Forensic Identification
BARBER; REBECCA A. L. ; et al.
U.S. patent application number 12/632608 was filed with the patent office on 2010-12-30 for forensic identification. This patent application is currently assigned to FORENSIC SCIENCE SERVICE LTD.. Invention is credited to JENNIFER A. ARNOLD, MICHAEL D. BARBER, REBECCA A. L. BARBER, TRUDY BURKE, PETER P. GILL, SHARON M. GILLBARD, MARC D. HAYWOOD, PETER E. JOHNSON, CAROLYN D. SMITH, ANDREW J. URQUHART.
| Application Number | 20100330559 12/632608 |
| Document ID | / |
| Family ID | 10815013 |
| Filed Date | 2010-12-30 |





| United States Patent Application | 20100330559 |
| Kind Code | A1 |
| BARBER; REBECCA A. L. ; et al. | December 30, 2010 |
FORENSIC IDENTIFICATION
Abstract
The invention provides allelic ladder mixtures and individual alleles suitable for use in such mixtures. The allelic ladder mixtures give improved identification and distinguishing capabilities, particularly suitable in forensic investigations.
| Inventors: | BARBER; REBECCA A. L.; (BIRMINGHAM, GB) ; BARBER; MICHAEL D.; (BIRMINGHAM, GB) ; JOHNSON; PETER E.; (BIRMINGHAM, GB) ; GILLBARD; SHARON M.; (BIRMINGHAM, GB) ; HAYWOOD; MARC D.; (BIRMINGHAM, GB) ; SMITH; CAROLYN D.; (BIRMINGHAM, GB) ; ARNOLD; JENNIFER A.; (BIRMINGHAM, GB) ; BURKE; TRUDY; (BIRMINGHAM, GB) ; URQUHART; ANDREW J.; (BIRMINGHAM, GB) ; GILL; PETER P.; (BIRMINGHAM, GB) |
| Correspondence Address: |
Workman Nydegger;1000 Eagle Gate Tower
60 East South Temple
Salt Lake City
UT
84111
US
|
| Assignee: | FORENSIC SCIENCE SERVICE
LTD. LONDON GB |
| Family ID: | 10815013 |
| Appl. No.: | 12/632608 |
| Filed: | December 7, 2009 |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | ||
|---|---|---|---|---|
| 11821259 | Jun 21, 2007 | 7645580 | ||
| 12632608 | ||||
| 11244331 | Oct 4, 2005 | |||
| 11821259 | ||||
| 09910183 | Jul 20, 2001 | 7087380 | ||
| 11244331 | ||||
| 09706525 | Nov 3, 2000 | |||
| 09910183 | ||||
| 09498567 | Feb 4, 2000 | |||
| 09706525 | ||||
| 09107029 | Jun 29, 1998 | |||
| 09498567 | ||||
| Current U.S. Class: | 435/6.14 ; 536/23.1 |
| Current CPC Class: | C12Q 1/6876 20130101; C07H 21/04 20130101; C12Q 1/6881 20130101; C12Q 2600/172 20130101; C12Q 1/6827 20130101; C12Q 1/6888 20130101; C12Q 2600/156 20130101; C12Q 1/6827 20130101; C12Q 2565/125 20130101; C12Q 2545/113 20130101; C12Q 2525/151 20130101 |
| Class at Publication: | 435/6 ; 536/23.1 |
| International Class: | C12Q 1/68 20060101 C12Q001/68; C07H 21/04 20060101 C07H021/04 |
Foreign Application Data
| Date | Code | Application Number |
|---|---|---|
| Jun 28, 1997 | GB | 9713597.4 |
Claims
1. An allelic ladder mixture comprising one or more of the following allelic ladders: i) an allelic ladder for locus HUMVWFA31/A comprising one or more alleles with a short tandem repeat sequence consisting of sequences: TABLE-US-00052 TCTA TCTG TCTA (TCTG).sub.4 (TCTA).sub.3-; SEQ ID NO 1 or TCTA (TCTG).sub.4 (TCTA).sub.7-; SEQ ID NO 2 or (TCTA).sub.2 (TCTG).sub.4 (TCTA).sub.3 TCCA SEQ ID NO 3 (TCTA).sub.3-;
ii) an allelic ladder for locus HUMTHO1 comprising an allele with a short tandem repeat sequence consisting of sequence: TABLE-US-00053 (TCAT).sub.4 CAT (TCAT).sub.7 TCGT TCAT-; SEQ ID NO 4
iii) an allelic ladder for locus D8S1179 comprising an allele with a short tandem repeat sequence consisting of sequence: TABLE-US-00054 (TCTA).sub.2 TCTG(TCTA).sub.16-; SEQ ID NO 6
iv) an allelic ladder for locus HUMFIBRA/FGA comprising one or more alleles with a short tandem repeat sequence consisting of sequences: TABLE-US-00055 SEQ ID NO 7 (TTTC).sub.3 TTTT TTCT (CTTT).sub.5 T (CTTT).sub.3 CTCC (TTCC).sub.2-; or SEQ ID NO 8 (TTTC).sub.3 TTTT TTCT (CTTT).sub.13 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2-; or SEQ ID NO 9 (TTTC).sub.3 TTTT TTCT (CTTT).sub.16 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2-; or SEQ ID NO 10 (TTTC).sub.4 TTTT TT (CTTT).sub.15 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 11 (TTTC).sub.4 TTTT TT (CTTT).sub.16 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 12 (TTTC).sub.4 TTTT TT (CTTT).sub.17 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 13 (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.4 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 14 (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 15 (TTTC).sub.4 TTTT TT (CTTT).sub.11 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 16 (TTTC).sub.4 TTTT TT (CTTT).sub.10 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 17 (TTTC).sub.4 TTTT TT (CTTT).sub.12 (CTGT).sub.5 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 18 (TTTC).sub.4 TTTT TT (CTTT).sub.14 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-;
v) an allelic ladder for locus D21S11 comprising one or more alleles with a short tandem repeat sequence consisting of sequences: TABLE-US-00056 SEQ ID NO 19 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA(TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.6 TCGTCT-; or SEQ ID NO 20 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT- or SEQ ID NO 21 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT-; or SEQ ID NO 22 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.8 TCGTCT-; or SEQ ID NO 23 (TCTA).sub.5 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT-; or SEQ ID NO 24 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT-; or SEQ ID NO 25 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT-; or SEQ ID NO 26 (TCTA).sub.6 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT-; or SEQ ID NO 27 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-; or SEQ ID NO 28 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TA TCTA TCGTCT-; or SEQ ID NO 29 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TA TCTA TCGTCT-; or SEQ ID NO 30 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TA TCTA TCGTCT-; or SEQ ID NO 31 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.14 TATCTA TCGTCT-; or SEQ ID NO 32 (TCTA).sub.10 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-; or SEQ ID NO 33 (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-; or SEQ ID NO 34 (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TCGTCT-; or SEQ ID NO 35 (TCTA).sub.13 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-;
vi) an allelic ladder for locus D18S51 comprising an allele with a short tandem repeat sequence consisting of sequence: TABLE-US-00057 (AGAA).sub.8-. SEQ ID NO 36
2. An allelic ladder mixture according to claim 1 in which the mixture includes allelic ladders for a plurality of loci selected from HUMVWFA31/A, HUMTHO1, D8S1179, HUMFIBRA/FGA, D21S11 and D18S51.
3. An allelic ladder mixture according to claim 1 the mixture including allelic ladders for at least four loci.
4. An allelic ladder mixture according to claim 1 the mixture includes an amelogenin sex test.
5. An allelic ladder mixture according to claim 1 in which the allelic ladders in the mixture each include at least 7 alleles.
6. An allelic ladder mixture according to claim 1 in which the ladders, if present in the mixture, are provided such that: the HUMVWFA31/A allelic ladder includes at least 9 alleles; the HUMTHO1 allelic ladder includes at least 7; the D8S1179 allelic ladder includes at least 9 alleles; the HUMFIBRA/FGA allelic ladder includes at least 18 alleles or is present as HUMFIBRA/F GA/LW and HUMFIBRA/FGA/HW with the HUMFIBRA/FGA/LW ladder including at least 16 alleles, the HUMFIBRA/FGA/HW ladder including at least 6 alleles; the D21S11 allelic ladder includes at least 14 alleles; and the D18S51 ladder includes at least 15 alleles.
7. An allelic ladder mixture according to claim 1 in which one or more of the allelic ladders in the mixture comprises at least 4 pairs of alleles 4 base pairs from each other.
8. An allelic ladder mixture according to claim 1 in which the ladders, if present in the mixture, are provided such that: the HUMVWFA31/A allelic ladder includes at least 7 pairs of alleles 4 base pairs from each other; the HUMTHO1 allelic ladder includes at least 5 pairs of alleles 4 base pairs from each other; the D8S1179 allelic ladder includes at least 8 pairs of alleles 4 base pairs from each other; the HUMFIBRA/FGA allelic ladder includes at least 17 pairs of alleles 4 base pairs from each other; the D21S11 allelic ladder includes at least 3 pairs of alleles 4 base pairs from each other; and the D18S51 ladder includes at least 13 pairs of alleles 4 base pairs from each other.
9. An allelic ladder mixture according to claim 8 in which the D21S11 allelic ladder includes at least 8 pairs of alleles 8 base pairs from each other.
10. An allelic ladder mixture according to claim 1 in which the ladders, if present, are provided such that the HUMVWFA31/A ladder includes alleles ranging from 130 base pairs upwards and/or from 166 base pairs downwards; the HUMTHO1 ladder includes alleles ranging from 150 base pairs upwards and/or 189 base pairs downwards; the D8S1179 ladder includes alleles ranging from 157 base pairs upwards and/or 201 base pairs downwards; the HUMFIBRA/FGA ladder includes alleles ranging from 173 base pairs upwards and/or 298 base pairs downwards; the D21S11 ladder includes alleles ranging from 203 base pairs upwards and/or 255 base pairs downwards; and the D18S51 ladder includes alleles ranging from 270 base pairs upwards and/or 326 downwards.
11. An allelic ladder mixture comprising an allelic ladder for one or more of the following loci, with lowest and highest allele designation as follows: TABLE-US-00058 Lowest Highest Locus Designation Designation a) HUMVWFA31/A 10 21 b) HUMTH01 4 13.3 c) D8S1179 7 19 d) HUMFIBRA/FGA 16.1 50.2 e) D21S11 53 81 f) D18S51 8 27
12. An allelic ladder mixture according to claim 11 in which the loci ladders have both the upper and lower limits specified.
13. An allelic ladder mixture according to claim 11 in which the mixture includes allelic ladders for loci HUMVWFA31/A, HUMTHO1, D8S1179, HUMFIBRA/F GA, D21S11 and D18S51.
14. A method of analysing one or more samples comprising: a) obtaining genomic DNA from the sample; b) amplifying the DNA; c) obtaining an indication of one or more of the constituent parts of the sample; and comparing the indications with an allelic ladder mixture comprising one or more of the following allelic ladders: i) an allelic ladder for locus HUMVWFA31/A comprising one or more alleles with a short tandem repeat sequence consisting of sequences: TABLE-US-00059 SEQ ID NO 1 TCTA TCTG TCTA (TCTG).sub.4 (TCTA).sub.3-; or SEQ ID NO 2 TCTA (TCTG).sub.4 (TCTA).sub.7-; or SEQ ID NO 3 (TCTA).sub.2 (TCTG).sub.4 (TCTA).sub.3 TCCA (TCTA).sub.3-
ii) an allelic ladder for locus HUMTHO1 comprising an allele with a short tandem repeat sequence consisting of sequence: TABLE-US-00060 (TCAT).sub.4 CAT (TCAT).sub.7 TCGT TCAT-; SEQ ID NO 4
iii) an allelic ladder for locus D8S1179 comprising an allele with a short tandem repeat sequence consisting of sequence: TABLE-US-00061 (TCTA).sub.2 TCTG (TCTA).sub.16-; SEQ ID NO 6
iv) an allelic ladder for locus HUMFIBRA/FGA comprising one or more alleles with a short tandem repeat sequence consisting of the sequences: TABLE-US-00062 SEQ ID NO 7 (TTTC).sub.3 TTTT TTCT (CTTT).sub.5 T (CTTT).sub.3 CTCC (TTCC).sub.2-; or SEQ ID NO 8 (TTTC).sub.3 TTTT TTCT (CTTT).sub.13 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2-; or SEQ ID NO 9 (TTTC).sub.3 TTTT TTCT (CTTT).sub.16 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2-; or SEQ ID NO 10 (TTTC).sub.4 TTTT TT (CTTT).sub.15 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 11 (TTTC).sub.4 TTTT TT (CTTT).sub.16 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 12 (TTTC).sub.4 TTTT TT (CTTT).sub.17 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 13 (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.4 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 14 (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 15 (TTTC).sub.4 TTTT TT (CTTT).sub.11 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 16 (TTTC).sub.4 TTTT TT (CTTT).sub.10 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 17 (TTTC).sub.4 TTTT TT (CTTT).sub.12 (CTGT).sub.5 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4-; or SEQ ID NO 18 (TTTC).sub.4 TTTT TT (CTTT).sub.14 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4-;
v) an allelic ladder for locus D21S11 comprising one or more alleles with a short tandem repeat sequence consisting of sequences: TABLE-US-00063 SEQ ID NO 19 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA(TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.6 TCGTCT-; or SEQ ID NO 20 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT-; or SEQ ID NO 21 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT-; or SEQ ID NO 22 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.8 TCGTCT-; or SEQ ID NO 23 (TCTA).sub.5 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT-; or SEQ ID NO 24 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT-; or SEQ ID NO 25 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT-; or SEQ ID NO 26 (TCTA).sub.6 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT-; or SEQ ID NO 27 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-; or SEQ ID NO 28 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TA TCTA TCGTCT-; or SEQ ID NO 29 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TA TCTA TCGTCT-; or SEQ ID NO 30 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TA TCTA TCGTCT-; or SEQ ID NO 31 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.14 TATCTA TCGTCT-; or SEQ ID NO 32 (TCTA).sub.10 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-; or SEQ ID NO 33 (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-; or SEQ ID NO 34 (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TCGTCT-; or SEQ ID NO 35 (TCTA).sub.13 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT-;
vi) an allelic ladder for locus D18S51 comprising an allele with a short tandem repeat sequence consisting of sequence: TABLE-US-00064 (AGAA).sub.8-. SEQ ID NO 36
15. A method according to claim 14 in which the DNA sample is one or more of a sample taken from the scene of a crime, a sample associated with the scene of a crime, a sample obtained from a suspect, a sample obtained from a human under consideration or a reference sample.
16. A method according to claim 14 in which the sample is amplified using a polymerase chain reaction and primers for one or more of loci HUMVWFA31/A, HUMTHO1, D8S1179, HUMFIBRA/FGA, D21S11 or D18S51 are employed.
17. One or more alleles comprising or consisting of sequences TCTA TCTG TCTA (TCTG).sub.4 (TCTA).sub.3; TCTA (TCTG).sub.4 (TCTA).sub.7; (TCTA).sub.2 (TCTG).sub.4 (TCTA).sub.3, TCCA (TCTA).sub.3; (TCAT).sub.4 CAT (TCAT).sub.7 TCGT TCAT; (TCTA).sub.8; (TCTA).sub.2 TCTG (TCTA).sub.16; (TTTC).sub.3 TTTT TTCT (CTTT).sub.5 T (CTTT).sub.3 CTCC (TTCC).sub.2; (TTTC).sub.3 TTTT TTCT (CTTT).sub.13 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2; (TTTC).sub.3 TTTT TTCT (CTTT).sub.16 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2; (TTTC).sub.4 TTTT TT (CTTT).sub.15 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT).sub.16 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT)17 (CTTC)3 (CTTT)3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.4 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT).sub.11 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT).sub.10 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT).sub.12 (CTGT).sub.5 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (TTTC).sub.4 TTTT TT (CTTT).sub.14 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4; (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.6 TCGTCT; (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT; (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT; (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.8 TCGTCT; (TCTA).sub.5 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT; (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT; (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT; (TCTA).sub.6 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT; (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT; (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TA TCTA TCGTCT; (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TA TCTA TCGTCT; (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TA TCTA TCGTCT; (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.14 TATCTA TCGTCT; (TCTA).sub.10 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT; (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT; (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TCGTCT; (TCTA).sub.13 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT; or (AGAA).sub.8; or at least 75% homologous thereto.
18. One or more alleles according to claim 16 in which the alleles are provided purified from alleles other than those of HUMVWFA31/A, HUMTHO1, D8S1179, HUMFIBRA/FGA, D21S11, D18S51 or AMG loci.
19. The use of an allelic ladder according to claim 1 for comparison with a DNA analysis result.
20. The use of claim 19, wherein the analysis is a DNA profile of a sample and the profile is based on analysis of one or more of loci HUMVWFA31/A, HUMTHO1, D9S1179, HUMFIBRA/FGA, D21S11, D18S51 or AMG.
Description
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a continuation of U.S. patent application Ser. No. 11/821,259, filed Jun. 21, 2007, which is a continuation of U.S. patent application Ser. No. 11/244,331, filed Oct. 4, 2005, which is a continuation of U.S. patent application Ser. No. 09/910,183, filed Jul. 20, 2001, now U.S. Pat. No. 7,087,380, issued Aug. 8, 2006, which is a continuation of U.S. patent application Ser. No. 09/706,525, filed Nov. 3, 2000 (abandoned), which is a continuation of U.S. patent application Ser. No. 09/498,567, filed Feb. 4, 2000 (abandoned), which is a continuation of U.S. patent application Ser. No. 09/107,029, filed Jun. 29, 1998 (abandoned), which claims priority to United Kingdom Application No. 9713597.4, filed Jun. 28, 1997, which for purposes of disclosure are incorporated herein by specific reference.
BACKGROUND OF THE INVENTION
[0002] 1. The Field of the Invention
[0003] The present invention is concerned with improvements in and relating to forensic identification, particularly where based on DNA profiling.
[0004] 2. The Relevant Technology
[0005] DNA profiling offers a versatile identification technique for a wide variety of applications including, anthropological, paternity and other forensic environments. The use of such profiling is significant in determining links, or their absence, between samples. Such samples might include those taken from known individuals and/or those taken from the scene of or linked to a crime.
[0006] DNA profiling based on the use of short tandem repeats (STR) or micro satellite loci is used in such applications. STR's are a class of polymorphic markers which consist of simple tandomly repeated sequences of between 1 and 6 base pairs in length. STR's in the non-coding part of the genome are generally considered.
[0007] In the human genome STR's occur every 6 to 10 kilo bases along the DNA. The length, however, varies greatly between individuals due to mutation and provides identifying characteristics as a result.
[0008] A variety of DNA profiling systems exist, including single locus analysis and multiple locus analysis where a number of STR loci are simultaneously amplified.
[0009] In analysing the results from an unknown sample it is generally considered against a ladder marker consisting of alleles derived from actual samples. The allelic ladder provides a reference point and allows correspondence of alleles to be identified clearly.
SUMMARY OF THE INVENTION
[0010] The present invention provides new alleles and new ladders incorporating them for a variety of loci. The present invention offers an improved range and coverage of markers as a result. The ladders include a number of rare alleles offering improved identification of the alleles in an unknown sample.
[0011] According to a first aspect of the invention we provide an allelic ladder mixture comprising one or more of the following allelic ladders:
[0012] i) an allelic ladder for locus HUMVWFA31/A comprising one or more of alleles comprising or consisting of sequences: [0013] TCTA TCTG TCTA (TCTG).sub.4 (TCTA).sub.3 (SEQ ID NO: 1); [0014] TCTA (TCTG).sub.4 (TCTA).sub.7 (SEQ ID NO: 2); or [0015] (TCTA).sub.2 (TCTG).sub.4 (TCTA).sub.3 TCCA (TCTA).sub.3 (SEQ ID NO: 3) or at least 75% homologous thereto;
[0016] ii) an allelic ladder for locus HUMTHO1 comprising or consisting of sequence: (TCAT).sub.4 CAT (TCAT).sub.7 TCGT TCAT (SEQ ID NO: 4); or at least 75% homologous thereto;
[0017] iii) an allelic ladder for locus D8S1179 comprising one or more of alleles: [0018] (TCTA).sub.8 (SEQ ID NO: 5); [0019] (TCTA).sub.2 TCTG(TCTA).sub.16 (SEQ ID NO: 6) or at least 75% homologous thereto;
[0020] iv) an allelic ladder for locus HUMFIBRA/FGA comprising one or more of alleles comprising or consisting of the sequences: [0021] (TTTC).sub.3 TTTT TTCT (CTTT).sub.5 T (CTTT).sub.3 CTCC (TTCC).sub.2 (SEQ ID NO: 7); [0022] (TTTC).sub.3 TTTT TTCT (CTTT).sub.13 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2 (SEQ ID NO: 8); [0023] (TTTC).sub.3 TTTT TTCT (CTTT).sub.16 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2 (SEQ ID NO: 9); [0024] (TTTC).sub.4 TTTT TT (CTTT).sub.15 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 10); [0025] (TTTC).sub.4 TTTT TT (CTTT).sub.16 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 11); [0026] (TTTC).sub.4 TTTT TT (CTTT).sub.17 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 12); [0027] (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.4 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 13); [0028] (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 14); [0029] (TTTC).sub.4 TTTT TT (CTTT).sub.11 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 15); [0030] (TTTC).sub.4 TTTT TT (CTTT).sub.10 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 16); [0031] (TTTC).sub.4 TTTT TT (CTTT).sub.12 (CTGT).sub.5 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 17); or [0032] (TTTC).sub.4 TTTT TT (CTTT).sub.14 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 18); or at least 75% homologous thereto;
[0033] v) an allelic ladder for locus D21S11 comprising one or more of alleles comprising or consisting of sequences: (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA(TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.6 TCGTCT (SEQ ID NO: 19); [0034] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT (SEQ ID NO: 20); [0035] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT (SEQ ID NO: 21); [0036] (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.8 TCGTCT (SEQ ID NO: 22); [0037] (TCTA).sub.5 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT (SEQ ID NO: 23); [0038] (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT (SEQ ID NO: 24); [0039] (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT (SEQ ID NO: 25); [0040] (TCTA).sub.6 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT (SEQ ID NO: 26); [0041] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 27); [0042] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TA TCTA TCGTCT (SEQ ID NO: 28); [0043] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.3 TCCATA (TCTA).sub.12 TA TCTA TCGTCT (SEQ ID NO: 29); [0044] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TA TCTA TCGTCT (SEQ ID NO: 30); [0045] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.14 TATCTA TCGTCT (SEQ ID NO: 31); [0046] (TCTA).sub.10 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 32); [0047] (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 33); [0048] (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 [0049] (TCTA).sub.13 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 35); or at least 75% homologous thereto;
[0050] vi) an allelic ladder for locus D18S51 comprising an allele comprising or consisting of sequence: [0051] (AGAA).sub.8 (SEQ ID NO: 36); or at least 75% homologous thereto.
[0052] Preferably the mixture includes allelic ladders for a plurality of loci. It is particularly preferred that the mixture include allelic ladders for at least four loci. Preferably the mixture includes allelic ladders for a plurality of loci selected from HUMVWFA31/A, HUMTHO1, D8 S1179, HUMFIBRA/FGA, D21S11 and D18S51. Preferably the mixture includes allelic ladders for at least four of these loci. In its most preferred form the mixture includes allelic ladders for all of these loci.
[0053] Preferably the mixture includes an amelogenin sex test.
[0054] Preferably one or more of the allelic ladders in the mixture includes at least 7 alleles and more preferably at least 12 alleles. Preferably a plurality, and particularly all, of the allelic ladders of the mixture include at least 8 and more preferably at least 10 alleles.
[0055] Preferably one or more or all of the ladders, if present in the mixture, may be provided such that: the HUMVWFA31/A allelic ladder includes at least 9, more preferably 11 and ideally 12 alleles; the HUMTHO1 allelic ladder includes at least 7, more preferably 9 and ideally 10 alleles; the D8S1179 allelic ladder includes at least 9, more preferably 12 and ideally 13 alleles; the HUMFIBRA/FGA allelic ladder includes at least 18, more preferably 26 and ideally 28 alleles or is present as HUMFIBRA/FGA/LW and HUMFIBRA/FGA/HW with the HUMFIBRA/FGA/LW ladder including at least 16 more preferably 18 and ideally 20 alleles, the HUMFIBRA/FGA/HW ladder including at least 6, more preferably at least 7 and ideally 8 alleles; the D21S11 allelic ladder includes at least 14, more preferably 16 and ideally 17 alleles; and the D18S51 ladder includes at least 15, more preferably 19 and ideally 20 alleles.
[0056] Preferably one or more of the allelic ladders in the mixture comprises at least 4 pairs of alleles 4 base pairs from each other. More preferably at least 10 pairs, and ideally at least 12 pairs of alleles are so provided. Preferably one or more or all the allelic ladders, if present in the mixture, may be provided such that: the HUMVWFA31/A allelic ladder includes at least 7, more preferably 10 and ideally 11 pairs of alleles 4 base pairs from each other; the HUMTHO1 allelic ladder includes at least 5, more preferably 6 and ideally 7 pairs of alleles 4 base pairs from each other; the D8S1179 allelic ladder includes at least 8, more preferably 11 and ideally 12 pairs of alleles 4 base pairs from each other; the HUMFIBRA/FGA allelic ladder includes at least 17, more preferably 20 and ideally 23 pairs of alleles 4 base pairs from each other; the D21S11 allelic ladder includes at least 3 and ideally 4 pairs of alleles 4 base pairs from each other; and the D18S51 ladder includes at least 13, more preferably 18 and ideally 19 pairs of alleles 4 base pairs from each other. The D21S11 allelic ladder may, or may further include, at least 8, more preferably 11 and ideally 12 pairs of alleles 8 base pairs from each other.
[0057] Preferably the allele sequences have at least 85% homogeneity with the listed sequences. More preferable levels of even 90% or at least 95% may be provided. Ideally the exact sequences listed are included within the alleles. In their most preferred form the alleles consist of the listed sequences.
[0058] The alleles may further include flanking sequences, i.e., between the primer and STR.
[0059] Preferably the HUMVWFA31/A ladder includes alleles ranging from 130, more preferably 126 and ideally 122 base pairs upwards and/or from 166 base pairs downwards. Preferably the HUMTHO1 ladder includes alleles ranging from 150 base pairs upwards and/or 189 base pairs downwards. Preferably the D8S1179 ladder includes alleles ranging from 157 base pairs upwards and/or 201, and more preferably 205 base pairs downwards. Preferably the HUMFIBRA/FGA ladder includes alleles ranging from 173 base pairs upwards and/or 298, more preferably 302 and ideally 310 base pairs downwards. Preferably the D21S11 ladder includes alleles ranging from 203 base pairs upwards and/or 255 or more preferably 259 base pairs downwards. Preferably the D18S51 ladder includes alleles ranging from 270 or more preferably 266 base pairs upwards and/or 326 or 330 or 334 or 338 or even 342 downwards.
[0060] According to a second aspect of the invention we provide an allelic ladder mixture comprising an allelic ladder for one or more of the following loci, with lowest molecular weight allele and/or uppermost molecular weight allele as follows:
TABLE-US-00001 Locus Low MW allele High MW allele a) HUMVWFA31/A 10 21 b) HUMTH01 4 13.3 c) D8S1179 7 19 d) HUMFIBRA/FGA 16.1 50.2 e) D21S11 53 81 f) D18S51 8 27
[0061] Preferably one or more of the loci ladders have both the upper and lower limits specified. Preferably all the loci ladders have the full ranges listed.
[0062] Preferably the mixture includes allelic ladders for a plurality of loci. It is particularly preferred that the mixture include allelic ladders for at least four loci. Preferably the mixture includes allelic ladders for a plurality of loci selected from HUMVWFA31/A, HUMTHO1, D8S1179, HUMFIBRA/FGA, D21S11 and D18S51. Preferably the mixture includes allelic ladders for at least four of these loci. In its most preferred form the mixture includes allelic ladders for all of these loci.
[0063] The intervals of alleles in the ladders and/or number of alleles in the ladders may be as specified in the first aspect of the invention. This aspect may include any of the other features specified elsewhere in the application.
[0064] The ladder mixtures of the first and/or second aspect of the invention may further include one or more of PARR buffer, primer(s), or Taq polymerase.
[0065] According to a third aspect of the invention we provide a method of analysing one or more samples comprising:--
[0066] a) obtaining genomic DNA from the sample;
[0067] b) amplifying the DNA;
[0068] c) obtaining an indication of one or more of the constituent parts of the sample; and comparing the indications with an allelic ladder mixture comprising one or more of the following allelic ladders:
[0069] i) an allelic ladder for locus HUMVWFA31/A comprising one or more of alleles comprising or consisting of sequences:
TABLE-US-00002 TCTA TCTG TCTA (TCTG).sub.4 (TCTA).sub.3; (SEQ ID NO: 1) TCTA (TCTG).sub.4 (TCTA).sub.7; (SEQ ID NO: 2) or (TCTA).sub.2 (TCTG).sub.4 (TCTA).sub.3 TCCA (SEQ ID NO: 3) (TCTA).sub.3
[0070] ii) an allelic ladder for locus HUMTHO1 comprising or consisting of sequence:
TABLE-US-00003 (TCAT).sub.4 CAT (TCAT).sub.7 TCGT TCAT; (SEQ ID NO: 4)
[0071] iii) an allelic ladder for locus D8S1179 comprising one or more of alleles comprising or consisting of sequences:
TABLE-US-00004 (TCTA).sub.8; (SEQ ID NO: 5) or (TCTA).sub.2 TCTG (TCTA).sub.16; (SEQ ID NO: 6)
[0072] iv) an allelic ladder for locus HUMFIBRA/FGA comprising one or more of alleles comprising or consisting of the sequences:
TABLE-US-00005 (SEQ ID NO: 7) (TTTC).sub.3 TTTT TTCT (CTTT).sub.5 T (CTTT).sub.3 CTCC (TTCC).sub.2; (SEQ ID NO: 8) (TTTC).sub.3 TTTT TTCT (CTTT).sub.13 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2; (SEQ ID NO: 9) (TTTC).sub.3 TTTT TTCT (CTTT).sub.16 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2; (SEQ ID NO: 10) (TTTC).sub.4 TTTT TT (CTTT).sub.15 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (SEQ ID NO: 11) (TTTC).sub.4 TTTT TT (CTTT).sub.16 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (SEQ ID NO: 12) (TTTC).sub.4 TTTT TT (CTTT).sub.17 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (SEQ ID NO: 13) (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.4 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4; (SEQ ID NO: 14) (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4; (SEQ ID NO: 15) (TTTC).sub.4 TTTT TT (CTTT).sub.11 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; (SEQ ID NO: 16) (TTTC).sub.4 TTTT TT (CTTT).sub.10 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4; (SEQ ID NO: 17) (TTTC).sub.4 TTTT TT (CTTT).sub.12 (CTGT).sub.5 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4; or (SEQ ID NO: 18) (TTTC).sub.4 TTTT TT (CTTT).sub.14 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4;
[0073] v) an allelic ladder for locus D21S11 comprising one or more of alleles comprising or consisting of sequences:
TABLE-US-00006 (SEQ ID NO: 19) (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA(TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.6 TCGTCT; (SEQ ID NO: 20) (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT; (SEQ ID NO: 21) (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT; (SEQ ID NO: 22) (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.8 TCGTCT; (SEQ ID NO: 23) (TCTA).sub.5 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT; (SEQ ID NO: 24) (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT; (SEQ ID NO: 25) (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT; (SEQ ID NO: 26) (TCTA).sub.6 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT; (SEQ ID NO: 27) (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT; (SEQ ID NO: 28) (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TA TCTA TCGTCT; (SEQ ID NO: 29) (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TA TCTA TCGTCT; (SEQ ID NO: 30) (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TA TCTA TCGTCT; (SEQ ID NO: 31) (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.14 TATCTA TCGTCT; (SEQ ID NO: 32) (TCTA).sub.10 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT; (SEQ ID NO: 33) (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT; (SEQ ID NO: 34) (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TCGTCT; or (SEQ ID NO: 35) (TCTA).sub.13 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT;
[0074] vi) an allelic ladder for locus D18S51 comprising an allele comprising or consisting of sequence:
TABLE-US-00007 (AGAA).sub.8; (SEQ ID NO: 36)
[0075] including allelic ladders or alleles 75% homologous thereto.
[0076] The allelic ladder mixture may possess other features specified in the first or second aspects of the invention or elsewhere in this application.
[0077] Preferably the DNA sample is one or more of a sample taken from the scene of a crime, a sample associated with the scene of a crime, a sample obtained from a suspect, a sample obtained from a human under consideration (for instance for paternity or maternity analysis) or a reference sample. The sample may be in the form of blood, hair, skin or bodily fluid.
[0078] Preferably the sample is amplified using a polymerase chain reaction. Preferably primers for one or more of loci HUMVWFA31/A, HUMTHO1, D8S1179, HUMFIBRA/FGA, D21S11 or D18S51 are employed. The primers may be dye or otherwise labelled.
[0079] According to a fourth aspect of the invention we provide one or more alleles comprising or consisting of sequences: [0080] TCTA TCTG TCTA (TCTG).sub.4 (TCTA).sub.3 (SEQ ID NO: 1); [0081] TCTA (TCTG).sub.4 (TCTA).sub.7 (SEQ ID NO: 2); [0082] (TCTA).sub.2 (TCTG).sub.4 (TCTA).sub.3 TCCA (TCTA).sub.3 (SEQ ID NO: 3); [0083] (TCAT).sub.4 CAT (TCAT).sub.7 TCGT TCAT (SEQ ID NO: 4); [0084] (TCTA).sub.8 (SEQ ID NO: 5); [0085] (TCTA).sub.2 TCTG (TCTA).sub.16 (SEQ ID NO: 6); [0086] (TTTC).sub.3 TTTT TTCT (CTTT).sub.5 T (CTTT).sub.3 CTCC (TTCC).sub.2 (SEQ ID NO: 7); [0087] (TTTC).sub.3 TTTT TTCT (CTTT).sub.13 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2 (SEQ ID NO: 8); [0088] (TTTC).sub.3 TTTT TTCT (CTTT).sub.16 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2 (SEQ ID NO: 9); [0089] (TTTC).sub.4 TTTT TT (CTTT).sub.15 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 10); [0090] (TTTC).sub.4 TTTT TT (CTTT).sub.16 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 11); [0091] (TTTC).sub.4 TTTT TT (CTTT).sub.17 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 12); [0092] (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.4 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 13); [0093] (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 14); [0094] (TTTC).sub.4 TTTT TT (CTTT).sub.11 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 15); [0095] (TTTC).sub.4 TTTT TT (CTTT).sub.10 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 16); [0096] (TTTC).sub.4 TTTT TT (CTTT).sub.12 (CTGT).sub.5 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 17); [0097] (TTTC).sub.4 TTTT TT (CTTT).sub.14 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 18); [0098] (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA(TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.6 TCGTCT (SEQ ID NO: 19); [0099] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT (SEQ ID NO: 20); [0100] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT (SEQ ID NO: 21); [0101] (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.8 TCGTCT (SEQ ID NO: 22); [0102] (TCTA).sub.5 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT (SEQ ID NO: 23); [0103] (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT (SEQ ID NO: 24); [0104] (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT (SEQ ID NO: 25); [0105] (TCTA).sub.6 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT (SEQ ID NO: 26); [0106] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 27); [0107] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TA TCTA TCGTCT (SEQ ID NO: 28); [0108] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TA TCTA TCGTCT (SEQ ID NO: 29); [0109] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TA TCTA TCGTCT (SEQ ID NO: 30); [0110] (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.14 TATCTA TCGTCT (SEQ ID NO: 31); [0111] (TCTA).sub.10 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 32); [0112] (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 33); [0113] (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TCGTCT (SEQ ID NO: 34); [0114] (TCTA).sub.13 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 35); or [0115] (AGAA).sub.8 (SEQ ID NO: 36); or at least 75% homologous thereto.
[0116] Preferably the alleles are provided purified from alleles other than those of HUMVWFA31/A, HUMTHO1, D8 S1179, HUMFIBRA/FGA, D21S11, D18S51 or AMG loci.
[0117] According to a fifth aspect of the invention we provide the use of an allelic ladder according to the first aspect of the invention and/or an allele according to the fourth aspect of the invention for comparison with a DNA analysis result.
[0118] The analysis may be a DNA profile of a sample. The profile may be based on analysis of one or more loci, in particular including one or more of HUMVWFA31/A, HUMTHO1, D8S1179, HUMFIBRA/FGA, D21S11, D18S51 or AMG. The sample may be from the scene of a crime, associated with the scene of a crime or comprise a bodily fluid sample. The sample may be used to compare two or more individuals, or samples arising therefrom, for instance in paternity and/or maternity analysis.
[0119] According to a sixth aspect of the invention we provide a method of producing an allelic ladder or mixture thereof by subjecting the ladders of the first, second or fourth aspects of the invention to PCR.
BRIEF DESCRIPTION OF THE DRAWINGS
[0120] The invention will now be described, by way of example only, and with reference to the accompanying figure in which:
[0121] FIGS. 1a and 1b illustrates the locus, allele designation and size for an embodiment of the invention;
[0122] FIG. 2a shows an electrophoretogram of the allelic ladder for Amelogenin (AMG);
[0123] FIG. 2b shows an electrophoretogram of the allelic ladder for HUMVWFA31/A;
[0124] FIG. 2c shows an electrophoretogram of the allelic ladder for HUMTHO1;
[0125] FIG. 2d shows an electrophoretogram of the allelic ladder for D8S1179;
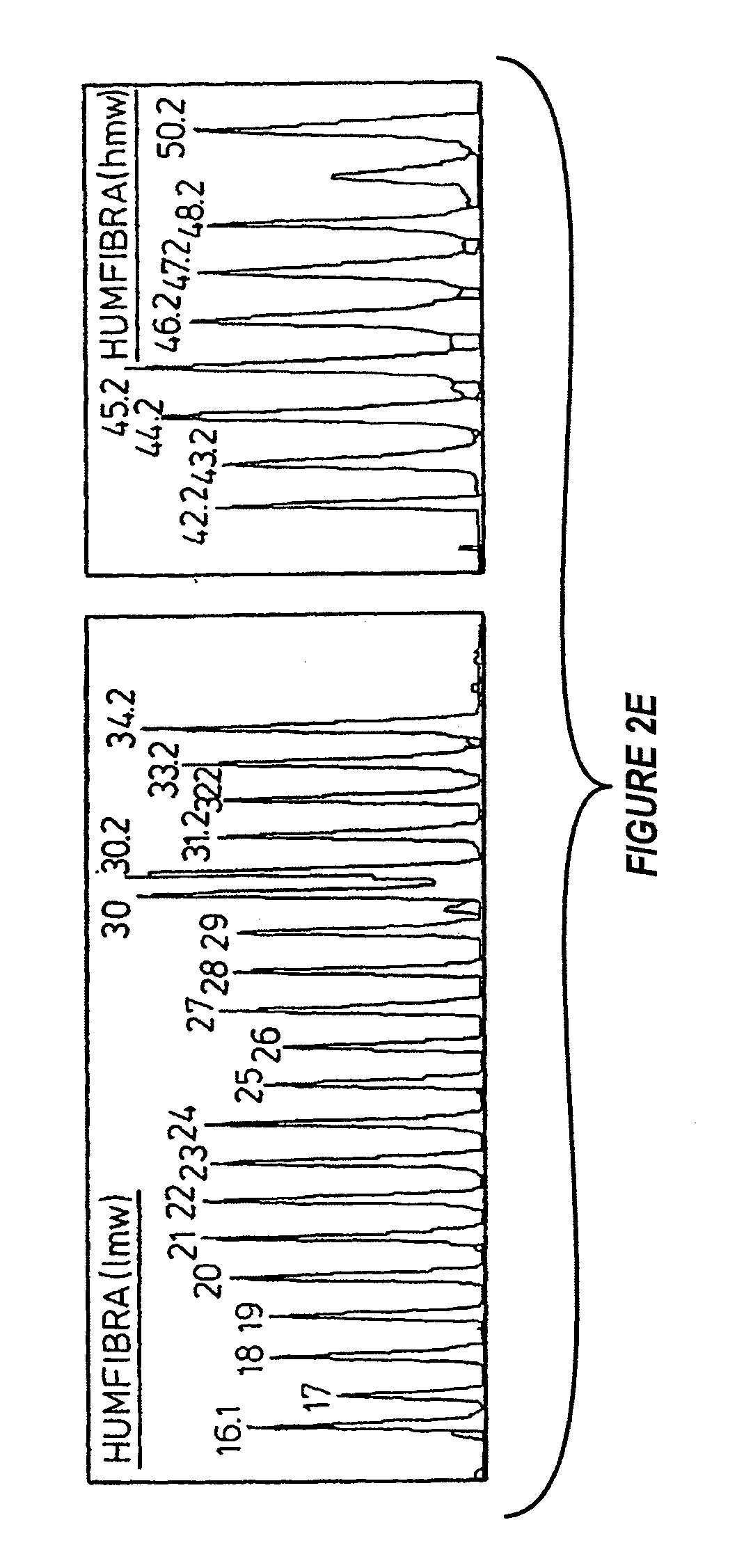
[0126] FIG. 2e shows an electrophoretogram of the allelic ladder for HUMFIBRA, low and high molecular weights;
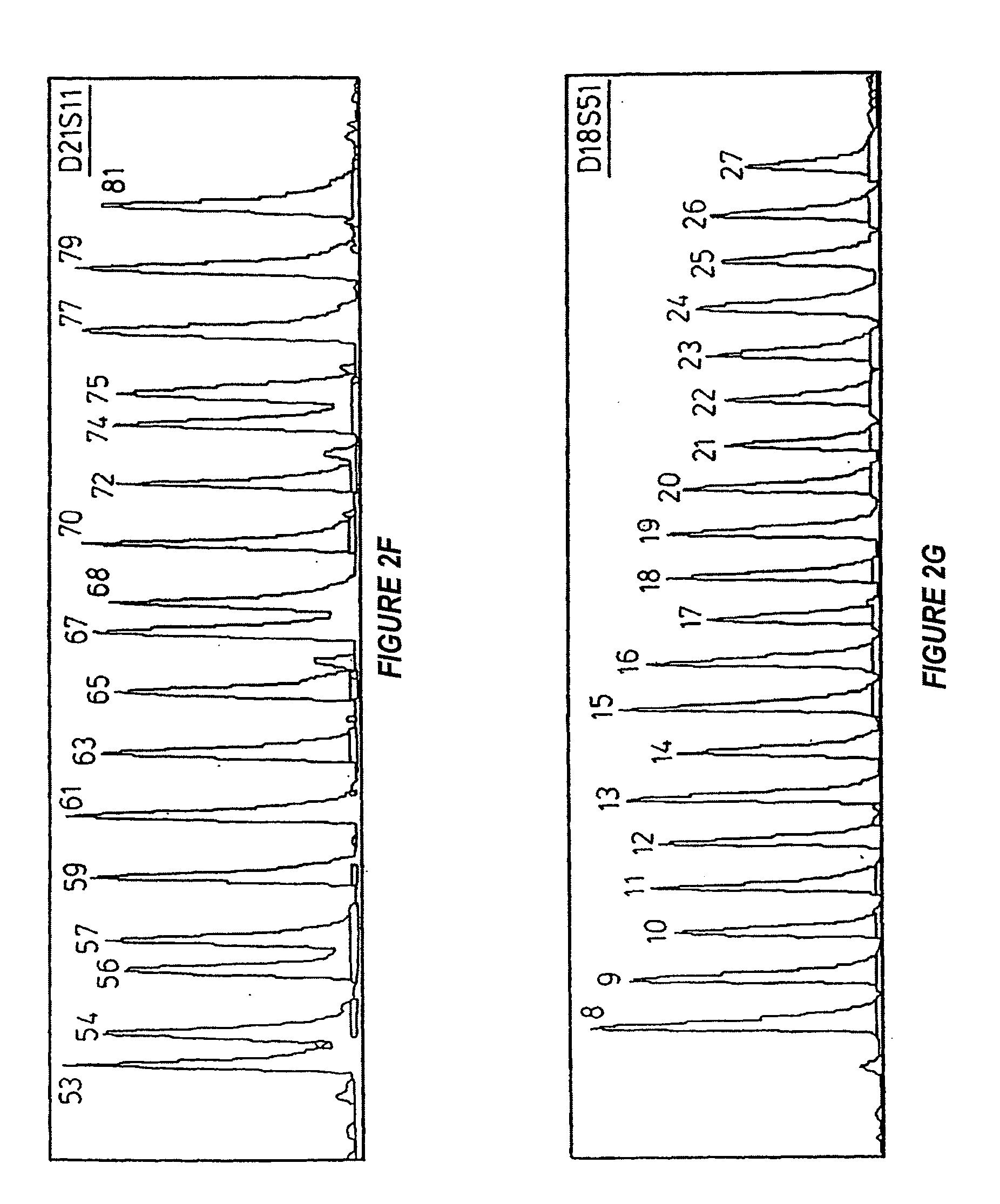
[0127] FIG. 2f shows an electrophoretogram of the allelic ladder for D21S11;
[0128] FIG. 2g shows an electrophoretogram of the allelic ladder for D18S51;
[0129] FIG. 3a shows the sequence of selected alleles forming the HUMVWFA31/A ladder (SEQ ID NOS 1-2 and 39, respectively, in order of appearance);
[0130] FIG. 3b shows the sequence of selected alleles forming the HUMTHO1 ladder (SEQ ID NO: 4);
[0131] FIG. 3c shows the sequence of selected alleles forming the D8S1179 ladder (SEQ ID NOS 5-6, respectively, in order of appearance);
[0132] FIG. 3d shows the sequence of selected alleles forming the HUMFIBRA ladder (SEQ ID NOS 7-18, respectively, in order of appearance);
[0133] FIG. 3e shows the sequence of selected alleles forming the D21S11 ladder (SEQ ID NOS 19-35, respectively, in order of appearance); and
[0134] FIG. 3f shows the sequence of selected alleles forming the D18S51 ladder (SEQ ID NO: 36).
DETAILED DESCRIPTION OF THE PREFERRED EMBODIMENTS
[0135] An allelic ladder mixture illustrative of the present invention is provided for loci HUMTHO1, D21S11, D8S1179, HUMVWFA31/A, HUMFIBRA/FGA and amelogenin sex test. The loci nomenclature is standard, corresponding to that used in the GENEBANK database.
[0136] The ladder mixture includes a significant number of alleles for each locus so as to provide a base line for comparison across a wide range. The loci, allelic designation and base pair sizes for the mixture are shown in FIGS. 1a and 1b. The nomenclature for the loci is discussed in Gill et al. 1996 Int. Journal Leg. Med. 109 14-22.
[0137] The allelic ladder mixture was presented in PARR buffer (containing Tris and 1.5 mM Mg ions at pH8.0) obtained from Cambio, primers obtained from Oswell and Taq polymerase from Perkin Elmer.
[0138] Electrophoretograms for the allelic ladders are shown in FIGS. 2a to 2g with the allelic number designations shown.
[0139] FIGS. 3a to 3f show the sequences for the alleles identified in FIGS. 2a to 2g.
[0140] The allelic ladder mixture discussed above was produced according to the following techniques. Buccal swabs and/or bloodstains were used as the sample sources. The genomic DNA was extracted using the chelex procedure described by Walsh et al. 1991 Bio. Techniques 1 91-98.
[0141] The recovered DNA was quantified by dot hybridisation using a higher primate specific probe, as disclosed in Walsh et al. 1992 Nucleic Acids Res. 20 5061-5065.
[0142] Each sample was then amplified according to the conditions set out below in Table 1 with unlabelled oligonucleotide primers, the sequences for which are disclosed in Urquhart et al. 1995 Bio Techniques 18 116-121 and Oldroyd et al. 1995 Electrophoresis 16 334-337.
TABLE-US-00008 TABLE 1 D18 95.degree. C. for 60 seconds D21 94.degree. C. for 30 seconds 60.degree. C. for 60 seconds 58.degree. C. for 60 seconds 72.degree. C. for 60 seconds 72.degree. C. for 30 seconds Method: 28 cycles + 72.degree. C. Method: 26 cycles + 72.degree. C. for 10 minutes then hold at for 10 minutes then hold at 4.degree. C. 4.degree. C. D8 94.degree. C. for 30 seconds TH01 94.degree. C. for 45 seconds 60.degree. C. for 60 seconds and 60.degree. C. for 60 seconds 72.degree. C. for 60 seconds VWA 72.degree. C. for 60 seconds Method: 30 cycles + 72.degree. C. Method: 28 cycles + 72.degree. C. for 10 minutes then hold at for 10 minutes then hold at 4.degree. C. 4.degree. C. FGA 93.degree. C. for 60 seconds Amelo 93.degree. C. for 30 seconds 60.degree. C. for 60 seconds 58.degree. C. for 75 seconds 72.degree. C. for 60 seconds 72.degree. C. for 15 seconds Method: 30 cycles + 72.degree. C. Method: 30 cycles + 72.degree. C. for 10 minutes then hold at for 10 minutes then hold at 4.degree. C. 4.degree. C.
[0143] Individual alleles were then isolated and sequence analysis was carried out according to the methods of Barber et al. 1996 Int. Journal Leg. Med. 108 180-185 and Barber and Parkin 1996 Int. Journal Leg. Med. 109 62-65. Both DNA strands of each allele reported were sequenced and the sequences provided in FIGS. 3a to 3g are the consensus results for this.
[0144] The illustrations of the alleles provided in FIGS. 3a to 3g follow the nomenclature recommended by the DNA commission of the International Society of Forensic Haemogenetics 1994 Int. Journal Leg. Med. 107 159-160 where the complete number of tandem repeats observed are designated by the digit. The longhand version of these sequences is provided at the end of the specific description.
[0145] To prepare the ladder cocktail amplification of the alleles is necessary. This process was performed by amplifying the purified single alleles described above using a labelled primer in each case. For the locus HUMFIBRA/FGA the ladder was produced from two separate mixes, discussed in more detail below. The primers used are disclosed in Urquhart et al. 1995 Bio Techniques 18 116-121 and Oldroyd et al. 1995 Electrophoresis 16 334-337 and were employed according to the conditions set out above in Table 1.
[0146] The singleplexs produced in this way were analysed on an Applied Biosystems 377 automated sequencer to confirm the sequences. The sequences obtained from the profiling system are one base longer than those determined form the DNA sequencing technique initially discussed above. This is due to the ability of DNA polymerase from Thermus aquaticus to catalyse a non-template mediated addition of a deoxyribonucleotide to the 3' hydroxyl of PCR products. This is generally known as the "n+1" product and can be generated in preference to the "n" product. The results reported here, however, refer to the "n" product rather than the "n+1" product for which the labelled primer PCR conditions have been optimised to produce.
[0147] The products of the amplification process for each locus were then diluted, mixed with one another and reanalysed to produce a single ladder for each loci having even peak heights. An initial level of 1000 Arbitary Units, AU, was increased to 1000-5000 AU to give greater signal strength and volume for the ladder.
[0148] The single ladders produced in this way were then mixed together to give the cocktail discussed above. The proportions of each ladder used are controlled to give balanced peak areas. The cocktail was then validated using Applied Biosystems 373A and Applied Biosystems 377 automated sequencers with Genescan and Genotyper software.
[0149] Allelic ladders according to the invention can be prepared by applying PCR amplification techniques to a pre-existing sample of the allelic ladder mixture. Alternatively the allelic ladders can be constructed from the sequence information provided herein.
[0150] The new ladders disclosed above significantly extends the range of alleles which can be identified in any DNA profiling system.
[0151] The allelic ladder mixture is used as a control sample alongside samples from known or unknown individuals which are then segregated according to size in a gel. Alleles in the sample under test can be designated by the known alleles in the control if they are within 0.5 bases of one another. Alleles falling outside this range are estimated based on their position relative to the ladder.
[0152] Using the standard nomenclature discussed above, the ladder range for each locus, defined by the extreme low molecular weight and extreme high molecular weight alleles are:
TABLE-US-00009 Locus Low MW allele High MW allele HUMVWFA31/A 10 21 HUMTH01 4 13.3 D8S1179 7 19 HUMFIBRA/FGA 16.1 50.2 D21S11 53 81 D18S51 8 27
[0153] The allelic ladders also enable the identification of certain rare and hence highly discriminatory alleles in DNA profiling, thus increasing the profiling systems power.
[0154] For the various locus certain alleles are of particular significance as follows:
[0155] Locus HUMTHO1
[0156] The primers used for this locus were labelled with 6-FAM. The polymorphic region of this locus is based around a tetranucleotide motif repeat, (TCAT).sub.n, where n=4 to 13. Particular alleles provided by the present invention include 4, 9.3, 10 and 13.3. The 9.3 and 13.3 alleles were found to have a deletion of a thiamine nucleotide at either the last base of the 4th repeat unit or the first base of the 5th repeat unit. The 13.3 allele notably possesses a non-consensus tetranucleotide (TCGT) at the 13th repeat.
[0157] Locus D21S11
[0158] The primers for this locus were also labelled with 6-FAM. The allele range extends from 53 to 81 and significantly includes alleles 53, 56, 57, 79 and 81. The polymorphic region of the D21S11 alleles is relatively complex in structure and is based around the tetranucleotide TCTR, where R is A or G (following the ambiguity codes of the Nomenclature Committee of the International Union of Biochemistry), as well as containing invariant hexa-, tri- and di-nucleotides. Both allele 54 and allele 56 deviate from this general structure in that they possess a deletion of a 14 base pair TA(TCTA).sub.3 (SEQ ID NO: 37) unit immediately prior to the invariant TCA tetranucleotide.
Locus D18S51
[0159] Again primers with a 6-FAM label were used. The ladder extends to 20 distinct alleles with particularly significant alleles at 8, 9, 23, 24, 25, 26 and 27. The polymorphic region is based around a simple tetranucleotide repeat motif (AGAA).sub.n (SEQ ID NO: 38), where n is 8 to 27.
[0160] Locus D8S1179
[0161] The primers used for this locus were labelled with TET. The ladder extends from alleles 7 to 19, based on 13 separate alleles. Significant alleles include 7, 15, 18 and 19. Different generalised structures were observed between the upper and lower molecular weight ends of the ladder. In the lower molecular weight area, 161 to 177 base pairs, a simple repeat region based on the tetranucleotide TCTA exists. In the higher weight region, 181 to 201 base pairs, a compound repeat region composed of the tetranucleotide TCTR was found.
Locus HUMVWFA31/A
[0162] HEX labelled primers were used for this locus. The ladder covers alleles between 10 and 21, based on 12 alleles in total. Noteworthy alleles 10, 11 and 12 are included. The polymorphic unit is generally composed of a compound repeat following the pattern (TCTR).sub.n. For the 13 and 14 alleles a non-consensus TCCA tetranucleotide at the 10th and 11th repeats was found.
Locus HUMFIBRA/FGA
[0163] This locus also employed HEX labelled primers. As mentioned above this ladder was constructed in two separate components. A low molecular weight and high molecular weight mix was used to produce the overall ladder. The low molecular weight mix ranges from allele 16.1 to 34.2 and the high molecular weight mix from allele 42.2 to 50.2.
[0164] The low MW mix includes significant alleles 16.1, 28, 30, 30.2, 31.2, 32.2, 33.2 and 34.2. The high MW mix includes noteworthy alleles 42.2, 43.2, 44.2, 45.2, 47.2, 48.2 and 50.2.
[0165] In general the HUMFIBRA/FGA alleles have a polymorphic unit based around the compound repeat YYBY, with the alleles in the upper part of the weight range being more complex in structure than those in the lower part. Within the general framework, allele 16.1 has a T nucleotide addition in the repeat region and allele 27 has a C to T transition in the 19th repeat unit (CTTT to CCTT). The upper MW allele range includes a stutter peak which is 4 base pairs smaller than the 50.2 allele. This artifact corresponds to allele 49.2 which has not currently been determined.
Amelogenin
[0166] Primers for this locus were once again labelled with 6-FAM. The sequence data revealed an X specific product of 105 base pairs and a Y specific product of 111 base pairs.
[0167] HUMTH01 allele sequences
TABLE-US-00010 13.3 (TCAT).sub.4 CAT (TCAT).sub.7 TCAT (SEQ ID NO: 4)
[0168] D21S11 alleles sequences
TABLE-US-00011 (SEQ ID NO: 19) 53 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA(TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.6 TCGTCT (SEQ ID NO: 20) 54 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT (SEQ ID NO: 21) 56 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT (SEQ ID NO: 22) 57 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.8 TCGTCT (SEQ ID NO: 23) 59 (TCTA).sub.5 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.9 TCGTCT (SEQ ID NO: 24) 61 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.10 TCGTCT (SEQ ID NO: 25) 63 (TCTA).sub.4 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT (SEQ ID NO: 26) 65 (TCTA).sub.6 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TCGTCT (SEQ ID NO: 27) 67 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 28) 68 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.11 TA TCTA TCGTCT (SEQ ID NO: 29) 70 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TA TCTA TCGTCT (SEQ ID NO: 30) 72 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TA TCTA TCGTCT (SEQ ID NO: 31) 74 (TCTA).sub.5 (TCTG).sub.6 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.14 TATCTA TCGTCT (SEQ ID NO: 32) 75 (TCTA).sub.10 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 33) 77 (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT (SEQ ID NO: 34) 79 (TCTA).sub.11 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.13 TCGTCT (SEQ ID NO: 35) 81 (TCTA).sub.13 (TCTG).sub.5 (TCTA).sub.3 TA (TCTA).sub.3 TCA (TCTA).sub.2 TCCATA (TCTA).sub.12 TCGTCT
[0169] D18S51 allele sequences
TABLE-US-00012 8 (AGAA).sub.8 (SEQ ID NO: 36)
[0170] D8S1179 allele sequences
TABLE-US-00013 7 (TCTA).sub.8 (SEQ ID NO: 5) 19 (TCTA).sub.2 TCTG (TCTA).sub.16 (SEQ ID NO: 6)
[0171] HUMVWAF31/A allele sequences
TABLE-US-00014 (SEQ ID NO: 1) 10 TCTA TCTG TCTA (TCTG).sub.4 (TCTA).sub.3 (SEQ ID NO: 2) 12 TCTA (TCTG).sub.4 (TCTA).sub.7 (SEQ ID NO: 3) 13 (TCTA).sub.2 (TCTG).sub.4 (TCTA).sub.3 TCCA (TCTA).sub.3
(Note also that the allele has an atypical 3' flanking sequence. The usual sequence is TCCA TCTA T. In this allele the sequence is (TCCA).sub.2T.
[0172] HUMFIBRA(FGA) allele sequences
TABLE-US-00015 (SEQ ID NO: 7) 16.1 (TTTC).sub.3 TTTT TTCT (CTTT).sub.5 T (CTTT).sub.3 CTCC (TTCC).sub.2 (SEQ ID NO: 8) 27 (TTTC).sub.3 TTTT TTCT (CTTT).sub.13 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2 (SEQ ID NO: 9) 30 (TTTC).sub.3 TTTT TTCT (CTTT).sub.16 CCTT (CTTT).sub.5 CTCC (TTCC).sub.2 (SEQ ID NO: 10) 31.2 (TTTC).sub.4 TTTT TT (CTTT).sub.15 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 11) 32.2 (TTTC).sub.4 TTTT TT (CTTT).sub.16 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 12) 33.2 (TTTC).sub.4 TTTT TT (CTTT).sub.17 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 13) 42.2 (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.4 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 14) 43.2 (TTTC).sub.4 TTTT TT (CTTT).sub.8 (CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 15) 44.2 (TTTC).sub.4 TTTT TT (CTTT).sub.11 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 16) 45.2 (TTTC).sub.4 TTTT TT (CTTT).sub.10 CTGT).sub.5 (CTTT).sub.13 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 17) 47.2 (TTTC).sub.4 TTTT TT (CTTT).sub.12 (CTGT).sub.5 (CTTT).sub.14 (CTTC).sub.3 (CTTT).sub.3 CTCC (TTCC).sub.4 (SEQ ID NO: 18) 48.2 (TTTC).sub.4 TTTT TT (CTTT).sub.14 (CTGT).sub.3 (CTTT).sub.14 (CTTC).sub.4 (CTTT).sub.3 CTCC (TTCC).sub.4
[0173] (1) GENERAL INFORMATION:
[0174] (i) APPLICANT: [0175] (A) NAME: Griffiths, Rebecca A. L. [0176] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0177] (C) CITY: Birmingham [0178] (D) STATE: W Midlands [0179] (E) COUNTRY: United Kingdom [0180] (F) POSTAL CODE (ZIP): B5 6QQ [0181] (A) NAME: Barber, Michael D. [0182] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0183] (C) CITY: Birmingham [0184] (D) STATE: W Midlands [0185] (E) COUNTRY: United Kingdom [0186] (F) POSTAL CODE (ZIP): B5 6QQ [0187] (A) NAME: Johnson, Peter E. [0188] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0189] (C) CITY: Birmingham [0190] (D) STATE: W Midlands [0191] (E) COUNTRY: United Kingdom [0192] (F) POSTAL CODE (ZIP): B5 6QQ [0193] (A) NAME: Gillbard, Sharon M. [0194] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0195] (C) CITY: Birmingham [0196] (D) STATE: W Midlands [0197] (E) COUNTRY: United Kingdom [0198] (F) POSTAL CODE (ZIP): B5 6QQ [0199] (A) NAME: Haywood, Marc D. [0200] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0201] (C) CITY: Birmingham [0202] (D) STATE: W Midlands [0203] (E) COUNTRY: United Kingdom [0204] (F) POSTAL CODE (ZIP): B5 6QQ [0205] (A) NAME: Smith, Carolyn D. [0206] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0207] (C) CITY: Birmingham [0208] (D) STATE: W Midlands [0209] (E) COUNTRY: United Kingdom [0210] (F) POSTAL CODE (ZIP): B5 6QQ [0211] (A) NAME: Arnold, Jennifer [0212] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0213] (C) CITY: Birmingham [0214] (D) STATE: W Midlands [0215] (E) COUNTRY: United Kingdom [0216] (F) POSTAL CODE (ZIP): B5 6QQ [0217] (A) NAME: Burke, Trudy [0218] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0219] (C) CITY: Birmingham [0220] (D) STATE: W Midlands [0221] (E) COUNTRY: United Kingdom [0222] (F) POSTAL CODE (ZIP): B5 6QQ [0223] (A) NAME: Urquhart, Andrew J. [0224] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0225] (C) CITY: Birmingham [0226] (D) STATE: W Midlands [0227] (E) COUNTRY: United Kingdom [0228] (F) POSTAL CODE (ZIP): B5 6QQ [0229] (A) NAME: Gill, Peter [0230] (B) STREET: c/o The Forensic Science Service, Priory House, Gooch St. [0231] (C) CITY: Birmingham [0232] (D) STATE: W Midlands [0233] (E) COUNTRY: United Kingdom [0234] (F) POSTAL CODE (ZIP): B5 6QQ
[0235] (ii) TITLE OF INVENTION: Improvements in and relating to forensic identification
[0236] (iii) NUMBER OF SEQUENCES: 36
[0237] (iv) COMPUTER READABLE FORM: [0238] (A) MEDIUM TYPE: Floppy disk [0239] (B) COMPUTER: IBM PC compatible [0240] (C) OPERATING SYSTEM: PC-DOS/MS-DOS [0241] (D) SOFTWARE: PatentIn Release #1.0, Version #1.30 (EPO)
[0242] (vi) PRIOR APPLICATION DATA: [0243] (A) APPLICATION NUMBER: GB 9713597.4 [0244] (B) FILING DATE: 28-JUN-1997
[0245] (2) INFORMATION FOR SEQ ID NO: 1:
[0246] (i) SEQUENCE CHARACTERISTICS: [0247] (A) LENGTH: 40 base pairs [0248] (B) TYPE: nucleic acid [0249] (C) STRANDEDNESS: single [0250] (D) TOPOLOGY: linear
[0251] (ii) MOLECULE TYPE: DNA (genomic)
[0252] (iii) HYPOTHETICAL: NO
[0253] (iv) ANTI-SENSE: NO
[0254] (vi) ORIGINAL SOURCE: [0255] (A) ORGANISM: Homo sapiens [0256] (I) ORGANELLE: Mitochondrion
TABLE-US-00016 [0256] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 1: TCTATCTGTC TATCTGTCTG TCTGTCTGTC TATCTATCTA 40
[0257] (2) INFORMATION FOR SEQ ID NO: 2:
[0258] (i) SEQUENCE CHARACTERISTICS: [0259] (A) LENGTH: 48 base pairs [0260] (B) TYPE: nucleic acid [0261] (C) STRANDEDNESS: single [0262] (D) TOPOLOGY: linear
[0263] (ii) MOLECULE TYPE: DNA (genomic)
[0264] (iii) HYPOTHETICAL: NO
[0265] (iv) ANTI-SENSE: NO
[0266] (vi) ORIGINAL SOURCE: [0267] (A) ORGANISM: Homo sapiens [0268] (I) ORGANELLE: Mitochondrion
TABLE-US-00017 [0268] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 2: TCTATCTGTC TGTCTGTCTG TCTATCTATC TATCTATCTA TCTATCTA 48
[0269] (2) INFORMATION FOR SEQ ID NO: 3:
[0270] (i) SEQUENCE CHARACTERISTICS: [0271] (A) LENGTH: 52 base pairs [0272] (B) TYPE: nucleic acid [0273] (C) STRANDEDNESS: single [0274] (D) TOPOLOGY: linear
[0275] (ii) MOLECULE TYPE: DNA (genomic)
[0276] (iii) HYPOTHETICAL: NO
[0277] (iv) ANTI-SENSE: NO
[0278] (vi) ORIGINAL SOURCE: [0279] (A) ORGANISM: Homo sapiens [0280] (I) ORGANELLE: Mitochondrion
TABLE-US-00018 [0280] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 3: TCTATCTATC TGTCTGTCTG TCTGTCTATC TATCTATCCA TCTATCTATC TA 52
[0281] (2) INFORMATION FOR SEQ ID NO: 4:
[0282] (i) SEQUENCE CHARACTERISTICS: [0283] (A) LENGTH: 55 base pairs [0284] (B) TYPE: nucleic acid [0285] (C) STRANDEDNESS: single [0286] (D) TOPOLOGY: linear
[0287] (ii) MOLECULE TYPE: DNA (genomic)
[0288] (iii) HYPOTHETICAL: NO
[0289] (iv) ANTI-SENSE: NO
[0290] (vi) ORIGINAL SOURCE: [0291] (A) ORGANISM: Homo sapiens [0292] (I) ORGANELLE: Mitochondrion
TABLE-US-00019 [0292] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 4: TCATTCATTC ATTCATCATT CATTCATTCA TTCATTCATT CATTCATTCG TTCAT 55
[0293] (2) INFORMATION FOR SEQ ID NO: 5:
[0294] (i) SEQUENCE CHARACTERISTICS: [0295] (A) LENGTH: 32 base pairs [0296] (B) TYPE: nucleic acid [0297] (C) STRANDEDNESS: single [0298] (D) TOPOLOGY: linear
[0299] (ii) MOLECULE TYPE: DNA (genomic)
[0300] (iii) HYPOTHETICAL: NO
[0301] (iv) ANTI-SENSE: NO
[0302] (vi) ORIGINAL SOURCE: [0303] (A) ORGANISM: Homo sapiens [0304] (I) ORGANELLE: Mitochondrion
TABLE-US-00020 [0304] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 5: TCTATCTATC TATCTATCTA TCTATCTATC TA 32
[0305] (2) INFORMATION FOR SEQ ID NO: 6:
[0306] (i) SEQUENCE CHARACTERISTICS: [0307] (A) LENGTH: 76 base pairs [0308] (B) TYPE: nucleic acid [0309] (C) STRANDEDNESS: single [0310] (D) TOPOLOGY: linear
[0311] (ii) MOLECULE TYPE: DNA (genomic)
[0312] (iii) HYPOTHETICAL: NO
[0313] (iv) ANTI-SENSE: NO
[0314] (vi) ORIGINAL SOURCE: [0315] (A) ORGANISM: Homo sapiens [0316] (I) ORGANELLE: Mitochondrion
TABLE-US-00021 [0316] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 6: TCTATCTATC TGTCTATCTA TCTATCTATC TATCTATCTA TCTATCTATC TATCTATCTA 60 TCTATCTATC TATCTA 76
[0317] (2) INFORMATION FOR SEQ ID NO: 7:
[0318] (i) SEQUENCE CHARACTERISTICS: [0319] (A) LENGTH: 65 base pairs [0320] (B) TYPE: nucleic acid [0321] (C) STRANDEDNESS: single [0322] (D) TOPOLOGY: linear
[0323] (ii) MOLECULE TYPE: DNA (genomic)
[0324] (iii) HYPOTHETICAL: NO
[0325] (iv) ANTI-SENSE: NO
[0326] (vi) ORIGINAL SOURCE: [0327] (A) ORGANISM: Homo sapiens [0328] (I) ORGANELLE: Mitochondrion
TABLE-US-00022 [0328] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 7: TTTCTTTCTT TCTTTTTTCT CTTTCTTTCT TTCTTTCTTT TCTTTCTTTC TTTCTCCTTC 60 CTTCC 65
[0329] (2) INFORMATION FOR SEQ ID NO: 8:
[0330] (i) SEQUENCE CHARACTERISTICS: [0331] (A) LENGTH: 108 base pairs [0332] (B) TYPE: nucleic acid [0333] (C) STRANDEDNESS: single [0334] (D) TOPOLOGY: linear
[0335] (ii) MOLECULE TYPE: DNA (genomic)
[0336] (iii) HYPOTHETICAL: NO
[0337] (iv) ANTI-SENSE: NO
[0338] (vi) ORIGINAL SOURCE: [0339] (A) ORGANISM: Homo sapiens [0340] (I) ORGANELLE: Mitochondrion
TABLE-US-00023 [0340] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 8: TTTCTTTCTT TCTTTTTTCT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT 60 CTTTCTTTCT TTCCTTCTTT CTTTCTTTCT TTCTTTCTCC TTCCTTCC 108
[0341] (2) INFORMATION FOR SEQ ID NO: 9:
[0342] (i) SEQUENCE CHARACTERISTICS: [0343] (A) LENGTH: 120 base pairs [0344] (B) TYPE: nucleic acid [0345] (C) STRANDEDNESS: single [0346] (D) TOPOLOGY: linear
[0347] (ii) MOLECULE TYPE: DNA (genomic)
[0348] (iii) HYPOTHETICAL: NO
[0349] (iv) ANTI-SENSE: NO
[0350] (vi) ORIGINAL SOURCE: [0351] (A) ORGANISM: Homo sapiens [0352] (I) ORGANELLE: Mitochondrion
TABLE-US-00024 [0352] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 9: TTTCTTTCTT TCTTTTTTCT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT 60 CTTTCTTTCT TTCTTTCTTT CTTTCCTTCT TTCTTTCTTT CTTTCTTTCT CCTTCCTTCC 120
[0353] (2) INFORMATION FOR SEQ ID NO: 10:
[0354] (i) SEQUENCE CHARACTERISTICS: [0355] (A) LENGTH: 126 base pairs [0356] (B) TYPE: nucleic acid [0357] (C) STRANDEDNESS: single [0358] (D) TOPOLOGY: linear
[0359] (ii) MOLECULE TYPE: DNA (genomic)
[0360] (iii) HYPOTHETICAL: NO
[0361] (iv) ANTI-SENSE: NO
[0362] (vi) ORIGINAL SOURCE: [0363] (A) ORGANISM: Homo sapiens [0364] (I) ORGANELLE: Mitochondrion
TABLE-US-00025 [0364] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 10: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 60 TTCTTTCTTT CTTTCTTTCT TTCTTCCTTC CTTCCTTTCT TTCTTTCTCC TTCCTTCCTT 120 CCTTCC 126
[0365] (2) INFORMATION FOR SEQ ID NO: 11:
[0366] (i) SEQUENCE CHARACTERISTICS: [0367] (A) LENGTH: 130 base pairs [0368] (B) TYPE: nucleic acid [0369] (C) STRANDEDNESS: single [0370] (D) TOPOLOGY: linear
[0371] (ii) MOLECULE TYPE: DNA (genomic)
[0372] (iii) HYPOTHETICAL: NO
[0373] (iv) ANTI-SENSE: NO
[0374] (vi) ORIGINAL SOURCE: [0375] (A) ORGANISM: Homo sapiens [0376] (I) ORGANELLE: Mitochondrion
TABLE-US-00026 [0376] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 11: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 60 TTCTTTCTTT CTTTCTTTCT TTCTTTCTTC CTTCCTTCCT TTCTTTCTTT CTCCTTCCTT 120 CCTTCCTTCC 130
[0377] (2) INFORMATION FOR SEQ ID NO: 12:
[0378] (i) SEQUENCE CHARACTERISTICS: [0379] (A) LENGTH: 134 base pairs [0380] (B) TYPE: nucleic acid [0381] (C) STRANDEDNESS: single [0382] (D) TOPOLOGY: linear
[0383] (ii) MOLECULE TYPE: DNA (genomic)
[0384] (iii) HYPOTHETICAL: NO
[0385] (iv) ANTI-SENSE: NO
[0386] (vi) ORIGINAL SOURCE: [0387] (A) ORGANISM: Homo sapiens [0388] (I) ORGANELLE: Mitochondrion
TABLE-US-00027 [0388] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 12: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 60 TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTCCTTCCT TCCTTTCTTT CTTTCTCCTT 120 CCTTCCTTCC TTCC 134
[0389] (2) INFORMATION FOR SEQ ID NO: 13:
[0390] (i) SEQUENCE CHARACTERISTICS: [0391] (A) LENGTH: 170 base pairs [0392] (B) TYPE: nucleic acid [0393] (C) STRANDEDNESS: single [0394] (D) TOPOLOGY: linear
[0395] (ii) MOLECULE TYPE: DNA (genomic)
[0396] (iii) HYPOTHETICAL: NO
[0397] (iv) ANTI-SENSE: NO
TABLE-US-00028 (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 13: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTGTCT 60 GTCTGTCTGT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 120 TTCTTCCTTC CTTCCTTCCT TTCTTTCTTT CTCCTTCCTT CCTTCCTTCC 170
[0398] (2) INFORMATION FOR SEQ ID NO: 14:
[0399] (i) SEQUENCE CHARACTERISTICS: [0400] (A) LENGTH: 174 base pairs [0401] (B) TYPE: nucleic acid [0402] (C) STRANDEDNESS: single [0403] (D) TOPOLOGY: linear
[0404] (ii) MOLECULE TYPE: DNA (genomic)
[0405] (iii) HYPOTHETICAL: NO
[0406] (iv) ANTI-SENSE: NO
[0407] (vi) ORIGINAL SOURCE: [0408] (A) ORGANISM: Homo sapiens [0409] (I) ORGANELLE: Mitochondrion
TABLE-US-00029 [0409] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 14: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTGTCT 60 GTCTGTCTGT CTGTCTTTCT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 120 TTCTTTCTTC CTTCCTTCCT TCCTTTCTTT CTTTCTCCTT CCTTCCTTCC TTCC 174
[0410] (2) INFORMATION FOR SEQ ID NO: 15:
[0411] (i) SEQUENCE CHARACTERISTICS: [0412] (A) LENGTH: 178 base pairs [0413] (B) TYPE: nucleic acid [0414] (C) STRANDEDNESS: single [0415] (D) TOPOLOGY: linear
[0416] (ii) MOLECULE TYPE: DNA (genomic)
[0417] (iii) HYPOTHETICAL: NO
[0418] (iv) ANTI-SENSE: NO
[0419] (vi) ORIGINAL SOURCE: [0420] (A) ORGANISM: Homo sapiens [0421] (I) ORGANELLE: Mitochondrion
TABLE-US-00030 [0421] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 15: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 60 TTCTTTCTGT CTGTCTGTCT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 120 TTCTTTCTTT CTTTCTTCCT TCCTTCCTTT CTTTCTTTCT CCTTCCTTCC TTCCTTCC 178
[0422] (2) INFORMATION FOR SEQ ID NO: 16:
[0423] (i) SEQUENCE CHARACTERISTICS: [0424] (A) LENGTH: 182 base pairs [0425] (B) TYPE: nucleic acid [0426] (C) STRANDEDNESS: single [0427] (D) TOPOLOGY: linear
[0428] (ii) MOLECULE TYPE: DNA (genomic)
[0429] (iii) HYPOTHETICAL: NO
[0430] (iv) ANTI-SENSE: NO
[0431] (vi) ORIGINAL SOURCE: [0432] (A) ORGANISM: Homo sapiens [0433] (I) ORGANELLE: Mitochondrion
TABLE-US-00031 [0433] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 16: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 60 TTCTGTCTGT CTGTCTGTCT GTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 120 TTCTTTCTTT CTTTCTTCCT TCCTTCCTTC CTTTCTTTCT TTCTCCTTCC TTCCTTCCTT 180 CC 182
[0434] (2) INFORMATION FOR SEQ ID NO: 17:
[0435] (i) SEQUENCE CHARACTERISTICS: [0436] (A) LENGTH: 190 base pairs [0437] (B) TYPE: nucleic acid [0438] (C) STRANDEDNESS: single [0439] (D) TOPOLOGY: linear
[0440] (ii) MOLECULE TYPE: DNA (genomic)
[0441] (iii) HYPOTHETICAL: NO
[0442] (iv) ANTI-SENSE: NO
[0443] (vi) ORIGINAL SOURCE: [0444] (A) ORGANISM: Homo sapiens [0445] (I) ORGANELLE: Mitochondrion
TABLE-US-00032 [0445] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 17: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 60 TTCTTTCTTT CTGTCTGTCT GTCTGTCTGT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 120 TTCTTTCTTT CTTTCTTTCT TTCTTTCTTC CTTCCTTCCT TTCTTTCTTT CTCCTTCCTT 180 CCTTCCTTCC 190
[0446] (2) INFORMATION FOR SEQ ID NO: 18:
[0447] (i) SEQUENCE CHARACTERISTICS: [0448] (A) LENGTH: 194 base pairs [0449] (B) TYPE: nucleic acid [0450] (C) STRANDEDNESS: single [0451] (D) TOPOLOGY: linear
[0452] (ii) MOLECULE TYPE: DNA (genomic)
[0453] (iii) HYPOTHETICAL: NO
[0454] (iv) ANTI-SENSE: NO
[0455] (vi) ORIGINAL SOURCE: [0456] (A) ORGANISM: Homo sapiens [0457] (I) ORGANELLE: Mitochondrion
TABLE-US-00033 [0457] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 18: TTTCTTTCTT TCTTTCTTTT TTCTTTCTTT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 60 TTCTTTCTTT CTTTCTTTCT GTCTGTCTGT CTTTCTTTCT TTCTTTCTTT CTTTCTTTCT 120 TTCTTTCTTT CTTTCTTTCT TTCTTTCTTC CTTCCTTCCT TCCTTTCTTT CTTTCTCCTT 180 CCTTCCTTCC TTCC 194
[0458] (2) INFORMATION FOR SEQ ID NO: 19:
[0459] (i) SEQUENCE CHARACTERISTICS: [0460] (A) LENGTH: 113 base pairs [0461] (B) TYPE: nucleic acid [0462] (C) STRANDEDNESS: single [0463] (D) TOPOLOGY: linear
[0464] (ii) MOLECULE TYPE: DNA (genomic)
[0465] (iii) HYPOTHETICAL: NO
[0466] (iv) ANTI-SENSE: NO
[0467] (vi) ORIGINAL SOURCE: [0468] (A) ORGANISM: Homo sapiens [0469] (I) ORGANELLE: Mitochondrion
TABLE-US-00034 [0469] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 19: TCTATCTATC TATCTATCTG TCTGTCTGTC TGTCTGTCTG TCTATCTATC TATATCTATC 60 TATCTATCAT CTATCTATCC ATATCTATCT ATCTATCTAT CTATCTATCG TCT 113
[0470] (2) INFORMATION FOR SEQ ID NO: 20:
[0471] (i) SEQUENCE CHARACTERISTICS: [0472] (A) LENGTH: 115 base pairs [0473] (B) TYPE: nucleic acid [0474] (C) STRANDEDNESS: single [0475] (D) TOPOLOGY: linear
[0476] (ii) MOLECULE TYPE: DNA (genomic)
[0477] (iii) HYPOTHETICAL: NO
[0478] (iv) ANTI-SENSE: NO
[0479] (vi) ORIGINAL SOURCE: [0480] (A) ORGANISM: Homo sapiens [0481] (I) ORGANELLE: Mitochondrion
TABLE-US-00035 [0481] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 20: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTGTCTATC TATCTATCAT 60 CTATCTATCC ATATCTATCT ATCTATCTAT CTATCTATCT ATCTATCTAT CGTCT 115
[0482] (2) INFORMATION FOR SEQ ID NO: 21:
[0483] (i) SEQUENCE CHARACTERISTICS: [0484] (A) LENGTH: 119 base pairs [0485] (B) TYPE: nucleic acid [0486] (C) STRANDEDNESS: single [0487] (D) TOPOLOGY: linear
[0488] (ii) MOLECULE TYPE: DNA (genomic)
[0489] (iii) HYPOTHETICAL: NO
[0490] (iv) ANTI-SENSE: NO
[0491] (vi) ORIGINAL SOURCE: [0492] (A) ORGANISM: Homo sapiens [0493] (I) ORGANELLE: Mitochondrion
TABLE-US-00036 [0493] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 21: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTGTCTATC TATCTATCAT 60 CTATCTATCC ATATCTATCT ATCTATCTAT CTATCTATCT ATCTATCTAT CTATCGTCT 119
[0494] (2) INFORMATION FOR SEQ ID NO: 22:
[0495] (i) SEQUENCE CHARACTERISTICS: [0496] (A) LENGTH: 121 base pairs [0497] (B) TYPE: nucleic acid [0498] (C) STRANDEDNESS: single [0499] (D) TOPOLOGY: linear
[0500] (ii) MOLECULE TYPE: DNA (genomic)
[0501] (iii) HYPOTHETICAL: NO
[0502] (iv) ANTI-SENSE: NO
[0503] (vi) ORIGINAL SOURCE: [0504] (A) ORGANISM: Homo sapiens [0505] (I) ORGANELLE: Mitochondrion
TABLE-US-00037 [0505] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 22: TCTATCTATC TATCTATCTG TCTGTCTGTC TGTCTGTCTG TCTATCTATC TATATCTATC 60 TATCTATCAT CTATCTATCC ATATCTATCT ATCTATCTAT CTATCTATCT ATCTATCGTC 120 T 121
[0506] (2) INFORMATION FOR SEQ ID NO: 23:
[0507] (i) SEQUENCE CHARACTERISTICS: [0508] (A) LENGTH: 125 base pairs [0509] (B) TYPE: nucleic acid [0510] (C) STRANDEDNESS: single [0511] (D) TOPOLOGY: linear
[0512] (ii) MOLECULE TYPE: DNA (genomic)
[0513] (iii) HYPOTHETICAL: NO
[0514] (iv) ANTI-SENSE: NO
[0515] (vi) ORIGINAL SOURCE: [0516] (A) ORGANISM: Homo sapiens [0517] (I) ORGANELLE: Mitochondrion
TABLE-US-00038 [0517] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 23: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTATCTATC TATATCTATC 60 TATCTATCAT CTATCTATCC ATATCTATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CGTCT 125
[0518] (2) INFORMATION FOR SEQ ID NO: 24:
[0519] (i) SEQUENCE CHARACTERISTICS: [0520] (A) LENGTH: 129 base pairs [0521] (B) TYPE: nucleic acid [0522] (C) STRANDEDNESS: single [0523] (D) TOPOLOGY: linear
[0524] (ii) MOLECULE TYPE: DNA (genomic)
[0525] (iii) HYPOTHETICAL: NO
[0526] (iv) ANTI-SENSE: NO
[0527] (vi) ORIGINAL SOURCE: [0528] (A) ORGANISM: Homo sapiens [0529] (I) ORGANELLE: Mitochondrion
TABLE-US-00039 [0529] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 24: TCTATCTATC TATCTATCTG TCTGTCTGTC TGTCTGTCTG TCTATCTATC TATATCTATC 60 TATCTATCAT CTATCTATCC ATATCTATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCGTCT 129
[0530] (2) INFORMATION FOR SEQ ID NO: 25:
[0531] (i) SEQUENCE CHARACTERISTICS: [0532] (A) LENGTH: 133 base pairs [0533] (B) TYPE: nucleic acid [0534] (C) STRANDEDNESS: single [0535] (D) TOPOLOGY: linear
[0536] (ii) MOLECULE TYPE: DNA (genomic)
[0537] (iii) HYPOTHETICAL: NO
[0538] (iv) ANTI-SENSE: NO
[0539] (vi) ORIGINAL SOURCE: [0540] (A) ORGANISM: Homo sapiens [0541] (I) ORGANELLE: Mitochondrion
TABLE-US-00040 [0541] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 25: TCTATCTATC TATCTATCTG TCTGTCTGTC TGTCTGTCTG TCTATCTATC TATATCTATC 60 TATCTATCAT CTATCTATCC ATATCTATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCTATCG TCT 133
[0542] (2) INFORMATION FOR SEQ ID NO: 26:
[0543] (i) SEQUENCE CHARACTERISTICS: [0544] (A) LENGTH: 137 base pairs [0545] (B) TYPE: nucleic acid [0546] (C) STRANDEDNESS: single [0547] (D) TOPOLOGY: linear
[0548] (ii) MOLECULE TYPE: DNA (genomic)
[0549] (iii) HYPOTHETICAL: NO
[0550] (iv) ANTI-SENSE: NO
[0551] (vi) ORIGINAL SOURCE: [0552] (A) ORGANISM: Homo sapiens [0553] (I) ORGANELLE: Mitochondrion
TABLE-US-00041 [0553] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 26: TCTATCTATC TATCTATCTA TCTATCTGTC TGTCTGTCTG TCTGTCTATC TATCTATATC 60 TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCTATCT ATCGTCT 137
[0554] (2) INFORMATION FOR SEQ ID NO: 27:
[0555] (i) SEQUENCE CHARACTERISTICS: [0556] (A) LENGTH: 141 base pairs [0557] (B) TYPE: nucleic acid [0558] (C) STRANDEDNESS: single [0559] (D) TOPOLOGY: linear
[0560] (ii) MOLECULE TYPE: DNA (genomic)
[0561] (iii) HYPOTHETICAL: NO
[0562] (iv) ANTI-SENSE: NO
[0563] (vi) ORIGINAL SOURCE: [0564] (A) ORGANISM: Homo sapiens [0565] (I) ORGANELLE: Mitochondrion
TABLE-US-00042 [0565] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 27: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTGTCTATC TATCTATATC 60 TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCTATCT ATCTATCGTC T 141
[0566] (2) INFORMATION FOR SEQ ID NO: 28:
[0567] (i) SEQUENCE CHARACTERISTICS: [0568] (A) LENGTH: 143 base pairs [0569] (B) TYPE: nucleic acid [0570] (C) STRANDEDNESS: single [0571] (D) TOPOLOGY: linear
[0572] (ii) MOLECULE TYPE: DNA (genomic)
[0573] (iii) HYPOTHETICAL: NO
[0574] (iv) ANTI-SENSE: NO
[0575] (vi) ORIGINAL SOURCE: [0576] (A) ORGANISM: Homo sapiens [0577] (I) ORGANELLE: Mitochondrion
TABLE-US-00043 [0577] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 28: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTGTCTATC TATCTATATC 60 TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCTATCT ATATCTATCG TCT 143
[0578] (2) INFORMATION FOR SEQ ID NO: 29:
[0579] (i) SEQUENCE CHARACTERISTICS: [0580] (A) LENGTH: 147 base pairs [0581] (B) TYPE: nucleic acid [0582] (C) STRANDEDNESS: single [0583] (D) TOPOLOGY: linear
[0584] (ii) MOLECULE TYPE: DNA (genomic)
[0585] (iii) HYPOTHETICAL: NO
[0586] (iv) ANTI-SENSE: NO
[0587] (vi) ORIGINAL SOURCE: [0588] (A) ORGANISM: Homo sapiens [0589] (I) ORGANELLE: Mitochondrion
TABLE-US-00044 [0589] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 29: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTGTCTATC TATCTATATC 60 TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCTATCT ATCTATATCT ATCGTCT 147
[0590] (2) INFORMATION FOR SEQ ID NO: 30:
[0591] (i) SEQUENCE CHARACTERISTICS: [0592] (A) LENGTH: 151 base pairs [0593] (B) TYPE: nucleic acid [0594] (C) STRANDEDNESS: single [0595] (D) TOPOLOGY: linear
[0596] (ii) MOLECULE TYPE: DNA (genomic)
[0597] (iii) HYPOTHETICAL: NO
[0598] (iv) ANTI-SENSE: NO
[0599] (vi) ORIGINAL SOURCE: [0600] (A) ORGANISM: Homo sapiens [0601] (I) ORGANELLE: Mitochondrion
TABLE-US-00045 [0601] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 30: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTGTCTATC TATCTATATC 60 TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCTATCT ATCTATCTAT ATCTATCGTC T 151
[0602] (2) INFORMATION FOR SEQ ID NO: 31:
[0603] (i) SEQUENCE CHARACTERISTICS: [0604] (A) LENGTH: 155 base pairs [0605] (B) TYPE: nucleic acid [0606] (C) STRANDEDNESS: single [0607] (D) TOPOLOGY: linear
[0608] (ii) MOLECULE TYPE: DNA (genomic)
[0609] (iii) HYPOTHETICAL: NO
[0610] (iv) ANTI-SENSE: NO
[0611] (vi) ORIGINAL SOURCE: [0612] (A) ORGANISM: Homo sapiens [0613] (I) ORGANELLE: Mitochondrion
TABLE-US-00046 [0613] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 31: TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG TCTGTCTATC TATCTATATC 60 TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT CTATCTATCT ATCTATCTAT 120 CTATCTATCT ATCTATCTAT CTATATCTAT CGTCT 155
[0614] (2) INFORMATION FOR SEQ ID NO: 32:
[0615] (i) SEQUENCE CHARACTERISTICS: [0616] (A) LENGTH: 157 base pairs [0617] (B) TYPE: nucleic acid [0618] (C) STRANDEDNESS: single [0619] (D) TOPOLOGY: linear
[0620] (ii) MOLECULE TYPE: DNA (genomic)
[0621] (iii) HYPOTHETICAL: NO
[0622] (iv) ANTI-SENSE: NO
[0623] (vi) ORIGINAL SOURCE: [0624] (A) ORGANISM: Homo sapiens [0625] (I) ORGANELLE: Mitochondrion
TABLE-US-00047 [0625] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 32: TCTATCTATC TATCTATCTA TCTATCTATC TATCTATCTA TCTGTCTGTC TGTCTGTCTG 60 TCTATCTATC TATATCTATC TATCTATCAT CTATCTATCC ATATCTATCT ATCTATCTAT 120 CTATCTATCT ATCTATCTAT CTATCTATCT ATCGTCT 157
[0626] (2) INFORMATION FOR SEQ ID NO: 33:
[0627] (i) SEQUENCE CHARACTERISTICS: [0628] (A) LENGTH: 161 base pairs [0629] (B) TYPE: nucleic acid [0630] (C) STRANDEDNESS: single [0631] (D) TOPOLOGY: linear
[0632] (ii) MOLECULE TYPE: DNA (genomic)
[0633] (iii) HYPOTHETICAL: NO
[0634] (iv) ANTI-SENSE: NO
[0635] (vi) ORIGINAL SOURCE: [0636] (A) ORGANISM: Homo sapiens [0637] (I) ORGANELLE: Mitochondrion
TABLE-US-00048 [0637] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 33: TCTATCTATC TATCTATCTA TCTATCTATC TATCTATCTA TCTATCTGTC TGTCTGTCTG 60 TCTGTCTATC TATCTATATC TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT 120 CTATCTATCT ATCTATCTAT CTATCTATCT ATCTATCGTC T 161
[0638] (2) INFORMATION FOR SEQ ID NO: 34:
[0639] (i) SEQUENCE CHARACTERISTICS: [0640] (A) LENGTH: 165 base pairs [0641] (B) TYPE: nucleic acid [0642] (C) STRANDEDNESS: single [0643] (D) TOPOLOGY: linear
[0644] (ii) MOLECULE TYPE: DNA (genomic)
[0645] (iii) HYPOTHETICAL: NO
[0646] (iv) ANTI-SENSE: NO
[0647] (vi) ORIGINAL SOURCE: [0648] (A) ORGANISM: Homo sapiens [0649] (I) ORGANELLE: Mitochondrion
TABLE-US-00049 [0649] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 34: TCTATCTATC TATCTATCTA TCTATCTATC TATCTATCTA TCTATCTGTC TGTCTGTCTG 60 TCTGTCTATC TATCTATATC TATCTATCTA TCATCTATCT ATCCATATCT ATCTATCTAT 120 CTATCTATCT ATCTATCTAT CTATCTATCT ATCTATCTAT CGTCT 165
[0650] (2) INFORMATION FOR SEQ ID NO: 35:
[0651] (i) SEQUENCE CHARACTERISTICS: [0652] (A) LENGTH: 169 base pairs [0653] (B) TYPE: nucleic acid [0654] (C) STRANDEDNESS: single [0655] (D) TOPOLOGY: linear
[0656] (ii) MOLECULE TYPE: DNA (genomic)
[0657] (iii) HYPOTHETICAL: NO
[0658] (iv) ANTI-SENSE: NO
[0659] (vi) ORIGINAL SOURCE: [0660] (A) ORGANISM: Homo sapiens [0661] (I) ORGANELLE: Mitochondrion
TABLE-US-00050 [0661] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 35: TCTATCTATC TATCTATCTA TCTATCTATC TATCTATCTA TCTATCTATC TATCTGTCTG 60 TCTGTCTGTC TGTCTATCTA TCTATATCTA TCTATCTATC ATCTATCTAT CCATATCTAT 120 CTATCTATCT ATCTATCTAT CTATCTATCT ATCTATCTAT CTATCGTCT 169
[0662] (2) INFORMATION FOR SEQ ID NO: 36:
[0663] (i) SEQUENCE CHARACTERISTICS: [0664] (A) LENGTH: 32 base pairs [0665] (B) TYPE: nucleic acid [0666] (C) STRANDEDNESS: single [0667] (D) TOPOLOGY: linear
[0668] (ii) MOLECULE TYPE: DNA (genomic)
[0669] (iii) HYPOTHETICAL: NO
[0670] (iv) ANTI-SENSE: NO
[0671] (vi) ORIGINAL SOURCE: [0672] (A) ORGANISM: Homo sapiens [0673] (I) ORGANELLE: Mitochondrion
TABLE-US-00051 [0673] (xi) SEQUENCE DESCRIPTION: SEQ ID NO: 36: AGAAAGAAAG AAAGAAAGAA AGAAAGAAAG AA 32
[0674] The present invention may be embodied in other specific forms without departing from its spirit or essential characteristics. The described embodiments are to be considered in all respects only as illustrative and not restrictive. The scope of the invention is, therefore, indicated by the appended claims rather than by the foregoing description. All changes which come within the meaning and range of equivalency of the claims are to be embraced within their scope.
Sequence CWU 1
1
39140DNAHomo sapiens 1tctatctgtc tatctgtctg tctgtctgtc tatctatcta
40248DNAHomo sapiens 2tctatctgtc tgtctgtctg tctatctatc tatctatcta
tctatcta 48352DNAHomo sapiens 3tctatctatc tgtctgtctg tctgtctatc
tatctatcca tctatctatc ta 52455DNAHomo sapiens 4tcattcattc
attcatcatt cattcattca ttcattcatt cattcattcg ttcat 55532DNAHomo
sapiens 5tctatctatc tatctatcta tctatctatc ta 32676DNAHomo sapiens
6tctatctatc tgtctatcta tctatctatc tatctatcta tctatctatc tatctatcta
60tctatctatc tatcta 76765DNAHomo sapiens 7tttctttctt tcttttttct
ctttctttct ttctttcttt tctttctttc tttctccttc 60cttcc 658108DNAHomo
sapiens 8tttctttctt tcttttttct ctttctttct ttctttcttt ctttctttct
ttctttcttt 60ctttctttct ttccttcttt ctttctttct ttctttctcc ttccttcc
1089120DNAHomo sapiens 9tttctttctt tcttttttct ctttctttct ttctttcttt
ctttctttct ttctttcttt 60ctttctttct ttctttcttt ctttccttct ttctttcttt
ctttctttct ccttccttcc 12010126DNAHomo sapiens 10tttctttctt
tctttctttt ttctttcttt ctttctttct ttctttcttt ctttctttct 60ttctttcttt
ctttctttct ttcttccttc cttcctttct ttctttctcc ttccttcctt 120ccttcc
12611130DNAHomo sapiens 11tttctttctt tctttctttt ttctttcttt
ctttctttct ttctttcttt ctttctttct 60ttctttcttt ctttctttct ttctttcttc
cttccttcct ttctttcttt ctccttcctt 120ccttccttcc 13012134DNAHomo
sapiens 12tttctttctt tctttctttt ttctttcttt ctttctttct ttctttcttt
ctttctttct 60ttctttcttt ctttctttct ttctttcttt cttccttcct tcctttcttt
ctttctcctt 120ccttccttcc ttcc 13413170DNAHomo sapiens 13tttctttctt
tctttctttt ttctttcttt ctttctttct ttctttcttt ctttctgtct 60gtctgtctgt
ctttctttct ttctttcttt ctttctttct ttctttcttt ctttctttct
120ttcttccttc cttccttcct ttctttcttt ctccttcctt ccttccttcc
17014174DNAHomo sapiens 14tttctttctt tctttctttt ttctttcttt
ctttctttct ttctttcttt ctttctgtct 60gtctgtctgt ctgtctttct ttctttcttt
ctttctttct ttctttcttt ctttctttct 120ttctttcttc cttccttcct
tcctttcttt ctttctcctt ccttccttcc ttcc 17415178DNAHomo sapiens
15tttctttctt tctttctttt ttctttcttt ctttctttct ttctttcttt ctttctttct
60ttctttctgt ctgtctgtct ttctttcttt ctttctttct ttctttcttt ctttctttct
120ttctttcttt ctttcttcct tccttccttt ctttctttct ccttccttcc ttccttcc
17816182DNAHomo sapiens 16tttctttctt tctttctttt ttctttcttt
ctttctttct ttctttcttt ctttctttct 60ttctgtctgt ctgtctgtct gtctttcttt
ctttctttct ttctttcttt ctttctttct 120ttctttcttt ctttcttcct
tccttccttc ctttctttct ttctccttcc ttccttcctt 180cc 18217190DNAHomo
sapiens 17tttctttctt tctttctttt ttctttcttt ctttctttct ttctttcttt
ctttctttct 60ttctttcttt ctgtctgtct gtctgtctgt ctttctttct ttctttcttt
ctttctttct 120ttctttcttt ctttctttct ttctttcttc cttccttcct
ttctttcttt ctccttcctt 180ccttccttcc 19018194DNAHomo sapiens
18tttctttctt tctttctttt ttctttcttt ctttctttct ttctttcttt ctttctttct
60ttctttcttt ctttctttct gtctgtctgt ctttctttct ttctttcttt ctttctttct
120ttctttcttt ctttctttct ttctttcttc cttccttcct tcctttcttt
ctttctcctt 180ccttccttcc ttcc 19419113DNAHomo sapiens 19tctatctatc
tatctatctg tctgtctgtc tgtctgtctg tctatctatc tatatctatc 60tatctatcat
ctatctatcc atatctatct atctatctat ctatctatcg tct 11320115DNAHomo
sapiens 20tctatctatc tatctatcta tctgtctgtc tgtctgtctg tctgtctatc
tatctatcat 60ctatctatcc atatctatct atctatctat ctatctatct atctatctat
cgtct 11521119DNAHomo sapiens 21tctatctatc tatctatcta tctgtctgtc
tgtctgtctg tctgtctatc tatctatcat 60ctatctatcc atatctatct atctatctat
ctatctatct atctatctat ctatcgtct 11922121DNAHomo sapiens
22tctatctatc tatctatctg tctgtctgtc tgtctgtctg tctatctatc tatatctatc
60tatctatcat ctatctatcc atatctatct atctatctat ctatctatct atctatcgtc
120t 12123125DNAHomo sapiens 23tctatctatc tatctatcta tctgtctgtc
tgtctgtctg tctatctatc tatatctatc 60tatctatcat ctatctatcc atatctatct
atctatctat ctatctatct atctatctat 120cgtct 12524129DNAHomo sapiens
24tctatctatc tatctatctg tctgtctgtc tgtctgtctg tctatctatc tatatctatc
60tatctatcat ctatctatcc atatctatct atctatctat ctatctatct atctatctat
120ctatcgtct 12925133DNAHomo sapiens 25tctatctatc tatctatctg
tctgtctgtc tgtctgtctg tctatctatc tatatctatc 60tatctatcat ctatctatcc
atatctatct atctatctat ctatctatct atctatctat 120ctatctatcg tct
13326137DNAHomo sapiens 26tctatctatc tatctatcta tctatctgtc
tgtctgtctg tctgtctatc tatctatatc 60tatctatcta tcatctatct atccatatct
atctatctat ctatctatct atctatctat 120ctatctatct atcgtct
13727141DNAHomo sapiens 27tctatctatc tatctatcta tctgtctgtc
tgtctgtctg tctgtctatc tatctatatc 60tatctatcta tcatctatct atccatatct
atctatctat ctatctatct atctatctat 120ctatctatct atctatcgtc t
14128143DNAHomo sapiens 28tctatctatc tatctatcta tctgtctgtc
tgtctgtctg tctgtctatc tatctatatc 60tatctatcta tcatctatct atccatatct
atctatctat ctatctatct atctatctat 120ctatctatct atatctatcg tct
14329147DNAHomo sapiens 29tctatctatc tatctatcta tctgtctgtc
tgtctgtctg tctgtctatc tatctatatc 60tatctatcta tcatctatct atccatatct
atctatctat ctatctatct atctatctat 120ctatctatct atctatatct atcgtct
14730151DNAHomo sapiens 30tctatctatc tatctatcta tctgtctgtc
tgtctgtctg tctgtctatc tatctatatc 60tatctatcta tcatctatct atccatatct
atctatctat ctatctatct atctatctat 120ctatctatct atctatctat
atctatcgtc t 15131155DNAHomo sapiens 31tctatctatc tatctatcta
tctgtctgtc tgtctgtctg tctgtctatc tatctatatc 60tatctatcta tcatctatct
atccatatct atctatctat ctatctatct atctatctat 120ctatctatct
atctatctat ctatatctat cgtct 15532157DNAHomo sapiens 32tctatctatc
tatctatcta tctatctatc tatctatcta tctgtctgtc tgtctgtctg 60tctatctatc
tatatctatc tatctatcat ctatctatcc atatctatct atctatctat
120ctatctatct atctatctat ctatctatct atcgtct 15733161DNAHomo sapiens
33tctatctatc tatctatcta tctatctatc tatctatcta tctatctgtc tgtctgtctg
60tctgtctatc tatctatatc tatctatcta tcatctatct atccatatct atctatctat
120ctatctatct atctatctat ctatctatct atctatcgtc t 16134165DNAHomo
sapiens 34tctatctatc tatctatcta tctatctatc tatctatcta tctatctgtc
tgtctgtctg 60tctgtctatc tatctatatc tatctatcta tcatctatct atccatatct
atctatctat 120ctatctatct atctatctat ctatctatct atctatctat cgtct
16535169DNAHomo sapiens 35tctatctatc tatctatcta tctatctatc
tatctatcta tctatctatc tatctgtctg 60tctgtctgtc tgtctatcta tctatatcta
tctatctatc atctatctat ccatatctat 120ctatctatct atctatctat
ctatctatct atctatctat ctatcgtct 1693632DNAHomo sapiens 36agaaagaaag
aaagaaagaa agaaagaaag aa 323714DNAHomo sapiens 37tatctatcta tcta
1438108DNAArtificial SequenceDescription of Artificial Sequence
Synthetic oligonucleotide 38agaaagaaag aaagaaagaa agaaagaaag
aaagaaagaa agaaagaaag aaagaaagaa 60agaaagaaag aaagaaagaa agaaagaaag
aaagaaagaa agaaagaa 1083965DNAHomo sapiens 39tctatctatc tgtctgtctg
tctgtctatc tatctatcca tctatctatc tatccatcca 60tccat 65
D00001

D00002

D00003

D00004

D00005

D00006

D00007

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.