 245 250 255Arg Gly Asp Cys Met
Tyr Ala Ser Leu Leu Gly Gly Pro Pro Ala Val 260 265 270Arg Pro Thr
Pro Cys Ala Pro Leu Pro Glu Cys Lys Gly Leu Pro Leu 275 280 285Asp
Glu Gly Pro Gly Lys Ser Thr Glu Glu Thr Ala Glu Tyr Ser Ser 290 295
300Phe Lys Gly Gly Tyr Ala Lys Gly Leu Glu Gly Glu Ser Leu Gly
Cys305 310 315 320Ser Gly Ser Ser Glu Ala Gly Ser Ser Gly Thr Leu
Glu Ile Pro Ser 325 330 335Ser Leu Ser Leu Tyr Lys Ser Gly Ala Leu
Asp Glu Ala Ala Ala Tyr 340 345 350Gln Asn Arg Asp Tyr Tyr Asn Phe
Pro Leu Ala Leu Ser Gly Pro Pro 355 360 365His Pro Pro Pro Pro Thr
His Pro His Ala Arg Ile Lys Leu Glu Asn 370 375 380Pro Leu Asp Tyr
Gly Ser Ala Trp Ala Ala Ala Ala Ala Gln Cys Arg385 390 395 400Tyr
Gly Asp Leu Gly Ser Leu His Gly Gly Ser Val Ala Gly Pro Ser 405 410
415Thr Gly Ser Pro Pro Ala Thr Thr Ser Ser Ser Trp His Thr Leu Phe
420 425 430Thr Ala Glu Glu Gly Gln Leu Tyr Gly Pro Gly Gly Gly Gly
Gly Ser 435 440 445Ser Ser Pro Ser Asp Ala Gly Pro Val Ala Pro Tyr
Gly Tyr Thr Arg 450 455 460Pro Pro Gln Gly Leu Thr Ser Gln Glu Ser
Asp Tyr Ser Ala Ser Glu465 470 475 480Val Trp Tyr Pro Gly Gly Val
Val Asn Arg Val Pro Tyr Pro Ser Pro 485 490 495Asn Cys Val Lys Ser
Glu Met Gly Pro Trp Met Glu Asn Tyr Ser Gly 500 505 510Pro Tyr Gly
Asp Met Arg Leu Asp Ser Thr Arg Asp His Val Leu Pro 515 520 525Ile
Asp Tyr Tyr Phe Pro Pro Gln Lys Thr Cys Leu Ile Cys Gly Asp 530 535
540Glu Ala Ser Gly Cys His Tyr Gly Ala Leu Thr Cys Gly Ser Cys
Lys545 550 555 560Val Phe Phe Lys Arg Ala Ala Glu Gly Lys Gln Lys
Tyr Leu Cys Ala 565 570 575Ser Arg Asn Asp Cys Thr Ile Asp Lys Phe
Arg Arg Lys Asn Cys Pro 580 585 590Ser Cys Arg Leu Arg Lys Cys Tyr
Glu Ala Gly Met Thr Leu Gly Ala 595 600 605Arg Lys Leu Lys Lys Leu
Gly Asn Leu Lys Leu Gln Glu Glu Gly Glu 610 615 620Asn Ser Asn Ala
Gly Ser Pro Thr Glu Asp Pro Ser Gln Lys Met Thr625 630 635 640Val
Ser His Ile Glu Gly Tyr Glu Cys Gln Pro Ile Phe Leu Asn Val 645 650
655Leu Glu Ala Ile Glu Pro Gly Val Val Cys Ala Gly His Asp Asn Asn
660 665 670Gln Pro Asp Ser Phe Ala Ala Leu Leu Ser Ser Leu Asn Glu
Leu Gly 675 680 685Glu Arg Gln Leu Val His Val Val Lys Trp Ala Lys
Ala Leu Pro Gly 690 695 700Phe Arg Asn Leu His Val Asp Asp Gln Met
Ala Val Ile Gln Tyr Ser705 710 715 720Trp Met Gly Leu Met Val Phe
Ala Met Gly
Trp Arg Ser Phe Thr Asn 725 730 735Val Asn Ser Arg Met Leu Tyr Phe
Ala Pro Asp Leu Val Phe Asn Glu 740 745 750Tyr Arg Met His Lys Ser
Arg Met Tyr Ser Gln Cys Val Arg Met Arg 755 760 765His Leu Ser Gln
Glu Phe Gly Trp Leu Gln Ile Thr Pro Gln Glu Phe 770 775 780Leu Cys
Met Lys Ala Leu Leu Leu Phe Ser Ile Ile Pro Val Asp Gly785 790 795
800Leu Lys Asn Gln Lys Phe Phe Asp Glu Leu Arg Met Asn Tyr Ile Lys
805 810 815Glu Leu Asp Arg Ile Ile Ala Cys Lys Arg Lys Asn Pro Thr
Ser Cys 820 825 830Ser Arg Arg Phe Tyr Gln Leu Thr Lys Leu Leu Asp
Ser Val Gln Pro 835 840 845Ile Ala Arg Glu Leu His Gln Phe Thr Phe
Asp Leu Leu Ile Lys Ser 850 855 860His Met Val Ser Val Asp Phe Pro
Glu Met Met Ala Glu Ile Ile Ser865 870 875 880Val Gln Val Pro Lys
Ile Leu Ser Gly Lys Val Lys Pro Ile Tyr Phe 885 890 895His Thr
Gln85895PRTMacaca mulatta 85Met Glu Val Gln Leu Gly Leu Gly Arg Val
Tyr Pro Arg Pro Pro Ser1 5 10 15Lys Thr Tyr Arg Gly Ala Phe Gln Asn
Leu Phe Gln Ser Val Arg Glu 20 25 30Val Ile Gln Asn Pro Gly Pro Arg
His Pro Glu Ala Ala Ser Ala Ala 35 40 45Pro Pro Gly Ala Ser Leu Gln
Gln Gln Gln Gln Gln Gln Gln Glu Thr 50 55 60Ser Pro Arg Gln Gln Gln
Gln Gln Gln Gln Gly Glu Asp Gly Ser Pro65 70 75 80Gln Ala His Arg
Arg Gly Pro Thr Gly Tyr Leu Val Leu Asp Glu Glu 85 90 95Gln Gln Pro
Ser Gln Pro Gln Ser Ala Pro Glu Cys His Pro Glu Arg 100 105 110Gly
Cys Val Pro Glu Pro Gly Ala Ala Val Ala Ala Gly Lys Gly Leu 115 120
125Pro Gln Gln Leu Pro Ala Pro Pro Asp Glu Asp Asp Ser Ala Ala Pro
130 135 140Ser Thr Leu Ser Leu Leu Gly Pro Thr Phe Pro Gly Leu Ser
Ser Cys145 150 155 160Ser Ala Asp Leu Lys Asp Ile Leu Ser Glu Ala
Ser Thr Met Gln Leu 165 170 175Leu Gln Gln Gln Gln Gln Glu Ala Val
Ser Glu Gly Ser Ser Ser Gly 180 185 190Arg Ala Arg Glu Ala Ser Gly
Ala Pro Thr Ser Ser Lys Asp Asn Tyr 195 200 205Leu Glu Gly Thr Ser
Thr Ile Ser Asp Ser Ala Lys Glu Leu Cys Lys 210 215 220Ala Val Ser
Val Ser Met Gly Leu Gly Val Glu Ala Leu Glu His Leu225 230 235
240Ser Pro Gly Glu Gln Leu Arg Gly Asp Cys Met Tyr Ala Pro Val Leu
245 250 255Gly Val Pro Pro Ala Val Arg Pro Thr Pro Cys Ala Pro Leu
Ala Glu 260 265 270Cys Lys Gly Ser Leu Leu Asp Asp Ser Ala Gly Lys
Ser Thr Glu Asp 275 280 285Thr Ala Glu Tyr Ser Pro Phe Lys Gly Gly
Tyr Thr Lys Gly Leu Glu 290 295 300Gly Glu Ser Leu Gly Cys Ser Gly
Ser Ala Ala Ala Gly Ser Ser Gly305 310 315 320Thr Leu Glu Leu Pro
Ser Thr Leu Ser Leu Tyr Lys Ser Gly Ala Leu 325 330 335Asp Glu Ala
Ala Ala Tyr Gln Ser Arg Asp Tyr Tyr Asn Phe Pro Leu 340 345 350Ala
Leu Ala Gly Pro Pro Pro Pro Pro Pro Pro Pro His Pro His Ala 355 360
365Arg Ile Lys Leu Glu Asn Pro Leu Asp Tyr Gly Ser Ala Trp Ala Ala
370 375 380Ala Ala Ala Gln Cys Arg Tyr Gly Asp Leu Ala Ser Leu His
Gly Ala385 390 395 400Gly Ala Ala Gly Pro Gly Ser Gly Ser Pro Ser
Ala Ala Ala Ser Ser 405 410 415Ser Trp His Thr Leu Phe Thr Ala Glu
Glu Gly Gln Leu Tyr Gly Pro 420 425 430Cys Gly Gly Gly Gly Gly Gly
Gly Gly Gly Gly Gly Gly Gly Ala Gly 435 440 445Glu Ala Gly Ala Val
Ala Pro Tyr Gly Tyr Thr Arg Pro Pro Gln Gly 450 455 460Leu Ala Gly
Gln Glu Gly Asp Phe Thr Ala Pro Asp Val Trp Tyr Pro465 470 475
480Gly Gly Met Val Ser Arg Val Pro Tyr Pro Ser Pro Thr Cys Val Lys
485 490 495Ser Glu Met Gly Pro Trp Met Asp Ser Tyr Ser Gly Pro Tyr
Gly Asp 500 505 510Met Arg Leu Glu Thr Ala Arg Asp His Val Leu Pro
Ile Asp Tyr Tyr 515 520 525Phe Pro Pro Gln Lys Thr Cys Leu Ile Cys
Gly Asp Glu Ala Ser Gly 530 535 540Cys His Tyr Gly Ala Leu Thr Cys
Gly Ser Cys Lys Val Phe Phe Lys545 550 555 560Arg Ala Ala Glu Gly
Lys Gln Lys Tyr Leu Cys Ala Ser Arg Asn Asp 565 570 575Cys Thr Ile
Asp Lys Phe Arg Arg Lys Asn Cys Pro Ser Cys Arg Leu 580 585 590Arg
Lys Cys Tyr Glu Ala Gly Met Thr Leu Gly Ala Arg Lys Leu Lys 595 600
605Lys Leu Gly Asn Leu Lys Leu Gln Glu Glu Gly Glu Ala Ser Ser Thr
610 615 620Thr Ser Pro Thr Glu Glu Thr Ala Gln Lys Leu Thr Val Ser
His Ile625 630 635 640Glu Gly Tyr Glu Cys Gln Pro Ile Phe Leu Asn
Val Leu Glu Ala Ile 645 650 655Glu Pro Gly Val Val Cys Ala Gly His
Asp Asn Asn Gln Pro Asp Ser 660 665 670Phe Ala Ala Leu Leu Ser Ser
Leu Asn Glu Leu Gly Glu Arg Gln Leu 675 680 685Val His Val Val Lys
Trp Ala Lys Ala Leu Pro Gly Phe Arg Asn Leu 690 695 700His Val Asp
Asp Gln Met Ala Val Ile Gln Tyr Ser Trp Met Gly Leu705 710 715
720Met Val Phe Ala Met Gly Trp Arg Ser Phe Thr Asn Val Asn Ser Arg
725 730 735Met Leu Tyr Phe Ala Pro Asp Leu Val Phe Asn Glu Tyr Arg
Met His 740 745 750Lys Ser Arg Met Tyr Ser Gln Cys Val Arg Met Arg
His Leu Ser Gln 755 760 765Glu Phe Gly Trp Leu Gln Ile Thr Pro Gln
Glu Phe Leu Cys Met Lys 770 775 780Ala Leu Leu Leu Phe Ser Ile Ile
Pro Val Asp Gly Leu Lys Asn Gln785 790 795 800Lys Phe Phe Asp Glu
Leu Arg Met Asn Tyr Ile Lys Glu Leu Asp Arg 805 810 815Ile Ile Ala
Cys Lys Arg Lys Asn Pro Thr Ser Cys Ser Arg Arg Phe 820 825 830Tyr
Gln Leu Thr Lys Leu Leu Asp Ser Val Gln Pro Ile Ala Arg Glu 835 840
845Leu His Gln Phe Thr Phe Asp Leu Leu Ile Lys Ser His Met Val Ser
850 855 860Val Asp Phe Pro Glu Met Met Ala Glu Ile Ile Ser Val Gln
Val Pro865 870 875 880Lys Ile Leu Ser Gly Lys Val Lys Pro Ile Tyr
Phe His Thr Gln 885 890 8958624DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 86tggggagaac catcctcacc ctgc
248724DNAartificialLNA oligomer Sequence/oligomer Sequence motif
87tccaggacca ggtagcctgt gggg 248824DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 88tgttcccctg gactcagatg ctcc
248924DNAartificialLNA oligomer Sequence/oligomer Sequence motif
89tggggcacaa ggagtgggac gcac 249024DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 90ttcggctgtg aagagagtgt gcca
249124DNAartificialLNA oligomer Sequence/oligomer Sequence motif
91cgcttttgac acaagtggga ctgg 249224DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 92catagtgaca cccagaagct tcat
249324DNAartificialLNA oligomer Sequence/oligomer Sequence motif
93gagtcatccc tgcttcataa catt 249424DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 94gattaccaag tttcttcagc ttcc
249524DNAartificialLNA oligomer Sequence/oligomer Sequence motif
95aggccttggc ccacttgacc acgt 249624DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 96agcatcctgg agttgacatt ggtg
249724DNAartificialLNA oligomer Sequence/oligomer Sequence motif
97gacacactgg ctgtacatcc ggga 249824DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 98gagccatcca aactcttgag agag
249924DNAartificialLNA oligomer Sequence/oligomer Sequence motif
99cagtgctttc atgcacagga attc 2410024DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 100attcgaagtt catcaaagaa tttt
2410124DNAartificialLNA oligomer Sequence/oligomer Sequence motif
101atcgagttcc ttgatgtagt tcat 2410224DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 102gcacttgcac agagatgatc tctg
2410324DNAartificialLNA oligomer Sequence/oligomer Sequence motif
103aatagatggg cttgactttc ccag 2410424DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 104ataacaggca gaagacatct gaaa
2410524DNAartificialLNA oligomer Sequence/oligomer Sequence motif
105attccccaag gcactgcaga ggag 2410624DNAartificialLNA oligomer
Sequence/oligomer Sequence motif 106atgggctgac attcatagcc ttca
24
References











D00001

D00002

D00003
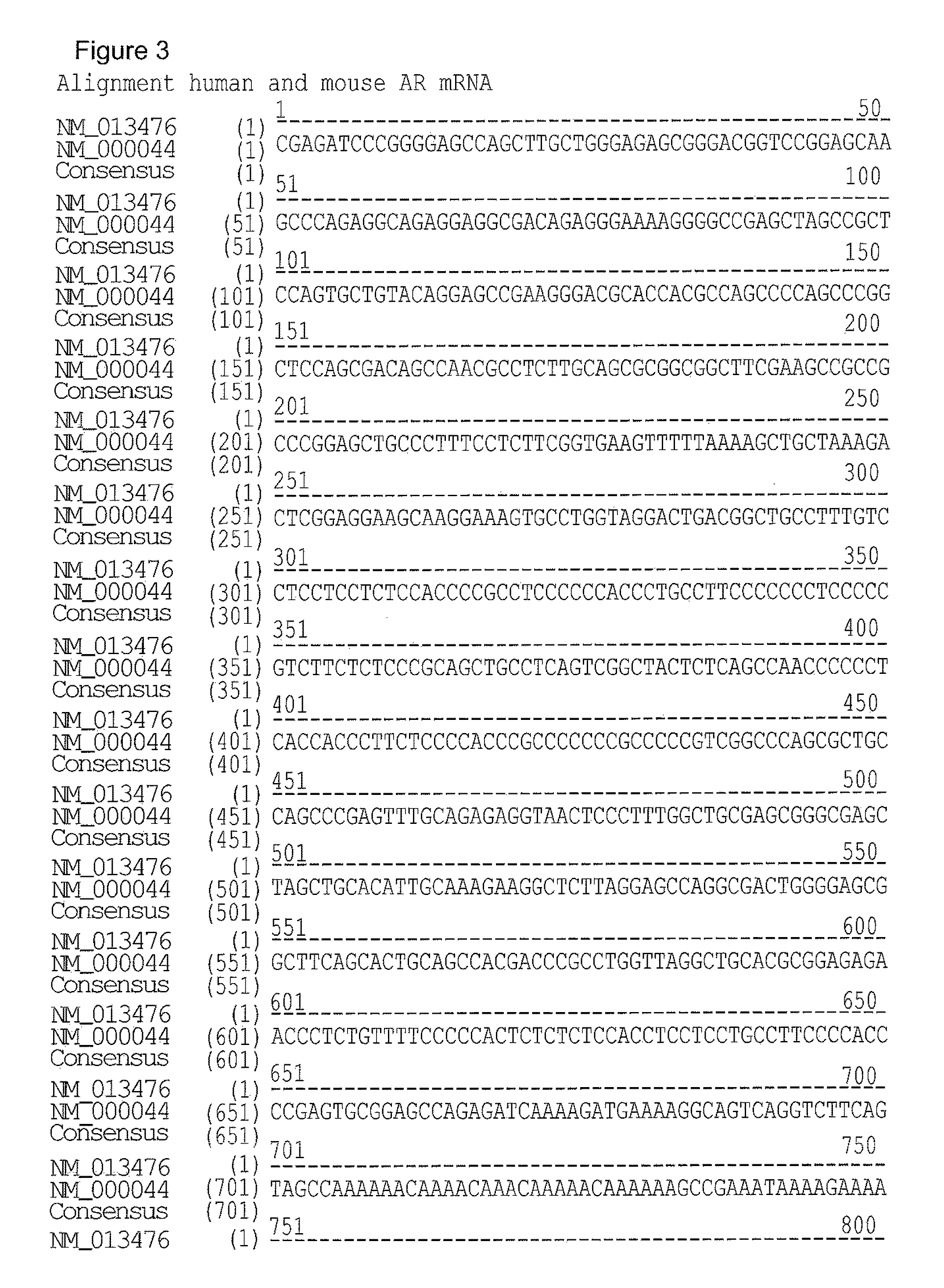

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015

D00016

D00017

D00018

D00019

D00020

D00021

D00022

P00001
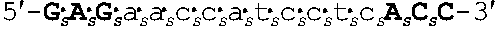
P00002

P00003

P00004

P00005

P00006

P00007

P00008
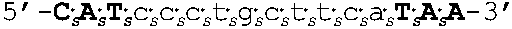
P00009

P00010

P00011

P00012

P00013

P00014

P00015

P00016

P00017

P00018

P00019

P00020

P00021

P00022

P00023

P00024
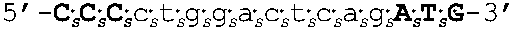
P00025
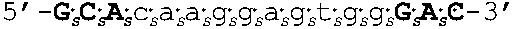
P00026
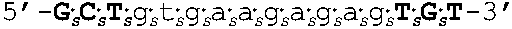
P00027

P00028

P00029

P00030

P00031

P00032

P00033

P00034

P00035

P00036

P00037

P00038

P00039

P00040
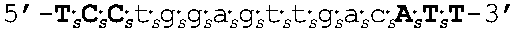
P00041

P00042

P00043

P00044

P00045

P00046

P00047

P00048

P00049
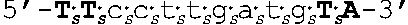
P00050

P00051

P00052

P00053

P00054

P00055

P00056

P00057

P00058

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.