Methods, models, systems, and apparatus for identifying target sequences for Cas enzymes or CRISPR-Cas systems for target sequences and conveying results thereof
Zhang , et al. February 23, 2
U.S. patent number 10,930,367 [Application Number 14/738,483] was granted by the patent office on 2021-02-23 for methods, models, systems, and apparatus for identifying target sequences for cas enzymes or crispr-cas systems for target sequences and conveying results thereof. This patent grant is currently assigned to THE BROAD INSTITUTE, INC., MASSACHUSETTS INSTITUTE OF TECHNOLOGY, PRESIDENT AND FELLOWS OF HARVARD COLLEGE. The grantee listed for this patent is THE BROAD INSTITUTE INC., MASSACHUSETTS INSTITUTE OF TECHNOLOGY, PRESIDENT AND FELLOWS OF HARVARD COLLEGE. Invention is credited to Patrick Hsu, Yinqing Li, David Arthur Scott, Joshua Asher Weinstein, Feng Zhang.










View All Diagrams
| United States Patent | 10,930,367 |
| Zhang , et al. | February 23, 2021 |
Methods, models, systems, and apparatus for identifying target sequences for Cas enzymes or CRISPR-Cas systems for target sequences and conveying results thereof
Abstract
Disclosed are thermodynamic and multiplication methods concerning CRISPR-Cas systems, and apparatus therefor.
| Inventors: | Zhang; Feng (Cambridge, MA), Li; Yinqing (Cambridge, MA), Scott; David Arthur (Cambridge, MA), Weinstein; Joshua Asher (Cambridge, MA), Hsu; Patrick (Cambridge, MA) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Applicant: |
|
||||||||||
| Assignee: | THE BROAD INSTITUTE, INC.
(Cambridge, MA) MASSACHUSETTS INSTITUTE OF TECHNOLOGY (Cambridge, MA) PRESIDENT AND FELLOWS OF HARVARD COLLEGE (Cambridge, MA) |
||||||||||
| Family ID: | 1000005379109 | ||||||||||
| Appl. No.: | 14/738,483 | ||||||||||
| Filed: | June 12, 2015 |
Prior Publication Data
| Document Identifier | Publication Date | |
|---|---|---|
| US 20150356239 A1 | Dec 10, 2015 | |
Related U.S. Patent Documents
| Application Number | Filing Date | Patent Number | Issue Date | ||
|---|---|---|---|---|---|
| PCT/US2013/074812 | Dec 12, 2013 | ||||
| 61836080 | Jun 17, 2013 | ||||
| 61758468 | Jan 30, 2013 | ||||
| 61769046 | Feb 25, 2013 | ||||
| 61802174 | Mar 15, 2013 | ||||
| 61806375 | Mar 28, 2013 | ||||
| 61814263 | Apr 20, 2013 | ||||
| 61819803 | May 6, 2013 | ||||
| 61828130 | May 28, 2013 | ||||
| 61736527 | Dec 12, 2012 | ||||
| 61748427 | Jan 2, 2013 | ||||
| 61791409 | Mar 15, 2013 | ||||
| 61835931 | Jun 17, 2013 | ||||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | G16B 20/00 (20190201); C12N 15/113 (20130101); C12N 9/22 (20130101); C12N 15/1089 (20130101); C12N 2320/11 (20130101); C12N 2310/20 (20170501); C12N 15/1082 (20130101) |
| Current International Class: | G16B 20/00 (20190101); C12N 9/22 (20060101); C12N 15/113 (20100101); C12N 15/10 (20060101) |
References Cited [Referenced By]
U.S. Patent Documents
| 5622856 | April 1997 | Natsoulis |
| 6251677 | June 2001 | Wilson et al. |
| 7601492 | October 2009 | Fu et al. |
| 7691995 | April 2010 | Zamore et al. |
| 8697359 | April 2014 | Zhang |
| 8771945 | July 2014 | Zhang |
| 8865406 | October 2014 | Zhang et al. |
| 8871445 | October 2014 | Cong et al. |
| 8889418 | November 2014 | Zhang et al. |
| 8932814 | January 2015 | Cong et al. |
| 8945839 | February 2015 | Zhang |
| 8993233 | March 2015 | Zhang et al. |
| 9549901 | January 2017 | Shi et al. |
| 9597357 | March 2017 | Gregory et al. |
| 9623071 | April 2017 | Guo et al. |
| 9834791 | December 2017 | Zhang et al. |
| 9873894 | January 2018 | Conway et al. |
| 10301651 | May 2019 | Doudna et al. |
| 2003/0186238 | October 2003 | Allawi et al. |
| 2004/0111221 | October 2004 | Beattie |
| 2005/0196851 | September 2005 | Uckun |
| 2005/0220796 | October 2005 | Dynan et al. |
| 2006/0178297 | August 2006 | Troy et al. |
| 2006/0234247 | October 2006 | Puttaraju et al. |
| 2007/0016012 | January 2007 | Hartlep |
| 2007/0244031 | October 2007 | Lu et al. |
| 2008/0293655 | November 2008 | Aygun et al. |
| 2010/0055798 | March 2010 | Battersby |
| 2010/0076057 | March 2010 | Sontheimer et al. |
| 2010/0081707 | April 2010 | Ali et al. |
| 2010/0093617 | April 2010 | Barrangou et al. |
| 2010/0233084 | September 2010 | Narasimhaswamy et al. |
| 2011/0016540 | January 2011 | Weinstein |
| 2011/0059502 | March 2011 | Chalasani |
| 2011/0189776 | August 2011 | Terns et al. |
| 2011/0223638 | September 2011 | Wiedenheft et al. |
| 2011/0239315 | September 2011 | Bonas et al. |
| 2012/0029891 | February 2012 | Behlke et al. |
| 2013/0096182 | April 2013 | Chatterjee et al. |
| 2013/0130248 | May 2013 | Haurwitz et al. |
| 2013/0315831 | November 2013 | Shi et al. |
| 2014/0068797 | March 2014 | Doudna et al. |
| 2014/0294771 | October 2014 | Schaffer et al. |
| 2014/0295557 | October 2014 | Joung et al. |
| 2014/0315985 | October 2014 | May et al. |
| 2014/0342456 | November 2014 | Mali |
| 2014/0356956 | December 2014 | Church et al. |
| 2014/0357530 | December 2014 | Zhang et al. |
| 2015/0045546 | February 2015 | Siksnys et al. |
| 2015/0071899 | March 2015 | Liu et al. |
| 2015/0071903 | March 2015 | Liu et al. |
| 2015/0071906 | March 2015 | Liu et al. |
| 2015/0232881 | August 2015 | Glucksmann et al. |
| 2015/0247150 | September 2015 | Zhang et al. |
| 2015/0291965 | October 2015 | Zhang et al. |
| 2015/0322457 | November 2015 | Kim et al. |
| 2015/0353905 | December 2015 | Weiss et al. |
| 2016/0017366 | January 2016 | Chen et al. |
| 2016/0024510 | January 2016 | Bikard |
| 2016/0024524 | January 2016 | Joung et al. |
| 2016/0130609 | May 2016 | Doudna et al. |
| 2016/0237456 | August 2016 | Church et al. |
| 2016/0251648 | September 2016 | Wang et al. |
| 2016/0281072 | September 2016 | Zhang |
| 2016/0298135 | October 2016 | Chen |
| 2016/0298137 | October 2016 | Chen |
| 2016/0324938 | November 2016 | Bikard |
| 2016/0340662 | November 2016 | Zhang et al. |
| 2017/0175144 | June 2017 | Zhang et al. |
| 2018/0127783 | May 2018 | Zhang et al. |
| 2018/0230495 | August 2018 | Doudna et al. |
| 2019/0010471 | January 2019 | Zhang et al. |
| 112015013784 | Jul 2017 | BR | |||
| 101228176 | Jul 2008 | CN | |||
| 103388006 | Nov 2013 | CN | |||
| 103668472 | Mar 2014 | CN | |||
| 104854241 | Aug 2015 | CN | |||
| 2591770 | May 2013 | EP | |||
| 2784162 | Jan 2014 | EP | |||
| 2764103 | Aug 2014 | EP | |||
| 2771468 | Sep 2014 | EP | |||
| 2 828 386 | Jan 2015 | EP | |||
| 2872170 | Dec 2005 | FR | |||
| 49/2015 | Dec 2015 | IN | |||
| 2004-519245 | Jul 2004 | JP | |||
| 2004-537285 | Dec 2004 | JP | |||
| 2005-509409 | Apr 2005 | JP | |||
| 2006-513694 | Apr 2006 | JP | |||
| 2006-518996 | Aug 2006 | JP | |||
| 2007-501626 | Feb 2007 | JP | |||
| 2009502170 | Jan 2009 | JP | |||
| 2009-536827 | Oct 2009 | JP | |||
| 2010-522547 | Jul 2010 | JP | |||
| 2012-508235 | Apr 2012 | JP | |||
| 2012-511332 | May 2012 | JP | |||
| 2012510812 | May 2012 | JP | |||
| 2012-529287 | Nov 2012 | JP | |||
| 2013-500045 | Jan 2013 | JP | |||
| 2013-518602 | May 2013 | JP | |||
| 2013-544077 | Dec 2013 | JP | |||
| 2014-526279 | Oct 2014 | JP | |||
| 2015-523856 | Aug 2015 | JP | |||
| 2016-500262 | Jan 2016 | JP | |||
| 2016-501531 | Jan 2016 | JP | |||
| 2016-501532 | Jan 2016 | JP | |||
| 2016500003 | Jan 2016 | JP | |||
| 2016-025710 | Feb 2016 | JP | |||
| 2016-504026 | Feb 2016 | JP | |||
| 2016502840 | Feb 2016 | JP | |||
| 2016-093196 | May 2016 | JP | |||
| 2016-516169 | Jun 2016 | JP | |||
| 2016-517954 | Jun 2016 | JP | |||
| 2016-131404 | Jul 2016 | JP | |||
| 2016-520317 | Jul 2016 | JP | |||
| 2016-521554 | Jul 2016 | JP | |||
| 2016-521975 | Jul 2016 | JP | |||
| 2016-521995 | Jul 2016 | JP | |||
| 2016-523082 | Aug 2016 | JP | |||
| 2016-524472 | Aug 2016 | JP | |||
| 2016182140 | Oct 2016 | JP | |||
| 2017-501151 | Jan 2017 | JP | |||
| 2017-501699 | Jan 2017 | JP | |||
| 6395765 | Sep 2018 | JP | |||
| 2009136452 | Apr 2011 | RU | |||
| WO-02/074968 | Sep 2002 | WO | |||
| WO-02/080851 | Oct 2002 | WO | |||
| WO-03/014318 | Feb 2003 | WO | |||
| WO-03/104414 | Dec 2003 | WO | |||
| WO-2004/029219 | Apr 2004 | WO | |||
| WO-2004/046321 | Jun 2004 | WO | |||
| WO-2004/062618 | Jul 2004 | WO | |||
| WO-2005/014791 | Feb 2005 | WO | |||
| 2005049642 | Jun 2005 | WO | |||
| 2007014275 | Feb 2007 | WO | |||
| WO-2007/134161 | Nov 2007 | WO | |||
| 2008093152 | Aug 2008 | WO | |||
| 2008108989 | Sep 2008 | WO | |||
| WO-2008/116860 | Oct 2008 | WO | |||
| 2010054108 | May 2010 | WO | |||
| 2010065123 | Jun 2010 | WO | |||
| WO-2010/068816 | Jun 2010 | WO | |||
| 2010079430 | Jul 2010 | WO | |||
| WO-2010/075424 | Jul 2010 | WO | |||
| WO-2010/143917 | Dec 2010 | WO | |||
| 2011011767 | Jan 2011 | WO | |||
| WO-2011/016840 | Feb 2011 | WO | |||
| WO-2011/036510 | Mar 2011 | WO | |||
| 2011064736 | Jun 2011 | WO | |||
| WO-2011/076873 | Jun 2011 | WO | |||
| WO-2011/100058 | Aug 2011 | WO | |||
| 2011146121 | Nov 2011 | WO | |||
| WO-2012/012738 | Jan 2012 | WO | |||
| WO-2012/031205 | Mar 2012 | WO | |||
| WO-2012/051343 | Apr 2012 | WO | |||
| 2012149470 | Nov 2012 | WO | |||
| 2012164565 | Dec 2012 | WO | |||
| WO-2013/044008 | Mar 2013 | WO | |||
| WO-2013/052681 | Apr 2013 | WO | |||
| WO-2013/071440 | May 2013 | WO | |||
| WO-2013/078400 | May 2013 | WO | |||
| 2013082519 | Jun 2013 | WO | |||
| 2013098244 | Jul 2013 | WO | |||
| 2013130824 | Sep 2013 | WO | |||
| 2013141680 | Sep 2013 | WO | |||
| 2013142578 | Sep 2013 | WO | |||
| WO-2013/155572 | Oct 2013 | WO | |||
| 2013176772 | Nov 2013 | WO | |||
| WO 2013/176772 | Nov 2013 | WO | |||
| WO-2014/165349 | Mar 2014 | WO | |||
| 2014065596 | May 2014 | WO | |||
| 2014089290 | Jun 2014 | WO | |||
| 2014093479 | Jun 2014 | WO | |||
| 2014093595 | Jun 2014 | WO | |||
| 2014093622 | Jun 2014 | WO | |||
| 2014093635 | Jun 2014 | WO | |||
| 2014093661 | Jun 2014 | WO | |||
| 2014093694 | Jun 2014 | WO | |||
| 2014093701 | Jun 2014 | WO | |||
| 2014093709 | Jun 2014 | WO | |||
| 2014093712 | Jun 2014 | WO | |||
| 2014093718 | Jun 2014 | WO | |||
| 2014099744 | Jun 2014 | WO | |||
| 2014099750 | Jun 2014 | WO | |||
| WO-2014/093655 | Jun 2014 | WO | |||
| WO-2015/031775 | Aug 2014 | WO | |||
| WO-2014/144761 | Sep 2014 | WO | |||
| WO-2014/186585 | Nov 2014 | WO | |||
| 2014204724 | Dec 2014 | WO | |||
| 2014204725 | Dec 2014 | WO | |||
| 2014204727 | Dec 2014 | WO | |||
| 2014204729 | Dec 2014 | WO | |||
| WO-2014/191518 | Dec 2014 | WO | |||
| WO-2014/197568 | Dec 2014 | WO | |||
| WO-2014/197748 | Dec 2014 | WO | |||
| WO-2014/204726 | Dec 2014 | WO | |||
| WO-2014/204728 | Dec 2014 | WO | |||
| WO-2015/006747 | Jan 2015 | WO | |||
| WO-2015/035136 | Mar 2015 | WO | |||
| WO-2015/048577 | Apr 2015 | WO | |||
| WO-2015/065964 | May 2015 | WO | |||
| WO-2015/070083 | May 2015 | WO | |||
| WO-2015/071474 | May 2015 | WO | |||
| 2015089419 | Jun 2015 | WO | |||
| WO-2015/089351 | Jun 2015 | WO | |||
| WO-2015/089364 | Jun 2015 | WO | |||
| WO-2015/089427 | Jun 2015 | WO | |||
| WO-2015/113063 | Jul 2015 | WO | |||
| WO-2016/022866 | Feb 2016 | WO | |||
| WO-2016/141224 | Sep 2016 | WO | |||
Other References
|
Tulpan et al. BMC Bioinfornnatics. 2010; 11: 105. Published online Feb. 24, 2010. doi: 10.1186/1471-2105-11-105. cited by examiner . Marraffini et al. US 2010/0076057. cited by examiner . Panyam, et al. "Biodegradable nanoparticles for drug and gene delivery to cells and tissue" Advanced Drug Delivery Reviews, 2003, 55:329-347. cited by applicant . Patterson, et al. "Codon optimization of bacterial luciferase (lux) for expression in mammalian cells" J. Ind. Microbio. Biotechnology, 2005, 32:115-123. cited by applicant . Pinera, et al. "RNA-guided gene activation by CRISPR-Cas9-based transcription factors" Nature Methods, 2013, 10(10):973-978. cited by applicant . Platt, et al. "CRISPR-Cas9 Knockin Mice for Genome Editing and Cancer Modeling" Cell, 2014, 159(2):440-455. cited by applicant . Porteus, et al. "Gene targeting using zinc finger nucleases" Nature Biotechnology, 2005, 23(8):967-973. cited by applicant . Pougach, et al. "Transcription, Processing and Function of CRISPR Cassettes in Escherichia coli" Mol. Microbiol, 2010, 77(6):1367-1379. cited by applicant . Qi, et al. "Repurposing CRISPR as an RNA-Guided Platform for Sequence-Specific Control of Gene Expression" Cell, 2013, 152(5):1173-1183. cited by applicant . Ran, et al, "In vivo genome editing using Staphylococcus aureus Cas9," Nature, 2015, doi:10.1038/nature14299. cited by applicant . Ran et al. "Double Nicking by RNA-Guided CRISPR Cas9 for Enhanced Genome Editing Specificity," Cell 154, 1-10, Sep. 12, 2013. cited by applicant . Ran, et al. "Genome engineering using the CRISPR-Cas9 system" Nature Protocols, 2013, 8(11):2281-2308. cited by applicant . Rand, et al. "Argonaute2 Cleaves the Anti-Guide Strand of siRNA during RISC Activation" Cell, 2005, 123:621-629. cited by applicant . Raymond, et al. "High-Efficiency FLP and .phi.C31 Site-Specific Recombination in Mammalian Cells" PLoS ONE, 2007, 2(1):e162. Doi. 10.1371/journal.pone.0000162. cited by applicant . Rebar, et al. "Induction of angiogenesis in a mouse model using engineered transcription factors" Nature Medicine, Dec. 2002, 8(12):1427-1432. cited by applicant . Reiss, et al. "RecA protein stimulates homologous recombination in plants" Proc. Natl. Acad. Sci. USA, 1996, 93:3094-3098. cited by applicant . Sanders, et al. "Use of a macromolecular crowding agent to dissect interactions and define functions in transcriptional activation by a DNA-tracking protein: Bacteriophage T4 gene 45 protein and late transcription" PNAS, 1994, 9:7703-7707. cited by applicant . Sapranauskas, et al. "The Streptococcus thermophilus CRISPR-Cas system provides immunity in Escherichia coli" Nucleic Acids Research, 2011, 39(21): 9275-9282. cited by applicant . Sauer, "Functional Expression of the cre-lox Site-Specific Recombination System in the Yeast Saccharomyces cerevisiae" Mol. Cell. Biol., 1987, 7(6):2087-2096. cited by applicant . Sauer, et al. "Site-specific DNA recombination in mammalian cells by the Cre recombinase of bacteriophage P1" Proc. Natl. Acad. Sci. U.S.A., 1988, 85:5166-5170. cited by applicant . Schiffer, et al. "Predictors of Hepatitis B Cure Using Gene Therapy to Deliver DNA Cleavage Enzymes: A Mathematical Modeling Approach" PLOS Computational Biology., 2013, 9(7):e1003131. www.ploscompbiol.org. cited by applicant . Scholze, et al. "TAL effector-DNA specificity" Virulence, 2010, 1(5):428-432, DOI:10.4161/viru.1.5.12863. cited by applicant . Schramm et al. "Recruitment of RNA polymerase III to its target promoters" Genes & Development, 2002, 16:2593-2620. cited by applicant . Seung Woo Cho, et al., "Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease" Nature Biotechnology, Mar. 2013, 31(3):230-232. cited by applicant . Seung Woo Cho, et al. "Supplementary Information: Targeted genome engineering in human cells with RNA-guided endonuclease" Nature Biotechnology, Mar. 2013, 31(3):1-10. cited by applicant . Seung Woo Cho, et al. "Analysis off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases" Genome Research, Nov. 2014, 24:132-141. cited by applicant . Shalem et al., "Genome-Scale CRISPR-Cas9 Knockout Screening in Human Cells", Science, 2014, 343;84-87. DOI:10.1126/science.1247005. cited by applicant . Shen, et al. "Efficient genome modification by CRISPR-Cas9 mickase with minimal off-target effects" 2014, Nature Methods, 11(4):399-404. cited by applicant . Shen, et al. "Generation of gene-modified mice via Cas9/RNA-mediated gene targeting" Cell Research, 2013, 23:720-723. cited by applicant . Sims, et al., "High-throughput RNA interference screening using pooled shRNA libraries and next generation sequencing", Genome Biology 12(10):R104, Oct. 2011. cited by applicant . Sontheimer, "Project 7: Establishing RNA-Directed DNA Targeting in Eukaryotic Cells" Physical Sciences-Onc., Nov. 16, 2011-Dec. 31, 2012, htt://groups.molbiosci.northwestern.edu/sontheimer/Sontheimer_cv.php) Molecular Biosciences. cited by applicant . Sosa, et A. "Animal transgenesis: an overiew" Brain Struct Funct, 2010, 214:91-109. cited by applicant . Stolfi, et al, "Tissue-specific genome editing in Ciona embryos by CRISPR/Cas9," Development, 2014, 141:4115-4120 doi:10.1242/dev.114488. cited by applicant . Swiech et al., "In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9," Nature Biotechnology, 2015, doi:10.1038/nbt.3055. cited by applicant . Terns, et al. "Crispr-based adaptive immune systems" Current Opinion in Microbiology, 2011, 14:321-327. cited by applicant . Tolia, et al. "Slicer and the Argonautes" Nature Chemical Biology, 2007, 3(1):36-43. cited by applicant . Trevino, et al. "Genome Editing Using Cas9 Nickases" Methods in Enxymology, 2014, 546:161-174. cited by applicant . Urnov, et al. "Highly efficient endogenous human gene correction using designed zinc-finger nucleases" Nature, 2005, 435:646-651. cited by applicant . Wang, et al. One-Step Generation of Mice Carrying Mutations in Multiple Genes by CRISPR/Cas-Mediated Genome Engineering, Cell, 2013, 153:910-918. cited by applicant . Wiedenheft, et al. "RNA-guided genetic silencing systems in bacteria and archaea" Nature, 2012, 432:331-338. cited by applicant . Wu, et al. "Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells"Nature Biotechnology, 2014, doi:10.1038/nbt.2889. cited by applicant . Xiao, et al. "Chromosomal deletions and inversions mediated by TALENs and CRIPPR/Cas in zebrafish" Nucleic Acids Research, 2013, 41(14):E141. doi:10.1093/nar/gkt464. cited by applicant . Xiao, et al. Production of High-Titer Recombinant Adeno-Associated Virus Vectors in the Absence of Helper Adenovirus. cited by applicant . Zetsche et al. "A split-Cas9 architecture for inducible genome editing and transcription modulation," Nature biotechnology, 2015, 33(2): 139-142. cited by applicant . Zhang, X. D., et al., "cSSMD: assessing collective activity for addressing off-target effects in genome-scale RNA interference screens", Bioinformatics (Oxford), 27(20);2775-2781, Oct. 2011. cited by applicant . Zhang, et al. "Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription" nature biotechnology, 2011, 29(2):149-154. cited by applicant . Zhu, et al. "Crystal structure of Cmr2 suggests a nucleotide cyclase-related enzyme in type III CRISPR-Cas sytems" FEBS Letters, 2012, 939-945. Doi:10.1016/j.febslet2012.02.036. cited by applicant . Gei ler, et al. "Trancscriptional Activators of Human Genes with Programmable DNA-Specificity" PLos One, 2011, 6(5):e19509. Doi:10.1371/hournal.pone.0019509. cited by applicant . Le Rhun, et al., Small RNAs in Streptococci, RNA Biology (Apr. 4, 2012) vol. 9, Issue 4, p. 414-426. cited by applicant . Clark, et al., A TALE of Two Nucleases: Gene Targeting for the Masses? Zebrafish (2011) vol. 8, No. 3, p. 147-149. cited by applicant . Mahfouz, et al., De Novo-engineered Transcription Activator-Like Effector (TALE) Hybrid Nuclease with Novel DNA Binding Specificity Creates Double-Strand Breaks, PNAS (Feb. 8, 2011) vol. 108, No. 6, p. 2623-2628. cited by applicant . Bobis-Wozowicz, et al., Targeted Genome Editing in Pluripotent Stem Cells Using Zinc-Finger Nucleases, Methods (2011) vol. 53, p. 339-346. cited by applicant . Swarthout, et al., Zinc Finger Nucleases: A New Era for Transgenic Animals, Annals of Neurosciences (Jan. 2011) vol. 18, No. 1, p. 25-28. cited by applicant . Goncalves, et al., Concerted Nicking of Donor and Chromosomal Acceptor DNA Promotes Homology-Directed Gene Targeting in Human Cells, Nucleic Acids Research (2012) vol. 40, No. 8, p. 3443-3455. cited by applicant . Van Nierop, et al., Stimulation of Homology-Directed Gene Targeting at an Endogenous Human Locus by a Nicking Endonuclease, Nucleic Acids Research (2009) vol. 37, No. 17, p. 5725-5736. cited by applicant . Lee, et al., Correction of the F508 Mutation in the Cystic Fibrosis Transmembrane Conductance Regulator Gene by Zinc-Finger Nuclease Homology-Directed Repair, Bioresearch Open Access (2012) vol. 1, No. 3, p. 99-108. cited by applicant . Gaj, et al., Targeted Gene Knockout by Direct Delivery of Zinc-Finger Nuclease Proteins, Nature Methods (Aug. 2012) vol. 9, No. 8, p. 805-807. cited by applicant . Porteus, et al., Chimeric Nucleases Stimulate Gene Targeting in Human Cells, Science (May 2, 2003) vol. 300, p. 763. cited by applicant . Sung, et al., The Importance of Valency in Enhancing the Import and Cell Routing Potential of Protein Transduction Domain-Containing Molecules, Biochimica et Biophysica Aeta (2006) vol. 1758, p. 355-363. cited by applicant . Dworetzky, et al., The Effects of Variations in the Number and Sequence of Targeting Signals on Nuclear Uptake, The Journal of Cell Biology (1988) vol. 107, p. 1279-1287. cited by applicant . Mincer, et al., Simulations of Nuclear Pore Transport Yield Mechanistic Insights and Quantitative Predictions, PNAS (Aug. 2, 2011) vol. 108, No. 31, p. E351-E358. cited by applicant . U.S. Appl. No. 61/734,256, filed Dec. 6, 2012, Fuqiang Chen. cited by applicant . U.S. Appl. No. 61/758,624, filed Jan. 30, 2013, Fuqiang Chen. cited by applicant . U.S. Appl. No. 61/761,046, filed Feb. 5, 2013, Scott Knight. cited by applicant . U.S. Appl. No. 61/761,422, filed Mar. 15, 2013, Scott Knight. cited by applicant . U.S. Appl. No. 61/735,876, filed Dec. 11, 2012, Blake A. Wiedenheft. cited by applicant . U.S. Appl. No. 61/799,531, filed Mar. 15, 2013, Blake A. Wiedenheft. cited by applicant . U.S. Appl. No. 61/738,355, filed Dec. 17, 2012, George M. Church. cited by applicant . U.S. Appl. No. 61/799,169, filed Mar. 13, 2012, Prashant Mali. cited by applicant . U.S. Appl. No. 61/613,373, filed Mar. 20, 2012, Virginijus Siksnys. cited by applicant . U.S. Appl. No. 61/625,420, filed Apr. 17, 2012, Virginijus Siksnys. cited by applicant . U.S. Appl. No. 61/652,086, filed May 5, 2012, Martin Jinek. cited by applicant . U.S. Appl. No. 61/716,256, filed Oct. 19, 2012, Martin Jinek. cited by applicant . U.S. Appl. No. 61/736,527, filed Dec. 12, 2012, F. Zhang. cited by applicant . U.S. Appl. No. 61/757,640, filed Jan. 28, 2013, Jinek. cited by applicant . U.S. Appl. No. 61/794,422, filed Mar. 15, 2013, Knight. cited by applicant . Andreas, et al. "Enhanced efficiency through nuclear localization signal fusion on phage C31-integrase: activity comparison with Cre and FLPe recombinase in mammalian cells" Nucleic Acids Research, 2002, 30(11):2299-2306. cited by applicant . Asuri, et al. "Directed Evolution of Adeno-Associated Virus for Enhanced Gene Delivery and Gene Targeting in Human Pluripotent Stem Cells" Molecular Therapy, Feb. 2012, 30(2):329-338. cited by applicant . Al-Attar et al., "Clustered regularly interspaced short palindromic repeats (CRISPRs): the hallmark of an ingenious antiviral defense mechanism in prokaryotes" Biol Chem., 2011, 392(4):277-289. cited by applicant . Baker, "Gene editing at CRISPR Speed" Nature Biotechnology, 2014, 32(4):309-312. cited by applicant . Banaszewska, et al. "Proprotein Convertase Subtilisin/Kexin Type 9: A New Target Molecule for Gene Therapy" Cellular & Molecular Biology Letters, Feb. 2012, 17(2):228-239. cited by applicant . Barrangou, "RNA-mediated programmable DNA cleavage" Nature Biotechnology, Sep. 2012, 30(9):836-388. cited by applicant . Bergemann, et al. "Excision of specific DNA-sequences from integrated retroviral vectors via site-specific recombination" Nucleic Acids Res., 1995, 23(21):4451-4456. cited by applicant . Bikard, et al. "CRISPR Interference Can Prevent Natural Transformation and Virulence Acquisition During In Vivo Bacterial Infection" Cell Host & Microbe, Aug. 2012, vol. 12:177-186. cited by applicant . Boch, et al. "Breaking the Code of DNA Binding Specificity of TAL-Type III Effecors" Science, 2009, 326:1509-1512. cited by applicant . Boch, et al. "Xanthomonas AvrBs3 Family-Type III Effectors: Discovery and Function" Annu. Rev. Phytopathol, 2010, Vo. 48:419-436. cited by applicant . Bogdanove, et al. "TAL Effectors:Customizable Proteins for DNA Targeting" Science, 2011, 333:1843-1846. cited by applicant . Briner, et al. "Guide RNA Functional Modules Direct Cas9 Activity and Orthogonality" Molecular Cell, Oct. 2014, 56:333-339. cited by applicant . Carroll, "A CRISPR Approach to Gene Targeting" Molecular Therapy, 2012, 20(9): 1658-1660. cited by applicant . Cermak, et al. Efficient design and assembly of custom TALEN and other TAL Effector-Based Constructs for DNA Targeting, Nucleic Acids Research (2011) vol. 39, No. 12, e82, p. 1-11. cited by applicant . Chen, et al. "Dynamic Imaging of Genomic Loci in Living Human Cells by an Optimized CRISPR/Cas System" Cell, Dec. 2013, vol. 155:1479-1491. cited by applicant . Jieliang Chen, et al. "An Efficient Antiviral Strategy for Targeting Hepatitis B Virus Genome Using Transcription Activator-Like Effector Nucelases" Molecular Therapy, 2014, 22(2):303-311. cited by applicant . Sidi Chen, et al., "Genome-wide CRISPR Screen in a Mouse Model of Tumor Growth and Metastasis," Cell, 2015, 160:1-15, http://dx.doi.org/10.1016/j.cell.2015.02.038. cited by applicant . Cho, et al. "Generation of Transgenic Mice" Curr Protoc Cell Biol., 2011, 19.11.doi:10.1002/0471143030.cb1911s42. cited by applicant . Choulika, et al., "Transfer of Single Gene-Containing Long Terminal Repeats into the Genome of Mammalian Cells by a Retroviral Vector Carrying the cre Gene and the loxP site" Journal of Virology, 1996, 70(3)1792-1798. cited by applicant . Christian, et al. "Targeting DNA Double-Strand Breaks With TAL Effector Nucleases" Genetics, Oct. 2010, vol. 186:757-761. cited by applicant . Christian, et al. "Supporting Information-Targeting DNA Double-Strand Breaks With TAL Effector Nucleases" Genetics, 2010, DOI:10.1534/110.120717:1SI-8SI. cited by applicant . Chylinski, et al. "The tractRNA and Cas9 Families of Type II CRISPR-Cas Immunity Systems" RNA Biology, May 2013, 10(5):726-737. cited by applicant . Cong, et al. "Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains" Nature Communications, 2012, 3:968, DOI:10/2038/ncomms1962. cited by applicant . Cong, et al, "Multiplex Genome Engineering Using CRISPR-Cas Systems" Science, 2013, 339:819-823. cited by applicant . Cong, et al, "Supplementary Material: Multiplex Genome Engineering Using CRISPR-Cas Systems" Science Express, Jan. 3, 2013. http://www.sciencemag.org/content/339/6121/819/suppl/DC1. cited by applicant . Connor, "Scientific split--the human genome breakthrough dividing former colleagues," Science, The Independent, Apr. 25, 2014, http://www.independent.co.uk/news/science/scientific-split--the-human-gen- ome-breakthrough-dividing-former-colleagues-9300456.html. cited by applicant . Dahlman, et al. "In vivo endothelial siRNA delivery using polymeric nanoparticles with low molecular weight" Nat. Nanotechnol., 2014, 9(8)648-655. doi:10.1038/nnano.2014.84. cited by applicant . Datensenko, et al. "Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system" Nature Communications, Jul. 10, 2012, 3:935, DOI:10.1038/ncomms1937. cited by applicant . Dingwall, et al. "Abstract: A Polypeptide Domain That Specifies Migration of Nucleoplasmin into the Nucleus" Cell, 1982, 30(2):449-58. cited by applicant . Deltcheva, et al. "CRISPR RNA maturation by trans-encoded small RNA and host Factor RNase III" Nature, Mar. 2011, vol. 471:602-609. cited by applicant . Deltcheva, et al. "Supplementary Information: CRISPR RNA Maturation by Trans-Encoded Small RNA and Host Factor RNase III" www.Nature.com/doi:10.1038/nature09886:1-35. cited by applicant . Drittanti, et al. "High throughput production, screening and anyalysis of adeno-associated viral vectors" Gene Therapy, 2000, 7:924-929. cited by applicant . Ebina, et al. "Harnessing the CRISPR/Cas9 system to disrupt latent HIV-1 provirus" Scientific Reports, 2013, 2:2510, doi:10.1038/srep02510. cited by applicant . Ellis, et al. Macromolecular Crowding: Obvious But Underappreciated, TRENDS in Biochemical Sciences, Oct. 2001, 26(10):597-604. cited by applicant . Ellis, et al. "Zinc-finger nuclease-mediated gene correction using single AAV vector transduction and enhanced by Food and Drug Administration-Approved Drugs" Gene Therapy, 2013, vol. 20:35-42. cited by applicant . Enyeart, et al., "Biotechnological applications of mobile group II introns and their reverse transcriptases: gene targeting, RNA-seq, and non-coding RNA analysis" Mobile DNA, 2014, 5:2, http://www.mobilednajournal.com/contents5/1/2. cited by applicant . Gabriel, et al. "An unbiased genome-wide analysis of zinc-finger nuclease specificity" Nature Biotechnology, Aug. 2011, 29(9):816-823. cited by applicant . Gaj, et al. "ZFN, TALEN, and CRISPR/Cas-Based Methods for Genome Engineering" Trends in Biotechnology, Jul. 2013, 31(7):397-405. cited by applicant . Garneau, et al. "The CRISPR-Cas bacterial immune systems cleaves bacteriophage and plasmid DNA" Nature, Nov. 2010, 468:67-71. cited by applicant . Gasiunas, et al. "Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria" PNAS, Sep. 2012, 109(39): E2579-E2586. cited by applicant . Gilbert, et al. "CRISPR-Mediated Modular RNA-Guided Regulation of Transcription in Eukaryotes" Cell, Jul. 2013, 154:442-451. cited by applicant . Goldfarb, et al. "Synthetic peptides as nuclear localization signals" Nature, Aug. 1986, 322(14):641-644. cited by applicant . Grens, Enzyme Improves CRISPR a smaller Cas9 protein enables in vivo genome engineering via viral vectors, The Scientist, Apr. 1, 2015. cited by applicant . Gustafsson, et al. "Codon Bias and heterologous protein expression" TRENDS in Biotechnology, Jul. 2004, 22(7):346-353. cited by applicant . Haft, et al. "Protein Families and Multiple CRISPR/Cas Subtypes Exist in Prokaryotic Genomes" PLoS Computational Biology, 2005, 1(6):0474-483. cited by applicant . Haft, et al. "A Guild of 45 CRISPR-Associated (Cas) Protein Families and Multiple CRISPR/Cas Subtypes Exist in Prokaryotic Genomes" PLoS Computational Biology, 2005, 1(6):0474-0483. cited by applicant . Hale, et al. "Essential Features and Rational Design of CRISPR RNAs that Function With the Cas RAMP Module Complex to Cleave RNAs" Molecular Cell, 2012, 45(3):292-302. cited by applicant . Hale, et al. "RNA-Guided RNA Cleavage by a CRISPR RNA-Cas Protein Complex" Cell, 2009, 139:945-956. cited by applicant . Hale, et al. "Prokaryotic siliencing (psi) RNAs in Pyrococcus furiosus", RNA, 2008, 14:2572-2579. cited by applicant . Handel, et al. "Versatile and Efficient Genome Editing in Human Cells by Combining Zinc-Finger Nucleases With Adeno-Associated Viral-Vectors" Human Gene Therapy, Mar. 2012, 23:321-329. cited by applicant . Hibbitt, et al. "RNAi-mediated knockdown of HMG CoA reductase enhances gene expression from physiologically regulated low-density lipoprotein receptor therapeutic vectors in vivo" Gene Therapy, 2012, 19:463-467. cited by applicant . Hou, et al., "Efficient genome engineering in human pluripotent stem cells using Cas9 from Neisseria meningitides," PNAS, 2013, 110(39):15644-15649. cited by applicant . Horvath et al. "RNA-guidded genome editing a la carte" Cell Research, 2013, 23:733-734, doi:10.1038/cr.2013.39. cited by applicant . Hsu, et al. "Development and Applications of CRISPR-Cas9 for Genome Engineering" Cell, Jun. 2014, 157:1262-1278. cited by applicant . Hsu et al., "DNA targeting specificity of RNA-guided Cas9 nucleases," Nature Biotechnology, 2013, 31(9):827-834. cited by applicant . Hsu et al., "Supplementary Information-DNA targeting specificity of RNA-guided Cas9 nucleases," Nature Biotechnology, 2013, doi:10.1038/nbt.2647. cited by applicant . Hwang Woong, et al. "Efficient genome editing in zebrafish using a CRISPR-Cas System" Nature Biotechnology, Mar. 2013, 31(3):227-229. cited by applicant . Hwang Woong, et al. "Efficient In Vivo Genome Editing Usng RNA-Guided Nucleases" Nat. Biotechnol., 2013, 31(3):227.229. doi. 1.1038/mbt.2501. cited by applicant . Janssen, et al., "Mouse Models of K-ras-Initiated Carcinogenesis" Biochimicia et Biophysica Acta, 2005, 1756:145-154. cited by applicant . Jiang, et al. "RNA-guided editing of bacterial genomes using CRISPR-Cas systems" Nature Biotechnology, 2013, 31(3):233-239. cited by applicant . Jinek, et al., "RNA-programmed genome editing in human cells;" 2013, eLife 2013:e00471, DOI:10.7554/eLife.00471. cited by applicant . Jinek, et al, "A Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity" Science, 2012, 337:816-821. cited by applicant . Jinek, et al., "Figures and figure supplements--A Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity" eLife 2014;2:e00471.DOI:10.7554/eLife 00471. cited by applicant . Kanasty, et al. "Delivery materials for siRNA therapeutics" Nature Materials, 2013, 12:967-977. cited by applicant . Karvelis, et al. "crRNA and tracrRNA guide Cas9-mediated DNA interference in Streptococcus thermophiles" RNA Biology, 2013, 10(5):841-851. cited by applicant . Karvelis, et al. "Supplemental Material to: crRNA and tracrRNA guide Cas9-mediated DNA interference in Streptococcus thermophiles" Landes Bioscience, 2013, 10(5), http://dx.doi.org/10.4161/rna.24203. cited by applicant . Kim, et al. "Crystal structure of Cas1 from Archaeoglobus fulgidus and characterization of its nucleolytic activity" Biochemical and Biophysical Research Communications, 2013, 441:720-725. cited by applicant . Konermann, et al, "Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex" Nature, 2015, 517:583-588. cited by applicant . Koornneef, et al. "Apoliprotein B Knockdown by AAV-Delivered shRNA Lowers Plasma Cholesterol in Mice" Molecular Therapy, Apr. 2011, 19( 4)731-740. cited by applicant . Lambowitz, et al. "Group II Introns: Mobile Ribozymes that Invade DNA" Cold Spring Hart Perspect Biol., 2011, 3:a003616. cited by applicant . Larson, et al. "CRISPR interference (CRISPRi) for sequence-specific control of gene expression" Nature Protocols, 2013, 8(11):2180-2196. cited by applicant . Lewis, et al., "The c-myc and PyMT oncogenes induce different tumor types in a somatic mouse model for pancreatic cancer" Genes & Development, 2003, 17:3127-3138. cited by applicant . Li et al. "In vivo genome editing restores hemostasis in a mouse model of hemophilia" Nature, 2011, 475(7355):217-221. doi: 10.1038/nature10177. cited by applicant . Li, et al. "Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotaina benthamiana using guide RNA and Cas9" Nature Biotechnology, 2013, 31(9):688-691. cited by applicant . Lombardo, et al., "Gene editing in human stem cells using zinc finger nucleases and integrase-defective lentiviral vector delivery" Nature Biotechnology, 2007, 25(11):1298-1306. cited by applicant . Luo, et al., "Highly parallel identification of essential genes in cancer cells", Proceeding of the National Academy of Sciences, 2008, 105(51);20380-20385. cited by applicant . Ma, et al. "A Guide RNA Sequence Design Platform for the CRISPR/Cas9 System for Model Organism Genomes", BioMed Research International, 2014, 2013:270805-4. http://dx.doi.org/10.1155/2014/270805. cited by applicant . Maeder, et al. "CRISPR RNA-guided activation of endogenous human genes" Nature Methods, 2013, 10(10):977-979. doi.10.1038/nmeth.2556. cited by applicant . Makarova, et al, "Evolution and classification of the CRISPR-Cas Systems" Nature Reviews Microbiology, 2011, 9(6):467-477. cited by applicant . Makarova et al. "Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systms" Biology Direct, 2011, 6:38. http:///www.biology-direct.com/content/6/1/38. cited by applicant . Mali, et al. "Supplementary Materials for--RNA-Guided Human Genome Engineering via Cas9" Science, 2013, 339:823-826. cited by applicant . Mali, et al. "RNA-Guided Human Genome engineering via Cas9" Science, 2013, DOI: 10.1126/SCIENCE.1232033. cited by applicant . Mali, et al. "CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering" nature biotechnology, 2013, 31(9):833-840. cited by applicant . Mali, et al. "Supplementary Information: Use of adjacent sgRNA:Cas9 complexes for transcriptional activation and genome engineering" Nature Biotechnoly, doi:10.1037/nbt.2675. cited by applicant . Malina, et al. "Repurposing CRISPR/Cas9 for in situ functional assays" Genes & Development, 2013, 27:2602-2614. cited by applicant . Marraffini, et al. "Self vs. non-self discrimination during CRISPR RNA-directed immunity" Nature, 2010, 463 (7280):568-571. cited by applicant . Mastroianni, et al. "Group II Intron-Based Gene Targeting Reactions in Eukaryotes" Plos One, 2008, 3(9):e3121. Doi:10.1371/journal.pone.0003121. cited by applicant . Meshorer, et al. "Chromatin in pluripotent embryonic stem cells and differentiation" Nature Reviews Molecular Cell Biology, 2006, 7:540-546. cited by applicant . Miller, et al. "A TALE nuclease architecture for efficient genome editing" Nature Biotechnology, 2011, 29(2):143-150. cited by applicant . Minton, "How can biochemical reactions within cells differ from those in test tubes?" Journal of Cell Science, 2006, 119:2863-2869. cited by applicant . Mojica et al. "Short motif sequences determine the targets of the prokaryotic CRISPR defence system", Microbiology, 2009, 155:733-740. cited by applicant . Morgan, et al. "Inducible Expression and Cytogenetic Effects of the EcoRI Restriction Endonuclease in Chinese Hamster Ovary Cells" Molecular and Cellular Biology, 1988, 8(10):4204-4211. cited by applicant . Mukhopadyay, "On the Same Wavelength," ASBMBTODAY, Aug. 2014, http://www.asbmb.org/asbmbtoday/201408/Features/Doudna/. cited by applicant . Nakamura, et al. "Codon usage tabulated from international DNA sequence databases: status for the year 2000" Nucleic Acids Research, 2000, 28(1): 292. cited by applicant . Nomura, et al., "Low-density lipoprotein receptor gene therapy using helper-dependent adenovirus produces long-term protection against atherosclerosis in a mouse model of familial hypercholesterolemia" Gene Therapy, 2004, 11:1540-1548. cited by applicant . Nishimasu et al. "Crystal Structure of Staphylococcus aureus Cas9," Cell 162, 1113-1126, Aug. 27, 2015. cited by applicant . Nishimasu, et al. Crystal Structure of Cas9 in Complex with Guide RNA and Target DNA, Cell, 2014, 156:935-949. cited by applicant . Oost, "New Tool for Genome Surgery" Science, Feb. 15, 2013, 399:768-770. cited by applicant . "The CRISPR Revolution," Catalyst Magazine, College of Chemistry, University of California, Berkeley, http://catalyst.berkeley.edu/slideshow/the-crispr-revolution/[Dec. 19, 2014 12:40:53] (Jul. 9, 2014). cited by applicant . Addgene Materials May 2015. cited by applicant . Addgene Materials Oct. 2014 including Addgene News 2013. cited by applicant . Addgene Reagent distribution list for Zhang Lab. cited by applicant . Arbitron, Inc. v. Kiefl, No. 09-CV-04013 PAC, 2010 WL 3239414, at *1 (S.D.N.Y. Aug. 13, 2010). cited by applicant . Autofluorescence MIT Flow Cytometry Core Facility (2018), 6 pages. cited by applicant . Baena-Lopez, L., et al., "Accelerated homologous recombination and subsequent genome modification in Drosophila," Development, vol. 140, No. 23, pp. 4818-4825, dated Dec. 2013, 18 pages, including Supplementary Material. cited by applicant . Balboa et al., "Conditionally Stabilized dCas9 Activator for Controlling Gene Expression in Human Cell Reprogramming and Differentiation. (plus Supplemental Information)", Stem Cell Reports, vol. 5, No. 3, Sep. 8, 2015, pp. 448-459 16PP. cited by applicant . Barrangou and Van Der Oost (Eds.), "CRISPR-Cas Systems," Springer Heidelberg (2013; written in 2012 before the publication of Cong et al.). cited by applicant . Barrangou, R. et al.: "CRISPR provides acquired resistance against viruses in prokaryotes," Science, vol. 315, pp. 1709-1712, dated Mar. 23, 2007, 5 pages. cited by applicant . Bennett et al., "Stable transgene expression in rod photoreceptors after recombinant adeno-associated virus-mediated gene transfer to monkey retina", Proc. Natl. Acad. Sci., USA, vol. 96, Aug. 1999, pp. 9920-9925. cited by applicant . Berns, K., et al. "A Large-Scale RNAi Screen in Human Cells Identifies New Components of the p53 Pathway," Nature 2004, vol. 428, pp. 431-437, dated Mar. 25, 2004, 7 pages. cited by applicant . Bhaya, D., et al., "CRISPR-Cas Systems in Bacteria and Archaea: Versatile Small RNAs for Adaptive Defense and Regulation," Annual Review of Genetics, vol. 45, No. 1, pp. 273-297, dated Dec. 15, 2011, 27 pages. cited by applicant . Bikard, et al. Supplementary Information for: "CRISPR Interference Can Prevent Natural Transformation and Virulence Acquisition During In Vivo Bacterial Infection," Cell Host & Microbe, vol. 12, No. 2, pp. 177-186, dated Aug. 16, 2012, 10 pages. cited by applicant . Bloom et al., "Inactivation of hepatitis B virus replication in cultured cells and in vivo with engineered transcription activator-like effector nucleases", Molecular Therapy, Aug. 20, 2013, vol. 21, No. 10, pp. 1889-1897. cited by applicant . Bohm, "The computer program LUDI: A new method for the de novo design of enzyme inhibitors", Journal of Computer-Aided Molecular Design, 1992, vol. 6, pp. 61-78. cited by applicant . Boutros et al.: "Genome-wide RNAi analysis of growth and viability in Drosophila cells," Science, American Association for the Advancement of Science, vol. 303, No. 5659, pp. 832-835, dated Feb. 6, 2004, 4 pages. cited by applicant . Branden, C., and Tooze, J., "Prediction, Engineering, and Design of Protein Structures," Introduction to Protein Structure, Chapter 16, p. 247, Garland Publishing, Inc., New York, dated 1991, 3 pages. cited by applicant . Brouns, S., et al., "Small CRISPR RNAs Guide Antiviral Defense in Prokaryotes," Science, vol. 321, pp. 960-964, dated Aug. 15, 2008, 6 pages. cited by applicant . Chinnasamy, D., et al., "Multicistronic lentiviral vectors containing the FMCV 2A Cleavage factor demonstrate robust expression of encoded genes at limiting MOI," Virology Journal, vol. 3, No. 4, dated Mar. 15, 2006, 16 pages. cited by applicant . Chou, JY, and Mansfield, BC, "Recombinant AAV-directed gene therapy for type I glycogen storage diseases," Expert Opinion on Biological Therapy, vol. 11, No. 8, pp. 1011-1024, dated Apr. 20, 2011, 21 pages. cited by applicant . Cockrell, "Berkeley's Wikipedian-in-residence is a first," NewsCenter, Feb. 25, 2014. cited by applicant . Community Corner, "CRISPR technology for gene therapy," Nature Medicine, vol. 20, No. 5, pp. 476-477, dated May 2014, 3 pages. cited by applicant . Cong, et al., Oct. 5, 2012 Manuscript including Supplementary Materials, "CRISPR-Assisted Mammalian Genome Engineering," published as "Multiplex Genome Engineering Using CRISPR-Cas Systems," Science, vol. 339, pp. 819-823, dated Feb. 15, 2013, 36 pages. cited by applicant . Cong, L., et al., "In Vivo Genome Engineering With AAV Vector Carrying CRISPR-Cas9 System," Molecular Therapy, vol. 22, Supplement 1, p. S214, dated May 23, 2014, 1 page. cited by applicant . Cong, L., et al., Supplementary Material for: "Multiplex Genome Engineering Using CRISPR-Cas Systems," Science Express, dated Jul. 5, 2012. cited by applicant . Cotropia, et al., "Copying in Patent Law," N.C.L. Rev., Stanford Public Law Working Paper No. 1270160 (2009), 87:1421. cited by applicant . Cummings et al., "Fourteen and counting: unraveling trinucleotide repeat diseases", Human Molecular Genetics, 2000, vol. 9, No. 6, pp. 909-916. cited by applicant . Daboussi, F., et al., "Chromosomal context and epigenetic mechanisms control the efficacy of genome editing by rare-cutting designer endonucleases," Nucleic Acids Research, vol. 40, No. 13, pp. 6367-6379, dated Jul. 13, 2012, 13 pages. cited by applicant . Daley, J., and Wilson, T., "Rejoining of DNA Double-Strand Breaks as a Function of Overhang Length," Molecular and Cellular Biology, vol. 25, No. 3, pp. 896-906, dated Feb. 2005, 11 pages. cited by applicant . Damian, M., and Porteus, M., "A Crisper Look at Genome Editing: RNA-guided Genome Modification," Molecular Therapy, vol. 21, No. 4, pp. 720-722, dated Apr. 2013, 3 pages. cited by applicant . Database UniProtKB/TrEMBL [online], Accession No. Q0P897, "The genome sequence of the food-borne pathogen Campylobacter jejuni reveals hypervariable sequences," Subname: Full=Putative CRISPR-associated protein, Oct. 3, 2012 uploaded, [retrieved on Nov. 22, 2017], URL, http://www.uniprot.org/uniprot/QOP897.txt?version=28. cited by applicant . Database WPI, Week 201437 Thomson Scientific, London, GB; AN 2014-J79552, XP-002737563, 2 pages. cited by applicant . Declaration of Feng Zhang for U.S. Appl. No. 14/054,414 dated Jan. 30, 2014, 10 pages. cited by applicant . Excerpt from Declaration of Feng Zhang, dated Sep. 9, 2015. cited by applicant . Federal Circuit decision in Dow Chemical Co. v. Nova Chemicals Corp., Appeal Nos. 2014-1431, 2014-1462 (Fed. Cir. Aug. 28, 2015) (Dow v. Nova), 25 pages. cited by applicant . Feldgarden et al., "Staphylococcus aureus M0408 acrHk-supercont1.1, whole genome shotgun sequence", NCBI Reference Sequence: NK_KB821326.1, Direct Submission, Dec. 10, 2012, pp. 1-4. cited by applicant . Fleming, J., et al.: "Adeno-Associated Virus and Lentivirus Vectors Mediate Efficient and Sustained Transduction of Cultured Mouse and Human Dorsal Root Ganglia Sensory Neurons," Human Gene Therapy, vol. 12, pp. 77-86, dated Jan. 1, 2001, 10 pages. cited by applicant . Fonfara et al., "Phylogeny of Cas9 determines functional exchangeability of dual-RNA and Cas9 among orthologous type II CRISPR-Cas systems", Nucleic Acids Research, 2014, vol. 42, No. 4, pp. 2577-2590. cited by applicant . Fu et al, "Targeted genome editing in human cells using CRISPR/Cas nucleases and truncated guide RNAsc", Jan. 1, 2014, The Use of CRISPR/Cas9 ZFNs and Talens in Generating Site-Specific Genome Alterations; Methods in Enzymology; ISSN 1557-7988, vol. 546, Elsevier, NL, pp. 21-45. cited by applicant . Gardlik, R., et al., "Vectors and delivery systems in gene therapy," Medical Science Monitor, vol. 11, No. 4, pp. RA110-121, dated Apr. 1, 2005, 12 pages. cited by applicant . Garriga-Canut, M., et al., "Synthetic zinc finger repressors reduce mutant huntingtin expression in the brain of R6/2 mice," Proceedings of the National Academy of Sciences, vol. 109, No. 45, pp. E3136-E3145, dated Nov. 6, 2012, 10 pages. cited by applicant . Geisinger et al., "In vivo blunt-end cloning through CRISPR /CAS9-facilitated non-nomoiogous end-joining", Nucleic Acid Research Advance Access, Jan. 13, 2016, pp. 1-15. cited by applicant . Habib, N., Assignment to Broad Institute, dated Jun. 9, 2014, 4 pages. cited by applicant . Hall, B., et al., "Overview: Generation of Gene Knockout Mice," Current Protocols in Cell Biology, unit 19.12, suppl. 44, pp. 1-17, dated 2009, 17 pages. cited by applicant . Harrison et al., "A CRISPR view of development", Genes & Development, vol. 28, No. 17, Sep. 1, 2014, pp. 1859-1872. cited by applicant . Holkers, M., et al., "Adenoviral vector DNA for accurate genome editing with engineered nucleases," Nature Methods, vol. 11, No. 10, pp. 1051-1057, Aug. 24, 2014, 8 pages (Only Abstract Available). cited by applicant . Houdebine, L., "The methods to generate transgenic animals and to control transgene expression," Journal of Biotechnology, vol. 98, pp. 145-160, dated 2002, 16 pages. cited by applicant . Huang v. California Institute of Technology, 2004 WL 2296330 (C.D. Cal. Feb. 18, 2004). cited by applicant . Hung, S., et al., "AAV-Mediated CRISPR/Cas Gene Editing of Retinal Cells in Vivo," Investigative Ophthalmology & Visual Science, vol. 57, No. 7, pp. 3470-3476, dated Jun. 2016, 7 pages. cited by applicant . Iwamoto et al., "A General Chemical Method to Regulate Protein Stability in the Mammalian Central Nervous System", Chemistry and Biology, Current Biology, London, GB, vol. 17, No. 9, Sep. 24, 2010, pp. 981-988. cited by applicant . Jackson, A., et al. "Widespread siRNA "off-target" transcript silencing mediated by seed region sequence complementarity," RNA 2006, vol. 12, No. 7, dated Mar. 16, 2006, 10 pages. cited by applicant . Jiang, W., et al., "RNA-guided editing of bacterial genomes using CRISPR-Cas systems," Nature Biotechnology, vol. 31, No. 3, pp. 233-239, dated Mar. 2013, 30 pages, including Supplementary Materials. cited by applicant . Jinek, M., et al, "A programmable Dual-RNA-Guided DNA Endonuclease in adaptive bacterial immunity," Science, vol. 337, n.6096, pp. 816-821, dated Aug. 17, 2012, 7 pages. cited by applicant . Joseph, T., and Osman, R., "Thermodynamic basis of selectivity in guide-target-mismatched RNA interference," Proteins, vol. 80, No. 5, pp. 1283-1298, dated May 2012, 26 pages. cited by applicant . Kim, E., et al., "Precision genome engineering with programmable DNA-nicking enzymes," Genome Research, vol. 22, pp. 1327-1333, dated 2012, 8 pages. cited by applicant . Kim, S., et al., "CRISPER RNAs trigger innate immune responses in human cells," Genome Research, pp. 1-7, dated Feb. 22, 2018, 8 pages. cited by applicant . Kleinstiver et al., "High-fidelity CRISP-Cas9 nucleases with no detectable genome-wide off-target effects", Nature, vol. 529, No. 7587, Jan. 28, 2016, pp. 490-495. cited by applicant . Kowalski, Thomas J., PowerPoint Presentation, "Interview Sep. 9, 2015". cited by applicant . Kumar, M., et al., "Systematic Determination of the Packaging Limit of Lentiviral Vectors," Human Gene Therapy, vol. 12, pp. 1893-1905, dated Oct. 10, 2001, 21 pages. cited by applicant . Kuwayama, H., "Enhancement of Homologous Recombination Efficiency by Homologous Oligonucleotides," Cell Interaction Sivakumar Gowder, IntechOpen, DOI: 10.5772/47779, dated Oct. 10, 2012, 12 pages. cited by applicant . Lebherz, C., et al., "Gene therapy with novel adeno-associated virus vectors substantially diminished atherosclerosis in a murine model of familial hypercholesterolemia," The Journal of Gene Medicine, vol. 6, pp. 663-672, dated Mar. 2, 2004, 10 pages. cited by applicant . Levitt, J., et al., "Intrinsic fluorescence and redox changes associated with apoptosis of primary human epithelial cells," Journal of Biomedical Optics, vol. 11, No. 6, pp. 064012 1-10, dated Nov./Dec. 2006, 10 pages. cited by applicant . Li et al., "Coevolution of CRISPR-Cas system with bacteria and phages", Hereditas, vol. 33, No. 3, Mar. 31, 2011, pp. 213-218. cited by applicant . Ma, M., et al., "A Guide RNA Sequence Design Platform for the CRISPR/Cas9 System for Model Organism Genomes," BioMed Research International, vol. 31, No. 3, pp. 822-824, dated Nov. 2013, 4 pages. cited by applicant . Maczuga, P., et al., "Embedding siRNA sequences targeting Apolipoprotein B100 in shRNA and miRNA scaffolds results in differential processing and in vivo efficacy," Molecular Therapy, vol. 21, No. 1, pp. 217-227, dated Jan. 2013, 11 pages. cited by applicant . Maeder, M., and Gersbach, C., "Genome-editing Technologies for Gene and Cell Therapy," Molecular Therapy, vol. 24, No. 3, pp. 430-446, dated Mar. 2016, 17 pages. cited by applicant . Maeder, M., et al., "Robust, synergistic regulation of human gene expression using TALE activators," Nature Methods, vol. 10, No. 3, pp. 243-245, dated Mar. 2013, 9 pages. cited by applicant . Mali et al., "CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering", Nature Biotechnology, Sep. 2013, vol. 31, No. 9, pp. 833-837. cited by applicant . Mali, et al.: "Supplementary Materials for--RNA-Guided Human Genome Engineering via Cas9" Science, vol. 339, pp. 823-826, dated Feb. 15, 2013, 8 pages. cited by applicant . Mali, P., et al., Supplementary Materials for: "CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering," Nature Biotechnology, vol. 31, No. 9, pp. 833-838, dated Aug. 1, 2013, 36 pages. cited by applicant . Manjunath, N., et al., "Newer Gene Editing Technologies toward HIV Gene Therapy," Viruses, vol. 5, No. 11, pp. 2748-2766, dated Nov. 14, 2013, 19 pages. cited by applicant . Manning v. Paradis, 296 F.3d 1098 (Fed. Cir. 2012). cited by applicant . Marraffini, L., "CRISPR-Cas Immunity against Phages: Its Effects on the Evolution and Survival of Bacterial Pathogens," PLOS, Pathogens, dated Dec. 12, 2013, 6 pages. cited by applicant . Marraffini, L., Assignment to Rockefeller University, dated Dec. 12, 2013, 3 pages. cited by applicant . Maxwell v. The Stanley Works, 2006 WL 1967012 *5 (M.D. Tenn. Jul. 11, 2006). cited by applicant . Mojica, F. J., et al., Supplementary Material for: "Short motif sequences determine the targets of the prokaryotic CRISPR defence system," Microbiology, vol. 155, No. 3, pp. 733-740, dated Mar. 1, 2009, 8 pages. cited by applicant . Morris et al., "Distributed automated docking of flexible ligands to proteins: Parallel applications of AutoDock 2.4*", Journal of Computer-Aided Molecular Design, 1996, vol. 10, pp. 293-304. cited by applicant . Muther, N., et al.: "Viral Hybrid Vectors for Somatic Integration--Are They the Better Solution?" Viruses, vol. 1, pp. 1295-1324, dated Dec. 15, 2009, 30 pages. cited by applicant . Opposition Against Appl. Ser. No. EP13818570.7 submitted by Schlich dated Oct. 26, 2015, 8 pages. cited by applicant . Opposition Against EP Appl. Ser. Appl. No. 2771468-B1 dated Oct. 26, 2015. cited by applicant . Ozawa, K., "Gene therapy using AAV," Uirusu, vol. 57, No. 1, pp. 47-55, dated Nov. 27, 2007, 13 pages (with English Abstract; No English Translation). cited by applicant . Pandika, et al., www.ozy.com/rising-stars-and-provocateurs/jennifer-doudna-crispr-code-kil- ler/4690; 2014 (Jul. 1, 2014). cited by applicant . Phillips, A., "The challenge of gene therapy and DNA delivery," The Journal of Pharmacy and Pharmacology, vol. 53, pp. 1169-1174, dated 2001, 6 pages. cited by applicant . Pougach, K.S., et al.: "CRISPR Adaptive Immunity Systems of Prokaryotes," Molecular Biology, vol. 46, No. 2, Apr. 2012, pp. 195-203, 1 page (English Abstract). cited by applicant . PowerPoint slide entitled "Development and Applications of CRISPR-Cas9 for Genome Editing" dated Sep. 9, 2015. cited by applicant . Pride, D., et al., "Analysis of Streptococcal CRISPRs from Human Saliva Reveals Substantial Sequence Diversity Within and Between Subjects Over Time," Genome Research, vol. 21, No. 1, pp. 126-136, dated Jan. 2011, 11 pages. cited by applicant . Qi, J., et al., "microRNAs regulate human embryonic stem cell division," Cell Cycle, vol. 8, No. 22, pp. 3729-3741, dated Nov. 15, 2009, 13 pages. cited by applicant . Radecke, S., et al., "Zinc-finger Nuclease-induced Gene Repair With Oligodeoxynucleotides: Wanted and Unwanted Target Locus Modifications," Molecular Therapy, vol. 18, No. 4, pp. 743-753, dated Apr. 2010, 11 pages. cited by applicant . Ran et al., "Double Nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity", Cell, Sep. 2013, vol. 154, pp. 1380-1389. cited by applicant . Rho, M., et al., "Diverse CRISPRs Evolving in Human Microbiomes," PLOS Genetics, vol. 8, No. 6, pp. e1002441, dated Jun. 2012, 12 pages. cited by applicant . Rodriguez et al., "AAV-CRISPR: A New Therapeutic Approach to Nucleotide Repeat Diseases", Molecular Therapy, vol. 22, Supplement 1, Abstract 247, May 2014, p. S94. cited by applicant . Rolling, "Recombinant AAV-mediated gene transter to the retina: gene therapy perspectives", Gene Therapy, Vo. 11, 2004, pp. S26-S32. cited by applicant . Rubin v. The General Hospital Corp., 2011-1439 (Fed. Cir. Mar. 28, 2013). cited by applicant . Sadowski, M., and Jones, D., "The sequence-structure relationship and protein function prediction," Current Opinion in Structural Biology, vol. 19, pp. 357-362, dated May 4, 2009, 6 pages. cited by applicant . Sanders, UC Berkeley Jan. 7, 2013 Press Release, available at http://newscenter.berkeley.edu/2013/01/07/cheap-and-easy-technique-to-sni- p-dna-could-revolutionize-gene-therapy/. cited by applicant . Seffernick, J., et al., "Melamine Deaminase and Atrazine Chlorohydrolase: 98 Percent Identical but Functionally Different," Journal of Bacteriology, vol. 183, No. 8, pp. 2405-2410, dated Apr. 2001, 6 pages. cited by applicant . Senis, E., et al., "CRISPR/Cas9-mediated genome engineering: An adeno-associated viral (AAV) vector toolbox," Biotechnology Journal, vol. 9, No. 11, Sp. Iss. SI, pp. 1402-1412, dated Sep. 4, 2014, 2 pages (Only Abstract Available). cited by applicant . Senturk et al., "A rapid and tunable method to temporally control cas9 expression enables the identification of essential genes and the interrogation of functional gene interactions in vitro and in vivo", Jul. 28, 2015, pp. 1-27, XP002756303, doi:10.1101/023366, Retrieved from the Internet: URL:http://biorxiv.org/content/early/2015/07/28/023366 [retrieved on Apr. 11, 2016). cited by applicant . Shengdar Tsai et al., "Dimeric CRISPR RNS-guided Fokl nucleases for highly specific genome editing", Nature Biotechnology, vol. 32, No. 6, Apr. 25, 2014, pp. 569-576. cited by applicant . Slaymaker et al., Rationally engineered Cas9 nucleases with improved specificity, Science, American Association for the Advancement of Science, US, vol. 351, No. 6268, Jan. 1, 2016, pp. 84-88. cited by applicant . Stoller, J. and Aboussouan, L., "Alpha1-antitrypsin deficiency," The Lancet, Seminar, vol. 365, No. 9478, pp. 2225-2236, dated Jun. 25, 2005, 12 pages. cited by applicant . Stratikopoulos, E., et al., "The hormonal action of IGF1 in postnatal mouse growth," Proceedings of the National Academy of Sciences, vol. 105, No. 49, pp. 19378-19383, dated Dec. 9, 2008, 6 pages. cited by applicant . Straub, C., et al., "CRISPR/Cas9-Mediated Gene Knock-Down in Post-Mitotic Neurons," PLOS One, vol. 9, No. 8, art. E105584, pp. 1-5, dated Aug. 2014, 6 pages. cited by applicant . Suzuki, K., et al., "In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration," Nature, vol. 540, art. 7631, pp. 144-149, dated Dec. 1, 2016, 44 pages. cited by applicant . Tang, T., et al., "A mouse knockout library for secreted and transmembrane proteins," Nature Biotechnology, vol. 28, No. 7, pp. 749-755, dated Jul. 2010, 9 pages. cited by applicant . Third Party Observations Concerning App. No. GB1420270.9 dated Jun. 30, 2015. cited by applicant . Third Party Observations Concerning Appl. No. EP2800811, dated Jul. 24, 2015. cited by applicant . Third Party Observations Concerning Appl. No. EP2800811, dated Sep. 4, 2015. cited by applicant . Third Party Observations Concerning Appl. No. GB1420270.9 dated Jun. 30, 2015. cited by applicant . Third Party Observations Concerning Appl. No. GB1420270.9, dated Jul. 13, 2015. cited by applicant . Third Party Observations in Accordance with Article 115 EPC, Appl. No. EP13824232.6, Pub. No. EP2764103A, Mar. 25, 2015. cited by applicant . Third Party Observations submitted by Broad Institute Inc. Concerning Appl. Ser. No. EP13793997.1 dated Jul. 24, 2015, 108 pages. cited by applicant . Third Party Observations submitted by Broad Institute Inc. Concerning Appl. Ser. No. EP13793997.1 dated Sep. 4, 2015, 25 pages. cited by applicant . Third Party Observations submitted by Regents of the University of California et al. Concerning App. No. GB1420270.9 dated Jul. 13, 2015, 18 pages. cited by applicant . Third Party-Observations, Appl. No. 1382432.6, Pub. No. EP2764103, Feb. 16, 2015. cited by applicant . Ultra-Precision Mfg. Ltd. v. Ford Motor Co., 2004 WL 3507671, *7, *11-12 (E.D. Mich. Mar. 30, 2004). cited by applicant . Van Der Oost, "New tool for genome surgery", Science, Feb. 2013, vol. 339, pp. 768-770. cited by applicant . Wienert, B., et al., "In vitro transcribed guide RNAs trigger an innate immune response via the RIG-I pathway," BioRxiv Preprint, dated Mar. 3, 2018, 28 pages. cited by applicant . Witkowski, A., et al., "Conversion of a Beta-Ketoacyl Synthase to a Malonyl Decarboxylase by Replacement of the Active-Site Cysteine with Glutamine," Biochemistry, vol. 38, pp. 11643-11650, dated Aug. 18, 1999, 8 pages. cited by applicant . Wittmann et al., "Engineered riboswitches: Expanding researchers' toolbox with synthetic RNA regulators", FEBS Letters, vol. 586, No. 15, Feb. 28, 2012, pp. 2076-2083. cited by applicant . Yanfang Fu et al., "Improving CRISPR-Cas nuclease specificity using truncated guide RNAs" (with Supplement Table), Nature Biotechnology, vol. 32, No. 3, Jan. 26, 2014, pp. 279-284. cited by applicant . Yin, H., et al., "Structure-guided chemical modification of guide RNA enables potent non-viral in vivo genome editing," Nature Biotechnology, vol. 35, pp. 1179-1187, dated Nov. 13, 2017, 22 pages. cited by applicant . Yu, W., et al., "Nrl knockdown by AAV-delivered CRISPR/Cas9 prevents retinal degeneration in mice," Nature Communications, vol. 8, art. 14716, dated Mar. 14, 2017, 15 pages. cited by applicant . Yusuke Miyazaki et al., Destabilizing Domains Derived from the Human Estrogen Receptor:, Journal of the American Chemical Society, vol. 134, No. 9, Mar. 7, 2012, pp. 3942-3945. cited by applicant . Zhang, F., PowerPoint Presentation: "Development and Applications of CRISPR-Cas9 for Genome Edition," Broad Institute/MIT, dated Sep. 9, 2015, 50 pages. cited by applicant . Schunder, et al. "First indication for a functional CRISPR/Cas system in Francisella tularensis" International Journal of Medical Microbiology, 2013, 303:1438-4221. cited by applicant . Singer, et al. "Applications of Lentiviral Vectors for shRNA Delivery and Transgenesis" Curr Gene Ther., 2008, 8(6):483-488. cited by applicant . Sung, et al. "An rpsL Cassette, Janus, for Gene Replacement through Negative Selectionin Streptococcus pneumoniae" Applied and Environmental Microbiology, 2001, 67(11):5190-5196. cited by applicant . Sung, Young Hoon, et al. "Mouse genetics: Catalogue and scissors" BMB Reports, 2012, 45(12):686-692. cited by applicant . Tinland, et al. "The T-DNA-linked VirD2 protein contains two distinct functional nuclear localization signals" Proc. Natl. Acad. Sci, 1992, 89:7442-7446. cited by applicant . Tiscornia, et al. "Development of Lentiviral Vectors Expressing siRNA" Gene Transfer-Delivery and Expression of DNA and RNA--A Laboratory Manual, 2007, Chapter 3:23-34. cited by applicant . Urrutia, et al. "KRAB-containing zing finger repressor proteins" Genome Biology, 2003, 4(10):231-231.8. cited by applicant . Urnov, et al. "Genome editing with engineered zinc finger nucleases" Nature Reviews, Genetics, 2010, 11:637-646. cited by applicant . Van Den Ackerveken, et al. "Recognition of the Bacterial Avirulence Protein AvrBs3 Occurs inside the Host Plant Cell" Cell, 1996, 87:1307-1316. cited by applicant . Vestergaard, et al. "CRISPR adaptive immune systems of Archaea" RNA Biology, 2014, 11(2):156-167. cited by applicant . Villion, et al. "The double-edged sword of CRISPR-Cas systems" Cell Research, 2013, 23:15-17. cited by applicant . Wang, et al. "Genetic Screens in Human Cells Using the CRISPR-Cas9 System", Science, 2014, 343:80-84. cited by applicant . Welch, et al. "Designing Genes for Successful Protein Expression" Methods in Enzymology, 2011, 498:43-66, DOI: 10.1016/B978-0-12-385120-8.00003-6. cited by applicant . Wolff, et al. "Nuclear security breached" Nature Biotechnology, 2001, 19:1118-1120. cited by applicant . Wu, Zhijian, et al. "Effect of Genome Size on AAV Vector Packaging" The American Society of Gene & Cell Therapy, (Jan. 2010), 18(1):80-86. cited by applicant . Xie, et al. "RNA-Guided Genome Editing in Plants Using a CRISPR-Cas System" Molecular Plant, 2013, 6(6):1975-1983. cited by applicant . Yaghmai, et al. "Optimized Regulation of Gene Expression Using Artificial Transcription Factors", Molecular Therapy, 2002, 5(6):685-694. cited by applicant . Yang, et al. "One-Step Generation of Mice Carrying Reporter and Conditional Alleles by CRISPR/Cas-Mediated Genome Engineering" Cell, 2013, 154:1370-1379. cited by applicant . Yi, et al. "Current Advances in Retroviral Gene Therapy" Current Gene Therapy, 2011, 11:218:228. cited by applicant . Yu, et al. "An efficient recombination system for chromosome engineering in Escherichia coli" PNAS, 2000, 97(11):5978-5983. cited by applicant . Yu, Zhongshen, et al. "Highly Efficient Genome Modifications Mediated by CRISPR/Cas9 in Drosophila" Genetics, 2013, 195:289-291. cited by applicant . Zetsche, et al. "CPF1 is a Single RNA-Guided Endonuclease of a Class 2 CRISPR-Cas System" Cell, 2015, 163:759-771. cited by applicant . Zhang, et al., Efficient Expression of CFTR Function with Adeno-Associated Virus Vectors that Carry Shortened CFTR Genes, PNAS (USA) (1998) vol. 95, p. 10158-10163. cited by applicant . Zhang, et al., "Optimized CRISPR Design", XP055167487, Oct. 23, 2013, URL:http//crispr.mit.edu/about[retrieved on Feb. 5, 2015]. cited by applicant . Zhang, et al. "Optogenetic interrogation of neural circuits: technology for probing mammalian brain structures" Nat Protoc., 2010, 5(3):439-456, doi:10.1038/nprot.2009.226. cited by applicant . Zolkiewska, et al. "ADAM Proteases:Ligand Processing and Modulation of the Notch Pathway" Cell Mol Life Sci, 2008, 65(13):2056-2068. cited by applicant . Notice of Opposition filed by Aug. 16, 2017 by Vossius against EP Patent No. 2840140. cited by applicant . Notice of Opposition filed Aug. 14, 2017 by Grund against EP Patent No. 2840140. cited by applicant . Notice of Opposition filed Aug. 11, 2017 by Schlich against EP Patent No. 2840140. cited by applicant . Notice of Opposition filed Aug. 16, 2017 by Mathys & Squire LLP against EP Patent No. 2840140. cited by applicant . The Broad Inst. v. The Regents of University of UCA--Decision on Motions for Patent Interference No. 106,048 filed Feb. 15, 2017. cited by applicant . Declaration of Paul Simons dated Dec. 22, 2015. cited by applicant . Holmes, "Understanding Scores" XP055167918, Oct. 23, 2013, https://groups.google.com/forum/#!profo_nt50txrP9Yb6e_LXccolb9hNf7gKeMLt6- rgaVQ4fOsQ/crispr/fkhX7Fu3r-l/rziHxKT76pYJ [retrieved on Feb. 6, 2015]. cited by applicant . Imagawa, et al. "Two nuclear localization signals are required for nuclear translocation of nuclear factor 1-A" FEBS Letters, 2000, 484118-124. cited by applicant . Jao, et al. "Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system" Proceeding of the National Academy of Sciences, 2013, www.pnas.org/cgi/doi/10.1073/pnas.1308335110. cited by applicant . Jinek, et al., "Supplementary Materials for a Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity" Science Express, 2012, DOI:10.1126/science.1225829. cited by applicant . Joshi, et al., "Evolution of I-Scel homing endonucleases with increased DNA recognition site specificity" Journal of Molecular Biology, 2011, 405(1):185-200; ePub: Oct. 26, 2010. cited by applicant . Joung, et al. "TALENs: a widely applicable technology for targeted genome editing" Nat Ref. Mol. Cell Biol., 2013, 14(1):49-55. doi:10.1038/nrm3586. cited by applicant . Kalderon, et al. "A Short Amino Acid Sequence Able to Specify Nuclear Location" Cell, 1984, 39:499-509. cited by applicant . Kiani, et al. "CAS9 gRNA engineering for genome editing, activation and repression" Nature Methods, Advanced Online Publication, 2015, DOI:10.1038/NMETH.3580. cited by applicant . Kinnevery, et al. "Emergence of Sequence Type 779 Methicillin-Resistant Staphylococcus aureus Harboring a Novel Pseudo Staphylococcal Cassette Chromosome mec (SCCmec)-SCC-SCC CRISPR Composite Element in Irish Hospitals" Antimicrobial Agents and Chemotherapy, 2013, 57(1):524-531. cited by applicant . Kondo, et al. "Highly Improved Gene Targeting by Germline-Specific Cas9 Expression in Drosphila" Genetics, 2013, 195:715-721. cited by applicant . Kosugi, et al. "Six Classes of Nuclear Localization Signals Specific to Different Binding Grooves of Importin .alpha..." The Journal of Biological Chemistry, 2009, 284(1):478-485. cited by applicant . Krauer, et al. "Identification of the nuclear localization signals within the Epstein-Barr virus EBNA-6 protein" Journal of General Virology, 2004, 85:165-172. cited by applicant . Kuhlman, et al. "Site-specific chromosomal integration of large synthetic constructs" Nucleic Acids Research, 2010, 38(6):1-10, doi:10.1093/nar/gkp1193. cited by applicant . Kuhlman, et al. "A place for everything Chromosomal intergration of large constructs" Bioengineered Bugs, Jul./Aug. 2010, 1(4)296-299. cited by applicant . Lanford, et al. "Induction of Nuclear Transport with a Synthetic Peptide Homologous to the SV40 T Antigen Transport Signal" Cell, 1986, 46:575-582. cited by applicant . Lange, et al. "Classical Nuclear Localization Signals: Definition, Function, and Interaction with Importin . . . " J. Biol. Chem., 2007, 282(8):5101-5105. cited by applicant . Lemay, et al. "Folding of the Adenine Riboswitch" Chemistry & Biology, 2006, 13:857-868. cited by applicant . Lewin, et al. "Nuclear localization sequences target proteins to the nucleus" Cells, 2006, 5:224. cited by applicant . Li, Ting, et al. "Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes" Nucleic Acids Research, 2011, 39(14):6315-6325. cited by applicant . Liu, et al. "Epstein-Barr Virus DNase Contains Two Nuclear Localization Signals Which Are Different in Sensitivity to the Hydrophobic Regions" Virology, 1998, 247:62-73. cited by applicant . Los, et al. "Halotag Technology: Cell Imaging and Protein Analysis" Cell Notes, 2006, 14:10-14. cited by applicant . Luo, Ming, et al. "Multiple Nuclear Localization Sequences Allow Modulation of 5-Lipoxygenase Nuclear Import" Traffic, 2004, 5:847-854. cited by applicant . Lyssenko, et al. "Cognate putative nuclear localization signal effects strong nuclear localization of a GFP reporter and facilitates gene expression studies in Caenorhabditis elegans" BioTechniques, 2007, 43:596-600. cited by applicant . Mahfouz, et al. "Targeted transcriptional repression using a chimeric TALE-SRDX repressor protein" Plant Mol Biol, 2012, 78:311-321. cited by applicant . Makarova, et al. "An updated evolutionary classification of CRISPR-Cas systems" Nature Reviews--Microbiology, 2015, 13:722-736. cited by applicant . Morbitzer, et al. "Regulation of selected genome loci using de novo-engineered transcription activator-like effector (TALE)-type transcription factors" PNAS, 2010, 108(50):21617-21622. cited by applicant . Morbitzer, et al. "Assembly of custom TALE-type DNA binding domains by modular cloning" 2011, 39(13):5790-5799. cited by applicant . Morin, et al. "Nuclear Localization of the Adenovirus DNA-Binding Protein: Requirement for Two Signals and Complementation during Viral Infection" Molecular and Cellular Biology, 1989, 9(10):4372-4380. cited by applicant . Moscou, et al. "A Simple Cipher Governs DNA Regognition by TAL Effectors" Science, 2009, 326:1501. cited by applicant . Mussolino, et al. "TALE nucleases: tailored genome engineering made easy" Current Opinion in Biotechnology, 2012, 23(5):644-650. cited by applicant . Nagarajan, et al. "A Hierarchy of Nuclear Localization Signals Governs the Import of the Regulatory Factor X Complex Subunits and MHC Class II Expression" The Journal of Immunology, 2004, 173:410.419. cited by applicant . Nakai, et al. "PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization" Trends in Biochem Sciences, 1999 24:34-35. cited by applicant . Noguchi, et al. "PDX-1 Protein Containing Its Own Antennapedia-Like Protein Transduction Domain Can Transduce Pancreatic Duct and Islet Cells" Diabetes, 2003, 52:1732-1737. cited by applicant . O'Hare, et al. "Transformation of mouse fibroblasts to methotrexate resistance by a recombinant plasmid expressing a prokaryotic dihydrofolate reductase" Proc. Natl. Acad. Sci., 1981, 78(3):1527-1531. cited by applicant . Park, et al. "Regulation of Ribosomal S6 Kinase 2 by Mammalian Target of Rapamycin", The Journal of Biological Chemistry, 2002, 277(35):31423-31429. cited by applicant . Perez-Pinera, et al. "Advances in Targeted Genome Editiong" Curr Opin Chem Biol., 2012, 16(3-4):268.277, doi:10.1016/j.cbpa.2012.06.007. cited by applicant . Planey, et al. "Mechanisms of Signal Transduction: Inhibition of Glucocorticoid-induced Apoptosis in 697 Pre-B Lymphocytes by the Mineralocorticoid Receptor N-terminal Domain", J. Biol. Chem., 2002, 277:42188-42196. cited by applicant . Posfai, et al. "Markerless gene replacement in Escherichia coli stimulated by a double-strand break in the chromosome"??, 1999, 27(22):4409-15. cited by applicant . Primo, et al. "Lentiviral vectors for cutaneous RNA managing" Experimental Dermatology, 2012, 21:162-170. cited by applicant . Radulovich, et al. "Modified gateway system for double shRNA expression and Cre/lox based gene expression" BMC Biotechnology, 2011, 11(24):1-9. cited by applicant . Roberts, et al. "The Effect of Protein Content on Nuclear Location Signal Function" Cell, 1987, 50:465-475. cited by applicant . Roberts, et al. "Nuclear location signal-mediated protein transport" Biochimica et Biophysica Acta, 1989, 1008:263-280. cited by applicant . Rodrigues, et al. "Red Fluorescent Protein (DsRed) as a Reporter in Saccharomyces cerevisiae" Journal of Bacteriology, 2001, 183(12):3791-3794. cited by applicant . Sambrook, et al., Molecular Cloning, A Laboratory Manual on the Web, 2001, Chapter 16. cited by applicant . Sanjana, et al. "A Transcription Activator-Like Effector (TALE) Toolbox for Genome Engineering" Nat. Protoc, 2011, 7(1):171-192, doi:10.1038/nprot.2011.431. cited by applicant . Sato, et al. "Generation of Adeno-Associated Virus Vector Enabling Functional Expressionof Oxytocin Receptor and Fluorescence Marker Genes Using the Human elF4G Internal Ribosome Entry Site Elemet" Biosci. Biotechno. Biochem, 2009, 73(9):2145-2148. cited by applicant . Sebo, et al. "A simplified and efficient germline-specific CRISPR/Cas9 system for Drosophila genomic engineering" Fly, 2014, 8(1):52-57. cited by applicant . Sharan, et al. "Recombineering: A Homologous Recombination-Based Method of Genetic Engineering" Nat. Protoc., 2009, 4(2):206-223, doi:10.1038/nprot.2008.227. cited by applicant . Shieh, et al. "Nuclear Targeting of the Maize R. Protein Requires Two Nuclear Localization Sequences" Plant Physiol, 1993, 101:353-361. cited by applicant . Siegl, et al. "I-Scel endonuclease: a new tool for DNA repair studies and genetic manipulations in streptomycetes" Appl Microbiol Bitotechnol, 2010, 87:1525-1532. cited by applicant . Joseph, et al. "Thermodynamic basis of selectivity in guide-target-mismatched RNA interference" NIH Public Access Author Manuscript 2012, 80(5):1283-1298, doi:10.1002/prot.24025. cited by applicant . A. Amsterdam et al., "Identification of 315 genes essential for early zebrafish development," proc Natl Acad Sci., vol. 101, Aug. 31, 2004, pp. 12792-12797, 6 pages. cited by applicant . A. Fire et al., "Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans," Nature, vol. 391, Feb. 19, 1998, pp. 806-811, 6 pages. cited by applicant . A. Subramanian et al., "Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles," Proc Natl Acad Sci., vol. 102, Oct. 25, 2005, pp. 15545-15550, 6 pages. cited by applicant . A.C. Spradling et al., "The Berkeley Drosophila Genome Project Gene Disruption Project: Single P-Element Insertions Mutating 25% of Vital Drosophila Genes," Genetics, vol. 153, Sep. 1999, pp. 135-177, 43 pages. cited by applicant . A.H. Tong et al., "Global mapping of the yeast genetic interaction network," Science, vol. 303, Feb. 6, 2004, pp. 808-813, 6 pages. cited by applicant . A.L. Lin and D.H Gutmann, "Advances in the treatment of neurofibromatosis-associated tumours," Nature, vol. 10, Nov. 2013, pp. 616-624, 9 pages. cited by applicant . A.P. Blanchard and L. Hood, "Sequence to array: probing the genome's secrets," Nat Biotechnol, vol. 14, Dec. 14, 1996, p. 1649. cited by applicant . Addgene, "gRNA_Cloning Vector", retrieved on Jan. 30, 2019, <https://www/addgenen.org/41824/>2 pages. cited by applicant . Allen, et al., "Liposomal drug delivery systems: From concept to clinical applications" Advanced Drug Delivery Reviews, vol. 65, 2013, pp. 36-48, 13 pages. cited by applicant . B. Langmead et al., "Ultrafast and memory-efficient alignment of short DNA sequences to the human genome," Genome biology, vol. 10, Mar. 4, 2009, 10 pages. cited by applicant . B.Langmead and S.L. Salzberg, "Fast gapped-read alignment with Bowtie 2," Nat Meth, vol. 9, 2012, pp. 357-359, 3 pages. cited by applicant . B.Scappini et al., "Changes associated with the development of resistance to imatinib (STI571) in two leukemia cell lines expressing p210 Bcr/Abl protein," Cancer, vol. 100, Apr. 1, 2004, pp. 1459-1471, 13 pages. cited by applicant . B.Sonnichsen et al., "Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans," Nature, vol. 434, Mar. 24, 2005, pp. 462-469, 8 pages. cited by applicant . Bae, T. and Schneewind, O. "Allelic replacement in Staphylococcus aureus with inducible counter-selection," Plasmid, vol. 55, 2006, pp. 58-63, 6 pages. cited by applicant . Botta, S. et al, "Transcriptional Repression with Zinc-Finger and Tale Protein Scaffold", Molecular Therapy, 2013, Supplement 1, p. S208, Abstract No. 539. cited by applicant . Brummelkamp TR et al., "A system for stable expression of short interfering RNAs in mammalian cells," Science, vol. 296, Apr. 19, 2002, pp. 550-553. cited by applicant . C. Cayrol et al., "The THAP-zinc finger protein THAP1 regulates endothelial cell proliferation through modulation of pRB/E2F cell-cycle target genes," Blood, vol. 109, 2007, pp. 584-594. cited by applicant . C. Trapnell et al., "Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks," Nature protocols, vol. 7, 2012, p. 562. cited by applicant . C. Trapnell et al., "TopHat: discovering splice junctions with RNA-Seq.," Bioinformatics, vol. 25, 2009, pp. 1105-1111. cited by applicant . C.J, Echeverri et al., "Minimizing the risk of reporting false positives in large-scale RNAi screens," Nature methods, vol. 3, Oct. 2006, p. 777. cited by applicant . C.M Johannessen et al., "COT drives resistance to RAF inhibition through MAP kinase pathway reactivation," Nature, vol. 468, Dec. 16, 2010, p. 968. cited by applicant . C.M. Johnston et al., "Large-scale population study of human cell lines indicate that dosage compensation is virtually complete," PLoS Genet., vol. 4, Jan. 2008, pp. 88-98, 11 pages. cited by applicant . Carte, J., et al., "Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes," Genes Dev., vol. 22, 2008, pp. 3489-3496. cited by applicant . Chadderton, N., et al., "Improved Retinal Function in a Mouse Model of Dominant Retinitis Pigmentosa Following AAV-delivered Gene Therapy", Molecular Therapy, vol. 17, Apr. 2009, pp. 593-599. cited by applicant . Chevalier et al., "Homing endonuclease: structural and functional insight into the catalysts of intron/intein mobility," Oxford University Press., vol. 29, 2001, pp. 3757-3774. cited by applicant . Cho, Minseon, et al., "Quantitative selection and parallel characterization of aptamers," PNAS, vol. 110, Nov. 12, 2013, pp. 18460-18465. cited by applicant . D.J.Burgess et al., "Topoisomerase levels determine chemotherapy response in vitro and in vivo," Proceedings of the National Academy of Sciences, vol. 105, Jul. 1, 2008, pp. 9053-9058. cited by applicant . Dean., "Recent Advances in Drug Design Methods: Where Will They Lead?", BioEssays, vol. 16, Sep. 1994, pp. 683-687. cited by applicant . Deveau, H. et al., "Phage response to CRISPR-encoded resistance in Streptococcus thermophilus," Journal of Bacteriology, vol. 190, Feb. 2008, pp. 1390-1400. cited by applicant . Deveau, H., et al., "CRISPR/Cas system and its role in phage-bacteria interactions," Annu. Rev. Microbiol., vol. 64, 2010, pp. 475-493. cited by applicant . E.S. Lander, "Initial impact of the sequencing of the human genome," Nature, vol. 470, Feb. 10, 2011, p. 187-197. cited by applicant . Edgar, R. and Qimron, U., "The Escherichia coli CRISPR system protects from .LAMBDA. lysogenization, lysogens, and prophage induction," Journal of Bacteriology, vol. 192, Dec. 2010, pp. 6291-6294. cited by applicant . Esvelt et. al., "Orthogonal Cas9 proteins for RNA-guided gene regulation and editing", Nature Methods, vol. 10, No. 11, Nov. 2013 (available online Sep. 29, 2013), pp. 1116-1123. cited by applicant . Fischer, S. et al., "An archaeal immune system can detect multiple Protospacer Adjacent Motifs (PAMs) to target invader DNA," J. Biol. Chem., vol. 287, Sep. 28, 2012, pp. 33351-33363. cited by applicant . Flannery, J. G., "Ribozyme-Mediated Gene Therapy for Autosomal Dominant Retinal Degeneration", Retinal Degenerative Diseases and Experimental Therapy, 1999, pp. 277-291. cited by applicant . G. Giaever et al., "Functional profiling of the Saccharomyces cerevisiae genome," Nature, vol. 418, Jul. 25, 2002, pp. 387-391. cited by applicant . G. Guo et al., "Mismatch repair genes identified using genetic screens in Blm-deficient embryonic stem cells," Nature, vol. 429, Jun. 24, 2004, p. 891. cited by applicant . Gao, et al., "A Sustained, Cytoplasmic Transgene Expression System delivered by Cationic Liposomes", Biochemical and Biophysical Research Communications, vol. 200, May 16, 1994, pp. 1201-1206. cited by applicant . GenBank: "CRISPR-associated protein Cas9/Csn1 [Staphylococcus aureus subsp. aureus]", GenBank: CCK74173.1, Year: 2012, http://www.ncbi.nlm.nih.gov/protein/403411236?sat=16&satkey=13804560, dated Dec. 14, 2016, 2 pages. cited by applicant . Gibson, D.G. et al., "Enzymatic assembly of DNA molecules up to several hundred kilobases," Nat methods, vol. 6, 2009, pp. 343-345. cited by applicant . Greenwald, D L, et al., "Engineered Zinc Finger Nuclease-Mediated Homologous Recombination of the Human Rhodopsin Gene", Investigative Ophthalmology & Visual Science, vol. 51, Dec. 2010, pp. 6374-6380. cited by applicant . Grosse, et al. "Meganuclease-medicated Inhibition of HSV1 Infection in Cultured Cells", Molecular Therapy, vol. 19, No. 4, Apr. 1, 2011, pp. 694-702. cited by applicant . Gudbergsdottir, S. et al., "Dynamic properties of the Sulfolobus CRISPR/Cas and CRISPR/Cmr systems when challenged with vector-borne viral and plasmid genes and protospacers," Mol. Microbiology, vol. 79, 2011, pp. 35-49. cited by applicant . H. Davies et al., "Mutations of the BRAF gene in human cancer," Nature, vol. 417, Jun. 27, 2002, p. 949-954. cited by applicant . H.W Cheung et al., "Systematic investigation of genetic vulnerabilities across cancer cell lines reveals lineage-specific dependencies in ovarian cancer," Proceedings of the National Academy of Sciences, vol. 108, Jul. 26, 2011, p. 12372-12377. cited by applicant . Hatoum-Aslan, A., et al., "Mature clustered, regularly interspaced, short palindromic repeats RNA (crRNA) length is measured by a ruler mechanism anchored at the precursor processing site," Proc. Natl. Acad. Sci., vol. 108, Dec. 27, 2011, pp. 21218-21222. cited by applicant . Haurwitz, R.E., et al., "Sequence- and structure-specific RNA processing by a CRISPR endonuclease," Science, vol. 329, 2010, pp. 1355-1358. cited by applicant . Havarstein, L.S., et al., "An unmodified heptadecapeptide pheromone induces competence for genetic transformation in Streptococcus pneumoniae, " Proc. Natl. Acad. Sci., vol. 92, Nov. 1995, pp. 11140-11144. cited by applicant . Hemann et al., "An epi-allelic series of p53 hypomorphs created by stable RNAi produces distinct tumor phenotypes in vivo," Nat Genetics, vol. 33, Mar. 2003, pp. 396-400. cited by applicant . Horinouchi, S. and Weisblum, B., "Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance," J. Bacteriology, vol. 150, May 1982, pp. 815-825. cited by applicant . Horton, R.M., "In Vitro recombination and Mutagenesis of DNA: SOEing Together Tailor-Made Genes," Methods Mol. Biology, vol. 15, 1993, pp. 251-261. cited by applicant . Horvath, P. and Barrangou, R. "CRISPR/Cas, the immune system of bacteria and archaea," Science, vol. 327, Jan. 8, 2010, pp. 167-170. cited by applicant . Hosaka, T. et al., "The novel mutation K87E in ribosomal protein S12 enhances protein synthesis activity during the late growth phase in Escherichia coli." Mol. Gen. Genomics, vol. 271, 2004, pp. 317-324. cited by applicant . Hoskins, J. et al., "Genome of the bacterium Streptococcus pneumoniae strain R6," Journal of Bacteriology, vol. 183, Oct. 2001, pp. 5709-5717. cited by applicant . Husmann, L.K.,et al., "Expression of the Arp protein, a member of the M protein family, is not sufficient to inhibit phagocytosis of Streptococcus pyogenes," Infection and immunity, vol. 63, Jan. 1995, pp. 345-348. cited by applicant . Hwang, W.Y., et al., "Efficient Genome Editing in Zebrafish Using a CRISPR-Cas System", Nature Biotechnology, vol. 31, No. 3, Jan. 29, 2013, pp. 227-229. cited by applicant . Ishino Y. et al., "Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product," J. Bacteriology, vol. 169, Dec. 1987, pp. 5429-5433, 5 pages. cited by applicant . J. Merkin et al., "Evolutionary dynamics of gene and isoform regulation in Mammalian tissues," Science, vol. 338, Dec. 21, 2012, p. 1593-1599, 7 pages. Includes Supplementary Information, 34 pages. cited by applicant . J.E. Carette et al., "Haploid genetic screens in human cells identify host factors used by pathogens," Science, vol. 326, Nov. 27, 2009, p. 1231-1235, 5 pages. cited by applicant . J.F. Rual et al., "Toward Improving Caenorhabditis elegans Phenome Mapping with an ORFeome-Based RNAi Library," Genome Research, vol. 14, 2004, pp. 2162-2168, 7 pages. cited by applicant . J.M. Engreitz et al., "The Xist IncRNA exploits three-dimensional genome architecture to spread across the X chromosome," Science, vol. 341, Aug. 16, 2013, pp. 1-8, 8 pages. cited by applicant . Jansen R. et al., "Identification of genes that are associated with DNA repeats in prokaryotes," Molecular Microbiology, vol. 43, 2002, pp. 1565-1575, 11 pages. cited by applicant . JL. Mummery-Widmer et al., "Genome-wide analysis of Notch signalling in Drosophila by transgenic RNAi," Nature, vol. 458, Apr. 23, 2009, pp. 987-992, 6 pages. Includes Supplementary information, 2 pages. cited by applicant . K. Yoshimoto et al., "Complex DNA repair pathways as possible therapeutic targets to overcome temozolomide resistance in glioblastoma," Front Oncology, vol. 2, Dec. 2012, pp. 1-8, 8 pages. cited by applicant . K.T Flaherty et al., "Inhibition of mutated, activated BRAF in metastatic melanoma," The New England Journal of Medicine, vol. 363, Aug. 26, 2010, pp. 1-22, 22 pages. cited by applicant . Kim et al., "High Cleavage Efficiency of a 2A Peptide Derived from Porcine Teschovirus-1 in Human Cell Lines, Zebrafish and Mice," PLoS One, vol. 6, Apr. 2011, pp. 1-8, 8 pages. cited by applicant . Koike-Yusa, H., et al., "Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library," Nat Biotechnology, vol. 32, Mar. 2014, pp. 267-273, 7 pages. Including Supplemental information, 3 pages. doi:10.1038/nbt.2800. cited by applicant . Koo et al., "Measuring and Reducing Off-Target Activities of Programmable Nucleases Including CRISPR-Cas9", Molecules and Cells, vol. 38, 2015, pp. 475-481, 7 pages. cited by applicant . Laganiere et. al., "An Engineered Zinc Finger Protein Activator of the Endogenous Glial Cell Line-Derived Neurotrophic Factor Gene Provides Functional Neuroprotection in a Rat Model of Parkinson's Disease", The Journal of Neuroscience, vol. 30, Dec. 8, 2010, pp. 16469-16474, 6 pages. cited by applicant . M. Booker et al., "False negative rates in Drosophila cell-based RNAi screens: a case study," BMC Genomics, vol. 12, 2011, pp. 1-11, 11 pages. cited by applicant . M. Costanzo et al., "The genetic landscape of a cell," Science, vol. 327, Jan. 22, 2010, pp. 425-431, 8 pages. cited by applicant . Madisen et al., "A robust and high-throughput Cre reporting and characterization system for the whole mouse brain," Nat. Neuroscience, vol. 13, Jan. 2010, pp. 133-140, 8 pages. cited by applicant . Marraffini, L.A., et al., "Sortases and the art of anchoring proteins to the envelopes of gram-positive bacteria," Microbiol. Mol. Biology Review vol. 70, Mar. 2006, pp. 192-221, 3 pages. cited by applicant . Martin, M., "Cutadapt removes adapter sequences from high-throughput sequencing reads," EMBnet.journal, vol. 17, 2011, pp. 10-12, 3 pages. cited by applicant . Moffat J et al., "A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen," Cell, vol. 124, Mar. 24, 2006, pp. 1283-1298, 16 pages. cited by applicant . Mojica F. J. M et al., "Biological significance of a family of regularly spaced repeats in the genomes of Archaea, Bacteria and mitochondria," Molecular Microbiology, vol. 36, 2000, pp. 244-246, 3 pages. cited by applicant . Motamedi, M.R., et al., "Double-strand-break repair recombination in Escherichia coli: physical evidence for a DNA replication mechanism in vivo," Genes Dev., vol. 13, 1999, pp. 2889-2903. cited by applicant . Musunuru, "Abstract 18593: Use of a CRISPR/Cas System for Cardiovascular Disease Modeling and Therapeutic Applications", Circulation, vol. 128, 2013, 4 pages (Meeting info: American Heart Association, 2013 Scientific Sessions and Resuscitation Science Symposium, Dallas, TX, US, Nov. 16-20, 2013). cited by applicant . Paddison et al., "A resource for large-scale RNA-interference-based screens in mammals," Nature, vol. 428, Mar. 25, 2004, pp. 427-431, 5 pages. cited by applicant . Podbielski, A., et al., "R. Novel series of plasmid vectors for gene inactivation and expression analysis in group A streptococci (GAS)," Gene, vol. 177, 1996, pp. 137-147, 11 pages. cited by applicant . R. Rad et al., "PiggyBac transposon mutagenesis: a tool for cancer gene discovery in mice," Science, vol. 330, Nov. 19, 2010, p. 1104-1107, 4 pages. cited by applicant . R.D Kolodner and G.T. Marsischky, "Eukaryotic DNA mismatch repair," Current Opinion in Genetics and Development, vol. 9, 1999, p. 89-96, 8 pages. cited by applicant . R.Renella et al., "Codanin-1 mutations in congenital dyserthropoietic anemia type 1 affect HP1.alpha. localization in erythroblasts," Blood, vol. 117, Jun. 2011, pp. 6928-6938, 11 pages. cited by applicant . Response to Third Party Observations in EP No. 13824232.6 filed Oct. 2, 2014, with Redlined and Clean Amended Claims, 14 pages. cited by applicant . S. Huang et al., "MED12 Controls the Response to Multiple Cancer Drugs through Regulation of TGF-.beta.; Receptor Signaling," Cell, vol. 151, 2012, pp. 937-950, 14 pages. cited by applicant . S. Konermann et al., "Optical control of mammalian endogenous transcription and epigenetic states," Nature, vol. 500, Aug. 22, 2013, pp. 472-476, 5 pages. Includes Supplemental Information, 13 pages. cited by applicant . S.H. Chen et al., "A Knockout Mouse Approach Reveals that TCTP Functions as an Essential Factor for Cell Proliferation and Survival in a Tissue- or Cell Type-specific Manner," Molecular Biology of the Cell, vol. 18, Jul. 2007, pp. 2525-2532, 8 pages. cited by applicant . S.R. Whittaker et al., "A Genome-Scale RNA Interference Screen Implicates NF1 Loss in Resistance to RAF Inhibition," Cancer Discovery, vol. 3, 2013, pp. 350-362, 14 pages. cited by applicant . S.S. Liu et al., "Identification and characterization of a novel gene, clorf109, encoding a CK2 substrate that is involved in cancer cell proliferation," Journal of Biomedical Science, vol. 19, 2012, 12 pages. cited by applicant . S.Xue and M. Barna, "Specialized ribosomes: a new frontier in gene regulation and organismal biology," Nat Rev Mol Cell Biology, vol. 13, Jun. 2012. pp. 355-369, 15 pages. cited by applicant . Sarra, G., et al., "Gene replacement therapy in the retinal degeneration slow (rds) mouse: the effect on retinal degeneration following partial transduction of the retina", Human Molecular Genetics, vol. 10, 2001, pp. 2353-2361, 9 pages. cited by applicant . Schiffer, et al. "Targeted DNA Mutagenesis for the Cure of Chronic Viral Infections" Journal of Virology, vol. 86, No. 17, Jun. 20, 2012, pp. 8920-8936. cited by applicant . Semenova, E. et al., "Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence," Proc. Natl. Acad. Sci., vol. 108, Jun. 21, 2011, pp. 10089-10103, 7 pages. cited by applicant . Spencer, J.M., et al., "Development of a Nuclease Screen to Improve Cas9 Targeting Specificity", Molecular Therapy, May 2015, vol. 23, Suppl. 1, S136(340). cited by applicant . Stewart SA et al., "Lentivirus-delivered stable gene silencing by RNAi in primary cells," RNA, vol. 9, 2003, pp. 493-501, 9 pages. cited by applicant . Swiech et al., "CRISPR-Mediated Genome Editing in the Mammalian Brain", Molecular Therapy, 747, vol. 22, 2014, p. S289. cited by applicant . Symington et al., "Double-Strand Break End Resection and Repair Pathway Choice", Annual Review of Genetics, vol. 45, 2011, pp. 247-271, 25 pages. cited by applicant . T. Horii et al., "Generation of an ICF Syndrome Model by Efficient Genome Editing of Human Induced Pluripotent Stem Cells Using the CRISPR System," International Journal of Molecular Sciences, vol. 14, 2013, p. 19774-19781, 9 pages. cited by applicant . T.J. Cradick et al., "CRISPR/Cas9 systems targeting .beta.-globin and CCR5 genes have substantial off-target activity," Nucleic Acids Research, vol. 41, 2013, 9584-9592, 9 pages. cited by applicant . T.Yan et al., "DNA mismatch repair (MMR) mediates 6-thioguanine genetoxicity by introducing single-strand breaks to signal a G2-M arrest in MMR-proficient RKO cells," Clinical Cancer Research, vol. 9, Jun. 2003, p. 2327-2334, 9 pages. cited by applicant . Takara Bio USA, Inc., "Lenti-X.TM. Tet-on .COPYRGT. 3G CRISPR/Cas9 System User Manual", 2016, pp. 1-35. cited by applicant . Third Party Observation for Application No. EP20130824232 dated Sep. 22, 2014, 19 pages. cited by applicant . Third Party Observation in Application No. PCT/US2013/074819 dated Apr. 10, 2015, 10 pages. cited by applicant . Third Party Observation Under Article 115 EPC in Application No. 13818570.7 dated Oct. 1, 2014. cited by applicant . Third-Party Observation for Application No. EP20130824232 dated Sep. 8, 2014, 47 pages. cited by applicant . V.N. Ngo et al., "A loss-of-function RNA interference screen for molecular targets in cancer," Nature, vol. 441, May 4, 2006, pp. 106-110, 5 pages. cited by applicant . Van Der Oost, J., et al., "CRISPR-based adaptive and heritable immunity in prokaryotes," Trends. Biochem. Sci., vol. 34, 2009, pp. 401-407, 7 pages. cited by applicant . Venken et al., "P[acman]: A BAC Transgenic Platform for Targeted Insertion of Large DNA Fragments in D. melanogaster", Science, vol. 314, Dec. 15, 2006, pp. 1747-1751, 5 pages. cited by applicant . W.G. Kaelin., "Use and Abuse of RNAi to Study Mammalian Gene Function," Science, vol. 337, Jul. 27, 2012, p. 421-422, 2 pages. cited by applicant . Wang, H.H. et al., "Genome-scale promoter engineering by coselection MAGE," Nat methods, vol. 9, Jun. 2012, pp. 591-593, 3 pages. cited by applicant . Wayengera, M., "Identity of zinc finger nucleases with specificity to herpes simplex virus type II genomic DNA; novel HSV-2 vaccine/therapy precursors", Theoretical Biology and Medical Modelling, vol. 8, No. 1, Jun. 24, 2011, p. 23. cited by applicant . Wayengera, M., "Zinc finger arrays binding human papillomavirus types 16 and 18 genomic DNA: precursors of gene-therapeutics for in-situ reversal of associated cervical neoplasia", Theoretical Biology and Medical Modeling, vol. 9, No. 1, Jul. 28, 2012, p. 30. cited by applicant . Weber et al., "TALENs Targeting HBV: Designer Endonuclease Therapies for Viral Infections", Molecular Therapy, vol. 21, Oct. 2013, pp. 1819-1821, 3 pages. cited by applicant . Wiedenheft, B. et al., "RNA-guided complex from a bacterial immune system enhances target recognition through seed sequence interactions," Proc. Natl. Acad. Sci., vol. 108, Jun. 21, 2011, 10092-10097, 7 pages. cited by applicant . X.Liu et al., "STAGA recruits Mediator to the MYC oncoprotein to stimulate transcription and cell proliferation," Molecular and cellular biology, vol. 28, Jan. 2008, p. 108-121, 14 pages. cited by applicant . Xiao, W., et al, "Gene Therapy Vectors Based on Adeno-Associated Virus Type 1", Journal of Virology, May 1999, vol. 73, No. 5, p. 3994-4003. cited by applicant . Xu, Zhi-Li et al., "Regulated gene expression from adenovirus vectors: a systematic comparison of various inducible systems," Gene, vol. 309, 2003, pp. 145-151, 7 pages. cited by applicant . Yang et al., "HIV-1 TAT-mediated protein transduction and subcellular localization using novel expression vectors," FEBS Letters, vol. 532, 2012, pp. 36-44, 9 pages. cited by applicant . Zahner, D. and Hakenbeck, R. "The Streptococcus pneumoniae beta-galactosidase is a surface protein," J. Bacteriology, vol. 182, Oct. 2000, pp. 5919-5921, 3 pages. cited by applicant . Zeng Y et al., "Both natural and designed micro RNAs can inhibit the expression of cognate mRNAs when expressed in human cells," Mol Cell., vol. 9, Jun. 2002, pp. 1327-1333, 7 pages. cited by applicant . Zhang, "Processing-Independent CRISPR RNAs Limit Natural Transformation in Neisseria meningitidis", Molecular Cell, vol. 50, May 23, 2013 pp. 488-503. cited by applicant . Zuris, et al., "Cationic lipid-mediated delivery proteins enables efficient protein-based genome editing in vitro and in vivo", Nature Biotechnology, vol. 33, No. 1, Jan. 2015, pp. 73-80. cited by applicant . Zuris, et al., "Efficient Delivery of Genome-Editing Proteins in Vitro and in Vivo" Nature Biotechnology, vol. 33, No. 1, Jan. 2015, pp. 1-26. cited by applicant . Zuris, et al., Supplementary Information--"Cationic lipid-mediated delivery proteins enables efficient protein-based genome editing in vitro and in vivo" Nature Biotechnology, vol. 33, No. 1, Jan. 2015, pp. 1-49. doi:10.1038/nbt.3081. cited by applicant . U.S. Appl. No. 14/054,414, filed Oct. 15, 2013. cited by applicant . U.S. Appl. No. 14/104,837, filed Dec. 12, 2013. cited by applicant . U.S. Appl. No. 14/104,900, filed Dec. 12, 2013. cited by applicant . U.S. Appl. No. 14/104,977, filed Dec. 12, 2013. cited by applicant . U.S. Appl. No. 14/104,990, filed Dec. 12, 2013. cited by applicant . U.S. Appl. No. 14/105,017, filed Dec. 12, 2013. cited by applicant . U.S. Appl. No. 14/105,031, filed Dec. 12, 2013. cited by applicant . U.S. Appl. No. 14/105,035, filed Dec. 12, 2013. cited by applicant . U.S. Appl. No. 14/183,429, filed Feb. 18, 2014. cited by applicant . U.S. Appl. No. 14/183,471, filed Feb. 18, 2014. cited by applicant . U.S. Appl. No. 14/183,486, filed Feb. 18, 2014. cited by applicant . U.S. Appl. No. 14/183,512, filed Feb. 18, 2014. cited by applicant . U.S. Appl. No. 14/222,930, filed Mar. 24, 2014. cited by applicant . U.S. Appl. No. 14/226,274, filed Mar. 26, 2014. cited by applicant . U.S. Appl. No. 14/256,912, filed Apr. 18, 2014. cited by applicant . U.S. Appl. No. 14/258,458, filed Apr. 22, 2014. cited by applicant . U.S. Appl. No. 14/259,420, filed Apr. 23, 2014. cited by applicant . U.S. Appl. No. 14/290,575, filed May 29, 2014. cited by applicant . U.S. Appl. No. 14/293,498, filed Jun. 2, 2014. cited by applicant . U.S. Appl. No. 14/293,674, filed Jun. 2, 2014. cited by applicant . U.S. Appl. No. 14/324,960, filed Jul. 7, 2014. cited by applicant . U.S. Appl. No. 14/463,253, filed Aug. 19, 2014. cited by applicant . U.S. Appl. No. 14/481,339, filed Sep. 9, 2014. cited by applicant . U.S. Appl. No. 14/497,627, filed Sep. 26, 2014. cited by applicant . U.S. Appl. No. 14/523,799, filed Oct. 24, 2014. cited by applicant . U.S. Appl. No. 14/681,382, filed Apr. 8, 2015. cited by applicant . U.S. Appl. No. 14/703,511, filed May 4, 2015. cited by applicant . U.S. Appl. No. 14/704,551, filed May 5, 2015. cited by applicant . U.S. Appl. No. 14/705,719, filed May 6, 2015. cited by applicant . U.S. Appl. No. 14/738,398, filed Jun. 12, 2015. cited by applicant . U.S. Appl. No. 14/738,483, filed Jun. 12, 2015. cited by applicant . U.S. Appl. No. 14/970,967, filed Dec. 16, 2015. cited by applicant . U.S. Appl. No. 14/971,169, filed Dec. 16, 2015. cited by applicant . U.S. Appl. No. 14/971,356, filed Dec. 16, 2015. cited by applicant . U.S. Appl. No. 14/972,523, filed Dec. 17, 2015. cited by applicant . U.S. Appl. No. 14/972,927, filed Dec. 17, 2015. cited by applicant . U.S. Appl. No. 14/973,062, filed Dec. 17, 2015. cited by applicant . U.S. Appl. No. 14/990,444, filed Jan. 7, 2016. cited by applicant . U.S. Appl. No. 14/991,083, filed Jan. 8, 2016. cited by applicant . U.S. Appl. No. 15/160,710, filed May 20, 2016. cited by applicant . U.S. Appl. No. 15/171,141, filed Jun. 2, 2016. cited by applicant . U.S. Appl. No. 15/172,636, filed Jun. 3, 2016. cited by applicant . U.S. Appl. No. 15/179,711, filed Jun. 10, 2016. cited by applicant . U.S. Appl. No. 15/179,799, filed Jun. 10, 2016. cited by applicant . U.S. Appl. No. 15/179,912, filed Jun. 10, 2016. cited by applicant . U.S. Appl. No. 15/179,938, filed Jun. 10, 2016. cited by applicant . U.S. Appl. No. 15/179,941, filed Jun. 10, 2016. cited by applicant . U.S. Appl. No. 15/217,489, filed Jul. 22, 2016. cited by applicant . U.S. Appl. No. 15/229,702, filed Aug. 5, 2016. cited by applicant . U.S. Appl. No. 15/230,025, filed Aug. 5, 2016. cited by applicant . U.S. Appl. No. 15/230,161, filed Aug. 5, 2016. cited by applicant . U.S. Appl. No. 15/330,876, filed Nov. 7, 2016. cited by applicant . U.S. Appl. No. 15/349,603, filed Nov. 11, 2016. cited by applicant . U.S. Appl. No. 15/430,260, filed Feb. 10, 2017. cited by applicant . U.S. Appl. No. 15/436,396, filed Feb. 17, 2017. cited by applicant . U.S. Appl. No. 15/620,098, filed Jun. 12, 2017. cited by applicant . U.S. Appl. No. 15/620,391, filed Jun. 12, 2017. cited by applicant . U.S. Appl. No. 15/633,126, filed Jun. 26, 2017. cited by applicant . U.S. Appl. No. 15/834,736, filed Dec. 7, 2017. cited by applicant . U.S. Appl. No. 15/838,064, filed Dec. 11, 2017. cited by applicant . U.S. Appl. No. 15/838,720, filed Dec. 12, 2017. cited by applicant . U.S. Appl. No. 15/844,528, filed Dec. 16, 2017. cited by applicant . U.S. Appl. No. 15/887,377, filed Feb. 2, 2018. cited by applicant . U.S. Appl. No. 15/967,464, filed Apr. 30, 2018. cited by applicant . U.S. Appl. No. 15/967,495, filed Apr. 30, 2018. cited by applicant . U.S. Appl. No. 15/967,510, filed Apr. 30, 2018. cited by applicant . U.S. Appl. No. 16/012,692, filed Jun. 19, 2018. cited by applicant . U.S. Appl. No. 16/158,295, filed Oct. 11, 2018. cited by applicant . U.S. Appl. No. 16/177,403, filed Oct. 31, 2018. cited by applicant . U.S. Appl. No. 16/178,551, filed Nov. 1, 2018. cited by applicant . U.S. Appl. No. 16/262,905, filed Jan. 30, 2019. cited by applicant . U.S. Appl. No. 61/748,427, filed Jan. 2, 2012, Zhang. cited by applicant . Abudayyeh, et al. "C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector" Science, 2016, 10.1126/science.aaf5573 (2016). cited by applicant . Bauer, et al., "An Erythroid Enhancer of BCL11A Subject to Genetic Variation Determines Fetal Hemoglobin Level" Science, 2013, 342:253-257. cited by applicant . Bassett, et al. "A Genome-Wide CRISPR Library for High-Throughput Genetic Screening in Drosophila Cells" Journal of Genetics and Genomics, 2015, 42:301-309. cited by applicant . Canver, et al. "BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis" Nature, 2015, 527:192-197, including Supplementary material. cited by applicant . "CRISPR-associated protein" UniProtKB/TrEMBL, Accession No. Q73QW6, uploaded Nov. 28, 2012, retrieved on Nov. 22, 2017, http://www.uniprot.org/uniprot/Q73Qw6.txt?version4. cited by applicant . CRISPR-associated protein, Csn1 family, UniProtKB/TrEMBL, Accession No. D0W2Z9, uploaded Oct. 3, 2012, retrieved Nov. 22, 2017, http://www.uniprot.org/uniprot/D0W2Z9.txt?version=4. cited by applicant . CRISPR-associated protein, Csn2 family [Staphylococcus pseudintermedius ED99] GenBank, Accession No. ADX75954, uploaded on Nov. 21, 2011, retrieved on Nov. 22, 2017, http://www.uniprot.org/uniprot/G1UFN3.txt?version=3. cited by applicant . "Crispr genome engineering" XP055167591, Oct. 5, 2013, https://web.archive.org/web/2013100500 [retrieved on Feb. 5, 2015. cited by applicant . "Csn1 family CRISPR-associated protein" UniProtKB/TrEMBL, Accession No. J3TRJ9, uploaded Oct. 31, 2012, retrieved on Dec. 14, 2017, http://www.uniprot.org/uniprot/J3TRJ9.txt?version=2. cited by applicant . Doench, et al., "Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation" Nature Biotechnology, 2014, 32(12):1262-1267, including Supplementary material. cited by applicant . Dong, et al. "The crystal structure of Cpf1 in complex with CRISPR RNA" Nature, 2016, 532:523-525, including Research Letter, doi:10.1038/nature17944. cited by applicant . Gao, et al. "Engineered Cpf1 Enzymes with Altered PAM Specificities" BioRxiv Preprint, XP-002769442, 2016, doi: http://dx.doi.org/10.1101/091611, 1-13, including Figure Legends. cited by applicant . Gao, et al. "Engineered Cpf1 variants with altered PAM specificities" Nature Biotechnology, 2017, 1-4, doi:10.1038/nbt.3900, advanced online publication including Supplementary Information. cited by applicant . Grissa, et al. "CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats" Nucleic Acids Research, 2007, 35:W52-W57, doi:10.1093/nar/gkm360. cited by applicant . Ho, et al. "Targeting non-coding RNAs with the CRISPR/Cas9 system in human cell lines" Nucleic Acids Research, 2015, 43(3):e17, doi:10.1093/narlgku1198. cited by applicant . Incontro, et al. "Efficient, Complete Deletion of Synaptic Proteins using CRISPR" Cell, Neuron, 2014, 83:1051-1057. cited by applicant . Kleinstiver, et al. "Engineered CRISPR-Cas9 nucleases with altered PAM specificities" Nature, 2015, 523:481-485, including Research Letter, doi:10.1038/nature14592. cited by applicant . Leenay, et al. "Identifying and Visualizing Functional PAM Diversity across CRISPR-Cas Systems" Molecular Cell, 2016, 62:137-147. cited by applicant . Li, Ping, et al. "Biallelic knockout of .alpha.-1,3 galactosyltransferase gene in porcine liver-derived cells using zing finger nucleases" Journal of Surgical Research, 2013, E39-E45. cited by applicant . Pattanayak, et al. "High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity" Nature Biotechnology, 2013, 31(9):839-843, including Supplementary Materials. cited by applicant . "Putative uncharacterized protein" UniProtKB/TrEMBL, Accession No. Q6NK13, uploaded Jun. 13, 2012, retrieved on Nov. 22, 2017, http://www.uniprot.org/uniprot/Q6NK13.txt?version=43. cited by applicant . "Putative CRISPR-associated protein", UniProtKB/TrEMBL, Accession No. Q0P897, uploaded Oct. 3, 2012, retrieved on Nov. 22, 2017, http://www.uniprot.org/uniprot/Q0P897.txt?version=28. cited by applicant . Putative CRISPR associated protein, UniProtKB/TrEMBL, Accession No. G1UFN3, uploaded Oct. 3, 2012, retrieved on Nov. 22, 2017, http://www.uniprot.org/uniprot/G1UFN3.txt?version=3. cited by applicant . Sander, et al. "CRISPR-Cas systems for editing, regulating and targeting genomes" Nature Biotechnology, 2014, 32:347-355. cited by applicant . Sanjana, et al. "Improved vectors and genome-wide libraries for CRISPR screening", HHS Public Access Author Manuscript, 2014, 11(8):783-784. cited by applicant . Sebastiani, et al. "BCL11A enhancer haplotypes and fetal hemoglobin in sickle cell anemia" Blood Cells, 2015, 54(3):1079-9796. cited by applicant . Shalem, et al. "High-throughput functional genomics using CRISP-Cas9" Nature Reviews Genetics, 2015, 16(5):1471-0056. cited by applicant . Trafton, et al. "CRISPR-carrying nanoparticles edit the genome" MIT News, Nov. 13, 2017, http://news.mit.edu/2017/crispr-carrying-nanoparticles-edit-genome-1113. cited by applicant . Type V CRISPR-associated protein Cpfi [Acidaminococcus sp. Bv3L6], 2017, NCBI Reference Sequence: WP_02173622.1, Non-redundant Protein Sequence. cited by applicant . Van Nierop, et al. "Stimulation of homology-directed gene targeting at an endogenous human locus by a nicking endonuclease" Nucleic Acids Research, 2009, 37(17):5725-5736. cited by applicant . Wu, Yuxuan, et al. "Correction of a Genetic Disease in Mouse via Use of CRISPR--Cas9" Cell Stem Cell, 2013, 13:659-662. cited by applicant . Yamano, et al. "Crystal Structure of Cpf1 in Complex with Guide RNA and Target DNA" Cell, 2016, 165:949-962. cited by applicant . Zhou, et al. "High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells" Nature, 2014, 509:487-491. cited by applicant . Press Release dated Jan. 15, 2018 "The Rockefeller University and Broad Institute of MIT and Harvard announce update to CRISPR-Cas9 Portfolio filed by Broad" Copyright by Broad Institute. cited by applicant . U.S. Appl. No. 16/445,150, filed Jun. 18, 2019. cited by applicant . U.S. Appl. No. 16/445,156, filed Jun. 18, 2019. cited by applicant . U.S. Appl. No. 16/525,531, filed Jul. 29, 2019. cited by applicant . U.S. Appl. No. 16/532,442, filed Aug. 5, 2019. cited by applicant . U.S. Appl. No. 16/535,043, filed Aug. 7, 2019. cited by applicant . U.S. Appl. No. 16/800,988, filed Feb. 25, 2020. cited by applicant . U.S. Appl. No. 16/844,548, filed Apr. 9, 2020. cited by applicant . U.S. Appl. No. 16/906,580, filed Jun. 19, 2020. cited by applicant . U.S. Appl. No. 16/938,110, filed Jul. 24, 2020. cited by applicant . U.S. Appl. No. 61,769,046, filed Feb. 25, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/835,931, filed Jun. 17, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/757,972, filed Jan. 29, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/768,959, filed Feb. 25, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/791,409, filed Mar. 15, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/717,324, filed Oct. 23, 2012, Seung Woo Kim. cited by applicant . U.S. Appl. No. 61/803,599, filed Mar. 20, 2013, Jin-Soo Kim. cited by applicant . U.S. Appl. No. 61/837,481, filed Jun. 20, 2013, Seung Woo Cho. cited by applicant . U.S. Appl. No. 61/765,576, filed Feb. 15, 2013, Wendell Lim. cited by applicant . U.S. Appl. No. 61/716,256, filed Oct. 10, 2012, Jinek. cited by applicant . U.S. Appl. No. 61/758,468, filed Jan. 30, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/802,174, filed Mar. 15, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/806,375, filed Mar. 28, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/814,263, filed Apr. 20, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/819,803, filed May 6, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/828,130, filed May 28, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/836,127, filed Jun. 17, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/842,322, filed Jul. 2, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/836,101, filed Jun. 17, 2013, Zhang. cited by applicant . U.S. Appl. No. 61/835,936, filed Jun. 17, 2013, Zhang. cited by applicant . Alberts, et al., Molecular Biology of the Cell, fourth edition, 2002, 671-676. cited by applicant . Au, et al. "Characterization of a baculovirus nuclear localization signal domain in the late express factor 3 protein" Virology, 2009, 385:209-217. cited by applicant . Ausubel, et al. "A Compendium of Methods from Current Protocols in Molecular Biology" Short Protocols in Molecular Biology, Fourth Edition, 1999, 9: 9-3-9-4. cited by applicant . Baiker, et al. "The Immediate-Early 63 Protein of Varicella-Zoster Virus: Analysis of Functional Domains Required for Replication in Vitro and for T-Cell and Skin Tropism in the SCIDhu Model in Vivo" Journal of Virology, 2004, 78(3):1181-1194. cited by applicant . Bassett, et al. "Highly Efficient Targeted Mutagenesis of Drosophila with the CRISPR/Cas9 System" Cell Reports, 2013, 4:220-228. cited by applicant . Beerli, et al. "Toward controlling gene expression at will: Specific regulation of the erbB-2/HER-2 promoter by using polydactyl zinc finger proteins constructed from modular building blocks", Proc. Natl. Acad. Sci., 1998, 95:14628-14633. cited by applicant . Beerli, et al. "Positive and negative regulation of endogenous genes by designed transcription factors:" PNAS, 2000, 97(4):1495-1500. cited by applicant . Beerli, et al. "Enginnering polydactyl zinc-finger transcription factors" Nature Biotechnology, 2002, 20:135-141. cited by applicant . Birch, et al. "Plant Transformation: Problems and Strategies for Practical Application" Annu. Rev. Plant Physiol. Plant Mol. Biol., 1997, 48:297-326. cited by applicant . Boden, et al. "Efficient Gene Transfer of HIV-1-Specific Short Hairpin RNA into Human Lymphocytic Cells Using Recombinant Adeno-associated Virus Vectors" Molecular Therapy, 2004, 9(3):396-402. cited by applicant . Bouard, et al. "Themed Section: Vector Design and Drug Delivery Review, Viral vectors: from virology to transgene expression" British Journal of Pharmacology, 2009, 157:153-165. cited by applicant . Brouns, "A Swiss Army Knife of Immunity" Science, 2012, 337:808-809, DOI:10.1126/science.1227253. cited by applicant . Campeau, et al. "A Versatile Viral System for Expression and Depletion of Proteins in Mammalian Cells" PLoS One, 2009, 4(8):e6529. cited by applicant . Carr, et al. "Genome engineering" Nature Biotechnology, 2009, 27(12):1151-1162. cited by applicant . Carroll, "Progress and prospects: Zinc-finger nucleases as gene therapy agents" Gene Therapy, 2008, 15:1463-1468. cited by applicant . Carroll, "Genome Engineering With Zing-Finger Nucleases" Genetics, 2011, 188:773.782. cited by applicant . Chan, et al. "Characterization of the Kinetochore Binding Domain of CENP-E Reveals Interactions with the Kinetochore Proteins CENP-F and hBuBR1" The Journal of Cell Biology, 1998, 143:49-63. cited by applicant . Chan, Wai-Ting, et al. "Toxin-Antitoxin Genes of the Gram-Positive Pathogen Streptococcus pneumoniae: So Few and Yet So Many" Microbiology and Molecular Biology Reviews, 2012, 76(4):773-791. cited by applicant . Chang, et al. "Genome editing with RNA-guided Cas9 nuclease in Zebrafish embryos" Cell Research, 2013, 23:465-472. cited by applicant . Chen, Fuqiang et al. "High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases" Nature Methods, 2011, 8(9):753-755, including Supplemental Online Methods. cited by applicant . Chylinksi, et al. "Classification and evolution of type II CRISPR-Cas systems" Nucleic Acids Research, 2014, 42(10):6091-6105, doi:10.1093lnarlgku241. cited by applicant . Chiu, et al. "Engineered GFP as a vital reporter in plants" Current Biology, 1996, 6(3):325-330. cited by applicant . Costantino, et al., "Enhanced levels of .lamda. Red-mediated recombinants in mismatch repair mutants" PNAS, 100 (26):15748-15753. cited by applicant . CRISPR-associated endonuclease Cas9; Oct. 21, 2012, XP002738511M, http://ibis/exam/dbfetch.jsp?id=uniprot:J7RUA5. cited by applicant . Dai, et al. "Genes:Structures and Regulation: The Transcription Factors GATA4 and dHAND Physically Interact to Synergistically Activate Cardiac Gene Expression through a p300-dependent Mechanism", J. Biol. Chem., 2002, 277:24390-24398. cited by applicant . DiCarlo, et al. "Genome engineering in Saccharomyces cerevisiae using CRISPTR-Cas systems", Nucleic Acids Research, 2013, 41(7):4336-4343, doi:10.1093/nar/gkt135. cited by applicant . Dingwall, et al. "The Nucleoplasmin Nuclear Location Sequence Is Larger and More Complex than That of SV-40 Large T Antigen" The Journal of Cell Biology, 1988, 107:841-849. cited by applicant . Do, et al. "Identification of multiple nuclear localization signals in murine EIf3, an ETS transcription factor" FEBS Letters, 2006, 580:1865-1871. cited by applicant . Dominguez, et al., "Beyond editing: repurposing CRISPR-Cas 9 for precision genome regulation and interrogation" Nat Rev Mol Cell Biol., 2016, 17(1):5-15, doi:10.1038/nrm.2015.2. cited by applicant . Ellis, Hilary, et al. "High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotids" PNAS, 2001, 98(12):6742-6746. cited by applicant . Espinoza, et al., "Characterization of the structure, function, and mechanism of B2 RNA, an ncRNA repressor of RNA polymerase II transcription" RNA, 2007, 13(4):583-596. cited by applicant . Fieck, et al. "Modifications of the E. coli Lac repressor for expression in eukaryotic cells: effects of nuclear signal sequences on protein activity and nuclear accumulation" Nucleic Acids Research, 1992, 20(7):1785-1791. cited by applicant . Fischer-Fantuzzi, et al. "Molecular and Cellular Biology" Mol. Cell. Biol., 1988, 8(12):5495, DOI:10.1128/MCB.8.12.5495. cited by applicant . "Fixes, extra genomes, and improvements to the CRISPR Design Tool" Google Groups, XP055167583, Oct. 21, 2013, URL:https://groups.google.com/forum/#!topic/crispr/g9Q8U1tNSis [retrieved on Feb. 5, 2015]. cited by applicant . Foecking, et al. "Powerful and versatile enhance-promoter unit for mammalian expression vectors" Gene, 1986, 101-105. cited by applicant . Freitas, et al. "Mechanisms and Signals for the Nuclear Import of Proteins" Current Genomics, 2009, 10:550-557. cited by applicant . Fu, et al. "High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells", Nature Biotechnology, 2013, 31(9):822-826. cited by applicant . Garcia-Bustos, et al. "Nuclear protein localization" Biochimica et Biophysica Acta, 1991, 1071:83-101. cited by applicant . Garg, et al. "Engineering synthetic TAL effectors with orthogonal target sites" Nucleic Acids Research, 2012, 40(15):7584-7595, doi:10.1093/nar/gks404. cited by applicant . Genbank, Accession No. HE980450--"Staphylococcus aureus subsp.aureus ORFX gene and pseudo SCCmec-SCC-SCCCRISPR element, strain M06/0171", http://www.ncbi.nlm.nih.gov/nuccore/HE980450, Retrieved Aug. 18, 2016. cited by applicant . Gomaa, et al. "Programmable removal of bacterial strains by use of genome-targeting CRISPR-Cas systems", MBio., 2014, 5(1):e00928-13. cited by applicant . Gratz, et al. "Genome Engineering of Drosophila with the CRISPR RNA-Guided Cas9 Nuclease" Genetics, 2013, 194:1029-1035. cited by applicant . Greenspan, et al. "Two Nuclear Location Signals in the Influenza Virus NS1 Nonstructural Protein" Journal of Virology, 1988, 62(8):3020-3026. cited by applicant . Greiger, et al. "Packaging Capacity of Adeno-Associated Virus Serotypes: Impact of Larger Genomes on Infectivity and Postentry Steps" Journal of Virology, 2005, 79(15): 9933-9944. cited by applicant . Guan, et al. "Heritable endogenous gene regulation in plants with designed polydactyl zinc finger transcription factors" PNAS, 2002, 99(20):13296-13301. cited by applicant . Haft, D.H., "HMM Summary Page: TIGR04330" 2012, XP-002757584, http://jcvi.org/cgi-bin/tigrfams/HmmReportPage.cgi?acc=TIGR04330. cited by applicant . Heintze, et al. "A CRISPR CASe for high-throughput silencing", Frontiers in Genetics, Oct. 2013, 4(193): DOI:10.3389/gfene.2013.00193. cited by applicant . Hicks, et al. "Protein Import Into the Nucleus: An Integrated View" Annu. Rev. Cell Dev. Biol., 1995, 11:155-88. cited by applicant . Hockemeyer, et al. "Highly efficient gene targeting of expressed and silent genes in human ESCs and iPSCs using zinc finger nucleases" Nat Biotechnol., 2009, 27(9):851-857, doi:10.1038/nbt.1562. cited by applicant . Holmes, "CRISPR Genome Engineering Resources" XP055167586, Oct. 2, 2013, https://groups.google/forum/#!top1c/crispr/5BpJj_Y3ylG [retreieved on Feb. 5, 2015]. cited by applicant . Decision on Motions--PTAB, The Regents of the University of California v. The Broad Institute, Inc., filed Sep. 10, 2020, in Patent Interference No. 106,115 (DK), 113 pages. cited by applicant . Redeclaration--PTAB, The Regents of the University of California v. The Broad Institute, Inc., filed Sep. 10, 2020, in Patent Interference No. 106,115 (DK), 3 pages. cited by applicant. |
Primary Examiner: Woitach; Joseph
Attorney, Agent or Firm: Foley & Lardner LLP
Government Interests
STATEMENT AS TO FEDERALLY SPONSORED RESEARCH
This invention was made with government support under Grant Nos. MH100706 and DK097768 awarded by the National Institutes of Health. The government has certain rights in the invention.
Parent Case Text
RELATED APPLICATIONS AND INCORPORATION BY REFERENCE
This application is a continuation-in-part of international patent application serial no. PCT/US2013/074812 filed Dec. 12, 2013 and published as WO2014093709, which claims priority to U.S. provisional patent application 61/836,080 entitled METHODS, SYSTEMS, AND APPARATUS FOR IDENTIFYING TARGET SEQUENCES FOR CAS ENZYMES OR CRISPR-CAS SYSTEMS FOR TARGET SEQUENCES AND CONVEYING RESULTS THEREOF filed on Jun. 17, 2013, and which also claims priority to US provisional patent applications 61/758,468; 61/769,046; 61/802,174; 61/806,375; 61/814,263; 61/819,803 and 61/828,130 each entitled ENGINEERING AND OPTIMIZATION OF SYSTEMS, METHODS AND COMPOSITIONS FOR SEQUENCE MANIPULATION, filed on Jan. 30, 2013; Feb. 25, 2013; Mar. 15, 2013; Mar. 28, 2013; Apr. 20, 2013; May 6, 2013 and May 28, 2013, respectively, and which also claims priority to US provisional patent applications 61/736,527 and 61/748,427, both entitled SYSTEMS METHODS AND COMPOSITIONS FOR SEQUENCE MANIPULATION filed on Dec. 12, 2012 and Jan. 2, 2013, respectively, and which also claims priority to US provisional patent applications 61/791,409 and 61/835,931 filed on Mar. 15, 2013 and Jun. 17, 2013 respectively.
Claims
What is claimed is:
1. A method for selecting and producing an engineered CRISPR complex for targeting and/or cleavage of a candidate target nucleic acid sequence within a eukaryotic cell, comprising the steps of: (a) determining amount, location and nature of mismatch(es) of guide sequence of potential CRISPR complex(es) and the candidate target nucleic acid sequence, (b) determining contribution of each of the amount, location and nature of mismatch(es) to hybridization free energy of binding between the target nucleic acid sequence and the guide sequence of potential CRISPR complex(es) from a training data set, (c) based on the contribution analysis of step (b), predicting cleavage at the location(s) of the mismatch(es) of the target nucleic acid sequence by the potential CRISPR complex(es), (d) selecting the CRISPR complex from potential CRISPR complex(es) based on whether the prediction of step (c) indicates that it is more likely than not that cleavage will occur at location(s) of mismatch(es) by the CRISPR complex; (e) producing the selected CRISPR complex or nucleic acid molecule(s) encoding the selected CRISPR complex for targeting and/or cleavage of the candidate target nucleic acid sequence within the eukaryotic cell; and (f) delivering the selected CRISPR complex or nucleic acid molecule(s) encoding the selected CRISPR complex into the eukaryotic cell, wherein the selected CRISPR complex targets and/or cleaves the candidate target nucleic acid sequence within the eukaryotic cell.
2. The method of claim 1 wherein the candidate target sequence is a DNA sequence, and the mismatch(es) are of RNA of potential CRISPR complex(es) and the DNA.
3. The method of claim 1, wherein step (b) is performed by determining known local free energies, .DELTA.Gij(k), between every guide RNA sequence i and target DNA nucleic acid sequence j at position k, calculating values of the effective free-energy Z.sub.ij using the relationship p.sub.ij.varies.e.sup.-.beta.Zij,where p.sub.ij is measured cutting frequency by guide RNA sequence i on target DNA nucleic acid sequence j in the training set and .beta. is a positive constant of proportionality, determining the weights which are position-dependent weights .alpha..sub.k by fitting the known value of .DELTA.Gij(k) and the calculated value of Z.sub.ij across each guide RNA/target DNA sequence pair in the training set in the sum across all N bases of the guide-sequence .times..alpha..times..DELTA..times..function. ##EQU00008## by writing the above equation in the matrix form {right arrow over (Z)}=G{right arrow over (.alpha.)} and wherein, step (c) is performed by estimating the effective free-energy Zest using the determined position dependent weights in the equation {right arrow over (Z.sub.est)}=G {right arrow over (.alpha.)} and determining estimated spacer-target cutting frequencies p.sub.est .alpha.e.sup.-.beta.Zest, to thereby predict cleavage.
4. The method of claim 2 wherein the distance, in bp, between the first and last base of the target sequence is 18.
5. The method of claim 1 wherein predicting cleavage comprises predicting whether cleavage is more likely than not to occur at location(s) of mismatch(es), and thereby predicting cleavage.
6. The method of claim 1, further comprising normalizing the calculated values of the effective free energy of hybridization Z for each guide RNA/target DNA sequence pair in the training set.
7. The method of claim 1, further comprising filtering out calculated value of the effective free energy of hybridization Z for each guide RNA/target DNA sequence pair in the training set which have a sequencing depth which is below a minimum sequencing depth.
8. The method of claim 1, wherein the method is implemented by a computer system comprising: a. a memory unit configured to receive and/or store sequence information of the candidate target nucleic acid sequence; and b. one or more processors alone or in combination programmed to perform steps (a) to (d).
9. The method of claim 1, wherein step (b) is performed by: defining a thermodynamic model having a set of weights linking effective free energy of hybridization Z to local free energies G; defining a training set of the guide sequence/target DNA sequence pairs; inputting known values of local free energies G for each guide sequence/target DNA sequence pair in the training set; calculating a value of effective free energy of hybridization Z for each guide sequence/target DNA sequence pair in the training set; determining the weights using a machine learning algorithm, and outputting the weights whereby the weights can be used to estimate the free energy of hybridization for any sequence.
10. The method of claim 1 wherein the guide sequence is comprised within a single guide RNA (sgRNA) or within a CRISPR-Cas system chimera RNA (chiRNA).
11. The method of claim 1, wherein the selected CRISPR complex generates a cleavage within the candidate target nucleic acid sequence within the eukaryotic cell.
Description
Reference is also made to international patent applications PCT/US2013/074611 filed Dec. 12, 2013 and published as WO 2014/093595; PCT/US2013/074743 filed Dec. 12, 2013 and published as WO 2014/093661; PCT/US2013/074790 filed Dec. 12, 2013 and published as WO 2014/093694; PCT/US2013/074825 filed Dec. 12, 2013 and published as WO 2014/093718; PCT/US2013/074667 filed Dec. 12, 2013 and published as WO 2014/093622; PCT/US2013/074691 filed Dec. 12, 2013 and published as WO 2014/093635; PCT/US2013/074736 filed Dec. 12, 2013 and published as WO 2014/093655; PCT/US2013/074819 filed Dec. 12, 2013 and published as WO 2014/093712; and PCT/US2013/074800 filed Dec. 12, 2013 and published as WO 2014/093701.
Reference is also made to US provisional patent applications 61/757,972, filed Jan. 29, 2013, 61/799,800 filed Mar. 15, 2013, 61/835,936, 61/835,973, 61/836,080, 61/836,101, 61/836,123, 61/836,127, and 61/847,537, each filed Jun. 17, 2013, 61/862,468 and 61/862,355, each filed Aug. 5, 2013; 61/871,301 filed Aug. 28, 2013; 61/969,777 filed Sep. 25, 2013; and 61/961,980 filed Oct. 28, 2013.
The foregoing applications, and all documents cited therein or during their prosecution ("appln cited documents") and all documents cited or referenced in the appln cited documents, and all documents cited or referenced herein ("herein cited documents"), and all documents cited or referenced in herein cited documents, together with any manufacturer's instructions, descriptions, product specifications, and product sheets for any products mentioned herein or in any document incorporated by reference herein, are hereby incorporated herein by reference, and may be employed in the practice of the invention. More specifically, all referenced documents are incorporated by reference to the same extent as if each individual document was specifically and individually indicated to be incorporated by reference.
FIELD OF THE INVENTION
The present invention generally relates to the engineering and optimization of systems, methods and compositions used for the control of gene expression involving sequence targeting, such as genome perturbation or gene-editing, that relate to Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and components thereof.
BACKGROUND OF THE INVENTION
The CRISPR/Cas or the CRISPR-Cas system (both terms are used interchangeably throughout this application) does not require the generation of customized proteins to target specific sequences but rather a single Cas enzyme can be programmed by a short RNA molecule to recognize a specific DNA target. Adding the CRISPR-Cas system to the repertoire of genome sequencing techniques and analysis methods may significantly simplify the methodology and accelerate the ability to catalog and map genetic factors associated with a diverse range of biological functions and diseases. To utilize the CRISPR-Cas system effectively for genome editing without deleterious effects, it is critical to understand methods, systems and apparatus for identifying target sequences for Cas enzymes or CRISPR-Cas systems for target sequences of interest and conveying the results, which are aspects of the claimed invention.
SUMMARY OF THE INVENTION
The CRISPR/Cas or the CRISPR-Cas system (both terms may be used interchangeably throughout this application) does not require the generation of customized proteins to target specific sequences but rather a single Cas enzyme can be programmed by a short RNA molecule to recognize a specific DNA target, in other words the Cas enzyme can be recruited to a specific DNA target using said short RNA molecule. Adding the CRISPR-Cas system to the repertoire of genome sequencing techniques and analysis methods may significantly simplify the methodology and accelerate the ability to catalog and map genetic factors associated with a diverse range of biological functions and diseases. To utilize the CRISPR-Cas system effectively for genome editing without deleterious effects, it is critical to understand aspects of engineering and optimization of these genome engineering tools, which are aspects of the claimed invention.
In some aspects the invention relates to a non-naturally occurring or engineered composition comprising a CRISPR/Cas system chimeric RNA (chiRNA) polynucleotide sequence, wherein the polynucleotide sequence comprises (a) a guide sequence capable of hybridizing to a target sequence in a eukaryotic cell, (b) a tracr mate sequence, and (c) a tracr sequence wherein (a), (b) and (c) are arranged in a 5' to 3' orientation, wherein when transcribed, the tracr mate sequence hybridizes to the tracr sequence and the guide sequence directs sequence-specific binding of a CRISPR complex to the target sequence, wherein the CRISPR complex comprises a CRISPR enzyme complexed with (1) the guide sequence that is hybridized to the target sequence, and (2) the tracr mate sequence that is hybridized to the tracr sequence,
or
an CRISPR enzyme system, wherein the system is encoded by a vector system comprising one or more vectors comprising I. a first regulatory element operably linked to a CRISPR/Cas system chimeric RNA (chiRNA) polynucleotide sequence, wherein the polynucleotide sequence comprises (a) one or more guide sequences capable of hybridizing to one or more target sequences in a eukaryotic cell, (b) a tracr mate sequence, and (c) one or more tracr sequences, and II. a second regulatory element operably linked to an enzyme-coding sequence encoding a CRISPR enzyme comprising at least one or more nuclear localization sequences, wherein (a), (b) and (c) are arranged in a 5' to 3'orientation, wherein components I and II are located on the same or different vectors of the system, wherein when transcribed, the tracr mate sequence hybridizes to the tracr sequence and the guide sequence directs sequence-specific binding of a CRISPR complex to the target sequence, wherein the CRISPR complex comprises the CRISPR enzyme complexed with (1) the guide sequence that is hybridized to the target sequence, and (2) the tracr mate sequence that is hybridized to the tracr sequence,
or
a multiplexed CRISPR enzyme system, wherein the system is encoded by a vector system comprising one or more vectors comprising I. a first regulatory element operably linked to (a) one or more guide sequences capable of hybridizing to a target sequence in a cell, and (b) at least one or more tracr mate sequences, II. a second regulatory element operably linked to an enzyme-coding sequence encoding a CRISPR enzyme, and III. a third regulatory element operably linked to a tracr sequence, wherein components I, II and III are located on the same or different vectors of the system, wherein when transcribed, the tracr mate sequence hybridizes to the tracr sequence and the guide sequence directs sequence-specific binding of a CRISPR complex to the target sequence, wherein the CRISPR complex comprises the CRISPR enzyme complexed with (1) the guide sequence that is hybridized to the target sequence, and (2) the tracr mate sequence that is hybridized to the tracr sequence, and wherein in the multiplexed system multiple guide sequences and a single tracr sequence is used.
Without wishing to be bound by theory, it is believed that the target sequence should be associated with a PAM (protospacer adjacent motif); that is, a short sequence recognized by the CRISPR complex. This PAM may be considered a CRISPR motif.
With regard to the CRISPR system or complex discussed herein, reference is made to FIG. 2. FIG. 2 shows an exemplary CRISPR system and a possible mechanism of action (A), an example adaptation for expression in eukaryotic cells, and results of tests assessing nuclear localization and CRISPR activity (B-F).
The invention provides a method of identifying one or more unique target sequences. The target sequences may be in a genome of an organism, such as a genome of a eukaryotic organism. Accordingly, through potential sequence-specific binding, the target sequence may be susceptible to being recognized by a CRISPR-Cas system. (Likewise, the invention thus comprehends identifying one or more CRISPR-Cas systems that identifies one or more unique target sequences.) The target sequence may include the CRISPR motif and the sequence upstream or before it. The method may comprise: locating a CRISPR motif, e.g., analyzing (for instance comparing) a sequence to ascertain whether a CRISPR motif. e.g., a PAM sequence, a short sequence recognized by the CRISPR complex, is present in the sequence; analyzing (for instance comparing) the sequence upstream of the CRISPR motif to determine if that upstream sequence occurs elsewhere in the genome; selecting the upstream sequence if it does not occur elsewhere in the genome, thereby identifying a unique target site. The sequence upstream of the CRISPR motif may be at least 10 bp or at least 11 bp or at least 12 bp or at least 13 bp or at least 14 bp or at least 15 bp or at least 16 bp or at least 17 bp or at least 18 bp or at least 19 bp or at least 20 bp in length, e.g., the sequence upstream of the CRISPR motif may be about 10 bp to about 20 bp, e.g., the sequence upstream is 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29 or 30 bp in length. The CRISPR motif may be recognized by a Cas enzyme such as a Cas9 enzyme, e.g., a SpCas9 enzyme. Further, the CRISPR motif may be a protospacer-adjacent motif (PAM) sequence, e.g., NGG or NAG. Accordingly, as CRISPR motifs or PAM sequences may be recognized by a Cas enzyme in vitro, ex vivo or in vivo, in the in silico analysis, there is an analysis, e.g., comparison, of the sequence in interest against CRISPR motifs or PAM sequences to identify regions of the sequence in interest which may be recognized by a Cas enzyme in vitro, ex vivo or in vivo. When that analysis identifies a CRISPR motif or PAM sequence, the next analysis e.g., comparison is of the sequences upstream from the CRISPR motif or PAM sequence, e.g., analysis of the sequence 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29 or 30 bp in length starting at the PAM or CRISPR motif and extending upstream therefrom. That analysis is to see if that upstream sequence is unique, i.e., if the upstream sequence does not appear to otherwise occur in a genome, it may be a unique target site. The selection for unique sites is the same as the filtering step: in both cases, you filter away all target sequences with associated CRISPR motif that occur more than once in the target genome.
Eukaryotic organisms of interest may include but are not limited to Homo sapiens (human), Mus musculus (mouse), Rattus norvegicus (rat), Danio rerio (zebrafish), Drosophila melanogaster (fruit fly), Caenorhabditis elegans (roundworm), Sus scrofa (pig) and Bos taurus (cow). The eukaryotic organism can be selected from the group consisting of Homo sapiens (human), Mus musculus (mouse), Rattus norvegicus (rat), Danio rerio (zebrafish), Drosophila melanogaster (fruit fly), Caenorhabditis elegans (roundworm), Sus scrofa (pig) and Bos taurus (cow). The invention also comprehends computer-readable medium comprising codes that, upon execution by one or more processors, implements a herein method of identifying one or more unique target sequences.
The invention further comprehends a computer system for identifying one or more unique target sequences, e.g., in a genome, such as a genome of a eukaryotic organism, the system comprising: a. a memory unit configured to receive and/or store sequence information of the genome; and b. one or more processors alone or in combination programmed to perform a herein method of identifying one or more unique target sequences (e.g., locate a CRISPR motif, analyze a sequence upstream of the CRISPR motif to determine if the sequence occurs elsewhere in the genome, select the sequence if it does not occur elsewhere in the genome), to thereby identifying a unique target site and display and/or transmit the one or more unique target sequences. The candidate target sequence may be a DNA sequence. Mismatch(es) can be of RNA of the CRISPR complex and the DNA. In aspects of the invention, susceptibility of a target sequence being recognized by a CRISPR-Cas system indicates that there may be stable binding between the one or more base pairs of the target sequence and guide sequence of the CRISPR-Cas system to allow for specific recognition of the target sequence by the guide sequence.
The CRISPR/Cas or the CRISPR-Cas system utilizes a single Cas enzyme that can be programmed by a short RNA molecule to recognize a specific DNA target, in other words the Cas enzyme can be recruited to a specific DNA target using said short RNA molecule. In certain aspects, e.g., when not mutated or modified or when in a native state, the Cas or CRISPR enzyme in CRISPR/Cas or the CRISPR-Cas system, effects a cutting at a particular position; a specific DNA target. Accordingly, data can be generated--a data training set--relative to cutting by a CRISPR-Cas system at a particular position in a nucleotide, e.g., DNA, sequence at a particular position for a particular Cas or CRISPR enzyme. Similarly, data can be generated--a data training set--relative to cutting by a CRISPR-Cas system at a particular position in a nucleotide, e.g., DNA, sequence of a particular mismatch of typical nucleic acid hybridization (e.g., rather than G-C at particular position, G-T or G-U or G-A or G-G) for the particular Cas. In generating such data sets, there is the concept of average cutting frequency. The frequency by which an enzyme will cut a nucleic acid molecule, e.g., DNA, is mainly a function of the length of the sequence it is sensitive to. For instance, if an enzyme has a recognition sequence of 4 base-pairs, out of sheer probability, with 4 positions, and each position having potentially 4 different values, there are 4.sup.4 or 256 different possibilities for any given 4-base long strand. Therefore, theoretically (assuming completely random DNA), this enzyme will cut 1 in 256 4-base-pair long sites. For an enzyme that recognizes a sequence of 6 base-pairs, the calculation is 4.sup.6 or 4096 possible combinations with this length, and so such an enzyme will cut 1 in 4096 6-base-pair long sites. Of course, such calculations take into consideration only that each position has potentially 4 different values, and completely random DNA. However, DNA is not completely random; for example, the G-C content of organisms varies. Accordingly, the data training set(s) in the invention come from observing cutting by a CRISPR-Cas system at a particular position in a nucleotide, e.g., DNA, sequence at a particular position for a particular Cas or CRISPR enzyme and observing cutting by a CRISPR-Cas system at a particular position in a nucleotide, e.g., DNA, sequence of a particular mismatch of typical nucleic acid hybridization for the particular Cas, in a statistically significant number of experiments as to the particular position, the CRISPR-Cas system and the particular Cas, and averaging the results observed or obtained therefrom. The average cutting frequency may be defined as the mean of the cleavage efficiencies for all guide RNA:target DNA mismatches at a particular location.
The invention further provides a method of identifying one or more unique target sequences, e.g., in a genome, such as a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system (and likewise, the invention also further provides a method of identifying a CRISPR-Cas system susceptible to recognizing one or more unique target sequences), wherein the method comprises: a) determining average cutting frequency at a particular position for a particular Cas from a data training set as to that Cas, b) determining average cutting frequency of a particular mismatch (e.g., guide-RNA/target mismatch) for the particular Cas from the data training set, c) multiplying the average cutting frequency at a particular position by the average cutting frequency of a particular mismatch to obtain a first product, d) repeating steps a) to c) to obtain second and further products for any further particular position (s) of mismatches and particular mismatches and multiplying those second and further products by the first product, for an ultimate product, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position (or optionally d) repeating steps a) to c) to obtain second and further products for any further particular position (s) of mismatches and particular mismatches and multiplying those second and further products by the first product, for an ultimate product, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position), and e) multiplying the ultimate product by the result of dividing the minimum distance between consecutive mismatches by the distance, in bp, between the first and last base of the target sequence, e.g., 15-20, such as 18, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position (or optionally e) multiplying the ultimate product by the result of dividing the minimum distance between consecutive mismatches by the distance, in bp, between the first and last base of the target sequence, e.g., 15-20, such as 18 and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position), to thereby obtain a ranking, which allows for the identification of one or more unique target sequences, to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. Steps (a) and (b) can be performed in either order. If there are no other products than the first product, that first product (of step (c) from multiplying (a) times (b)) is what is used to determine or obtain the ranking.
The invention also comprehends method of identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises: a) creating a data training set as to a particular Cas, b) determining average cutting frequency at a particular position for the particular Cas from the data training set, c) determining average cutting frequency of a particular mismatch for the particular Cas from the data training set, d) multiplying the average cutting frequency at a particular position by the average cutting frequency of a particular mismatch to obtain a first product, e) repeating steps b) to d) to obtain second and further products for any further particular position (s) of mismatches and particular mismatches and multiplying those second and further products by the first product, for an ultimate product, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position (or optionally e) repeating steps b) to d) to obtain second and further products for any further particular position (s) of mismatches and particular mismatches and multiplying those second and further products by the first product, for an ultimate product, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position), and f) multiplying the ultimate product by the result of dividing the minimum distance between consecutive mismatches by 18 and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position (or optionally f) multiplying the ultimate product by the result of dividing the minimum distance between consecutive mismatches by the distance, in bp, between the first and last base of the target sequence, e.g., 15-20, such as 18, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position), to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. Steps (a) and (b) can be performed in either order. Steps (a) and (b) can be performed in either order. If there are no other products than the first product, that first product (of step (c) from multiplying (a) times (b)) is what is used to determine or obtain the ranking.
The invention also comprehends a method of identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises: a) determining average cutting frequency of guide-RNA/target mismatches at a particular position for a particular Cas from a training data set as to that Cas, and/or b) determining average cutting frequency of a particular mismatch-type for the particular Cas from the training data set, to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. The method may comprise determining both the average cutting frequency of guide-RNA/target mismatches at a particular position for a particular Cas from a training data set as to that Cas, and the average cutting frequency of a particular mismatch-type for the particular Cas from the training data set. Where both are determined, the method may further comprise multiplying the average cutting frequency at a particular position by the average cutting frequency of a particular mismatch-type to obtain a first product, repeating the determining and multiplying steps to obtain second and further products for any further particular position(s) of mismatches and particular mismatches and multiplying those second and further products by the first product, for an ultimate product, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position, and multiplying the ultimate product by the result of dividing the minimum distance between consecutive mismatches by the distance, in bp, between the first and last base of the target sequence and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position, to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. The distance, in bp, between the first and last base of the target sequence may be 18. The method may comprise creating a training set as to a particular Cas. The method may comprise determining the average cutting frequency of guide-RNA/target mismatches at a particular position for a particular Cas from a training data set as to that Cas, if more than one mismatch, repeating the determining step so as to determine cutting frequency for each mismatch, and multiplying frequencies of mismatches to thereby obtain a ranking, which allows for the identification of one or more unique target sequences.
The invention further comprehends a method of identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises: a) determining average cutting frequency of guide-RNA/target mismatches at a particular position for a particular Cas from a training data set as to that Cas, and average cutting frequency of a particular mismatch-type for the particular Cas from the training data set, to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. The invention additionally comprehends a method of identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises: a) creating a training data set as to a particular Cas, b) determining average cutting frequency of guide-RNA/target mismatches at a particular position for the particular Cas from the training data set, and/or c) determining average cutting frequency of a particular mismatch-type for the particular Cas from the training data set, to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. The invention yet further comprehends a method of identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises: a) creating a training data set as to a particular Cas, b) determining average cutting frequency of guide-RNA/target mismatches at a particular position for the particular Cas from the training data set, and average cutting frequency of a particular mismatch-type for the particular Cas from the training data set, to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. Accordingly, in these embodiments, instead of multiplying cutting-frequency averages uniquely determined for a mismatch position and mismatch type separately, the invention uses averages that are uniquely determined, e.g., cutting-frequency averages for a particular mismatch type at a particular position (thereby without multiplying these, as part of preparation of training set). These methods can be performed iteratively akin to the steps in methods including multiplication, for determination of one or more unique target sequences.
The invention in certain aspects provides a method for selecting a CRISPR complex for targeting and/or cleavage of a candidate target nucleic acid sequence within a cell, comprising the steps of: (a) determining amount, location and nature of mismatch(es) of guide sequence of potential CRISPR complex(es) and the candidate target nucleic acid sequence, (b) determining contribution of each of the amount, location and nature of mismatch(es) to hybridization free energy of binding between the target nucleic acid sequence and the guide sequence of potential CRISPR complex(es) from a training data set, (c) based on the contribution analysis of step (b), predicting cleavage at the location(s) of the mismatch(es) of the target nucleic acid sequence by the potential CRISPR complex(es), and (d) selecting the CRISPR complex from potential CRISPR complex(es) based on whether the prediction of step (c) indicates that it is more likely than not that cleavage will occur at location(s) of mismatch(es) by the CRISPR complex Step (b) may be performed by: determining local thermodynamic contributions, .DELTA.G.sub.ij(k), between every guide sequence i and target nucleic acid sequence j at position k, wherein .DELTA.G.sub.ij(k) is estimated from a biochemical prediction algorithm and .alpha..sub.k is a position-dependent weight calculated from the training data set, estimating values of the effective free-energy Z.sub.ij using the relationship p.sub.ij.varies.e.sup.-.beta.Z.sup.ij, wherein p.sub.ij is measured cutting frequency by guide sequence i on target nucleic acid sequence j and .beta. is a positive constant of proportionality, determining position-dependent weights .alpha..sub.k by fitting across spacer/target-pairs with the sum across all N bases of the guide-sequence
.times..times..alpha..times..DELTA..times..times..function. ##EQU00001##
and wherein, step (c) is performed by determining the position-dependent weights from the effective free-energy {right arrow over (Z.sub.est)}=G{right arrow over (.alpha.)} between each spacer and every potential target in the genome, and determining estimated spacer-target cutting frequencies p.sub.est.varies.e.sup.-.beta.Z.sup.est to thereby predict cleavage. Beta is implicitly fit by fitting the values of alpha (that are completely free to be multiplied--in the process of fitting--by whichever constant is suitable for Z=sum(alpha*Delta G).
The invention also comprehends the creation of a training data set. A training data set is data of cutting frequency measurements, obtained to maximize coverage and redundancy for possible mismatch types and positions. There are advantageously two experimental paradigms for generating a training data set. In one aspect, generating a data set comprises assaying for Cas, e.g., Cas9, cleavage at a constant target and mutating guide sequences. In another aspect, generating a data set comprises assaying for Cas, e.g., Cas9, cleavage using a constant guide sequence and testing cleavage at multiple DNA targets. Further, the method can be performed in at least two ways: in vivo (in cells, tissue, or living animal) or in vitro (with a cell-free assay, using in vitro transcribed guide RNA and Cas, e.g., Cas9 protein delivered either by whole cell lysate or purified protein). Advantageously the method is performed by assaying for cleavage at a constant target with mismatched guide RNA in vivo in cell lines. Because the guide RNA may be generated in cells as a transcript from a RNA polymerase III promoter (e.g. U6) driving a DNA oligo, it may be expressed as a PCR cassette and transfect the guide RNA directly (FIG. 24c) along with CBh-driven Cas9 (PX165, FIG. 24c). By co-transfecting Cas9 and a guide RNA with one or several mismatches relative to the constant DNA target, one may assess cleavage at a constant endogenous locus by a nuclease assay such as SURVEYOR nuclease assay or next-generation deep sequencing. This data may be collected for at least one or multiple targets within a loci of interest, e.g., at least 1, at least 5, at least 10, at least 15 or at least 20 targets from the human EMX1 locus. In this manner, a data training set can be readily generated for any locus of interest. Accordingly, there are at least two ways for generating a data training set--in vivo (in cell lines or living animal) or in vitro (with a cell-free assay, using in vitro transcribed guide RNA and Cas, e.g., Cas9, protein delivered either by whole cell lysate or purified protein). Also, the experimental paradigm can differ--e.g. with mutated guide sequences or with a constant guide and an oligo library of many DNA targets. These targeting experiments can be done in vitro as well. The readout would simply be running a gel on the result of the in vitro cleavage assay--the results will be cleaved and uncleaved fractions. Alternatively or additionally, these fractions can be gel-isolated and sequencing adapters can be ligated prior to deep sequencing on these populations.
The invention comprehends computer-readable medium comprising codes that, upon execution by one or more processors, implements a herein method. The invention further comprehends a computer system for performing a herein method. The system can include I. a memory unit configured to receive and/or store sequence information of the genome; and II. one or more processors alone or in combination programmed to perform the herein method, whereby the identification of one or more unique target sequences is advantageously displayed or transmitted. The eukaryotic organism can be selected from the group consisting of Homo sapiens (human), Mus musculus (mouse), Rattus norvegicus (rat), Danio rerio (zebrafish), Drosophila melanogaster (fruit fly), Caenorhabditis elegans (roundworm), Sus scrofa (pig) and Bos taurus (cow). The target sequence can be a DNA sequence, and the mismatch(es) can be of RNA of the CRISPR complex and the DNA.
The invention also entails a method for selecting a CRISPR complex for targeting and/or cleavage of a candidate target nucleic acid sequence, e.g., within a cell, comprising the steps of: (a) determining amount, location and nature of mismatch(es) of potential CRISPR complex(es) and the candidate target nucleic acid sequence, (b) determining the contribution of the mismatch(es) based on the amount and location of the mismatch(es), (c) based on the contribution analysis of step (b), predicting cleavage at the location(s) of the mismatch(es), and (d) selecting the CRISPR complex from potential CRISPR complex(es) based on whether the prediction of step (c) indicates that it is more likely than not that cleavage will occur at location(s) of mismatch(es) by the CRISPR complex. The cell can be from a eukaryotic organism as herein discussed. The determining steps can be based on the results or data of the data training set(s) in the invention that come from observing cutting by a CRISPR-Cas system at a particular position in a nucleotide, e.g., DNA, sequence at a particular position for a particular Cas or CRISPR enzyme and observing cutting by a CRISPR-Cas system at a particular position in a nucleotide, e.g., DNA, sequence of a particular mismatch of typical nucleic acid hybridization for the particular Cas, in a statistically significant number of experiments as to the particular position, the CRISPR-Cas system and the particular Cas, and averaging the results observed or obtained therefrom. Accordingly, for example, if the data training set shows that at a particular position the CRISPR-Cas system including a particular Cas is rather promiscuous, i.e., there can be mismatches and cutting, the amount and location may be one position, and nature of the mismatch between the CRISPR complex and the candidate target nucleic acid sequence may be not serious such that the contribution of the mismatch to failure to cut/bind may be negligible and the prediction for cleavage may be more likely than not that cleavage will occur, despite the mismatch. Accordingly, it should be clear that the data training set(s) are not generated in silico but are generated in the laboratory, e.g., are from in vitro, ex vivo and/or in vivo studies. The results from the laboratory work, e.g., from in vitro, ex vivo and/or in vivo studies, are input into computer systems for performing herein methods.
In the herein methods the candidate target sequence can be a DNA sequence, and the mismatch(es) can be of RNA of potential CRISPR complex(es) and the DNA. In aspects of the invention mentioned herein, the amount of mismatches indicates the number of mismatches in DNA: RNA base pairing between the DNA of the target sequence and the RNA of the guide sequence. In aspects of the invention the location of mismatches indicates the specific location along the sequence occupied by the mismatch and if more than one mismatch is present if the mismatches are concatenated or occur consecutively or if they are separated by at least one of more residues. In aspects of the invention the nature of mismatches indicates the nucleotide type involved in the mismatched base pairing. Base pairs are matched according to G-C and A-U Watson-Crick base pairing.
The invention further involves a method for predicting the efficiency of cleavage at candidate target nucleic acid sequence, e.g., within a target in a cell, by a CRISPR complex comprising the steps of: (a) determining amount, location and nature of mismatch(es) of the CRISPR complex and the candidate target nucleic acid sequence, (b) determining the contribution of the mismatch(es) based on the amount and location of the mismatch(es), and (c) based on the contribution analysis of step (b), predicting whether cleavage is more likely than not to occur at location(s) of mismatch(es), and thereby predicting cleavage. As with other herein methods, the candidate target sequence can be a DNA sequence, and the mismatch(es) can be of RNA of the CRISPR complex and the DNA. The cell can be from a eukaryotic organism as herein discussed.
The invention even further provides a method for selecting a candidate target sequence, e.g., within a nucleic acid sequence, e.g., in a cell, for targeting by a CRISPR complex, comprising the steps of: determining the local thermodynamic contributions, .DELTA.G.sub.ij(k), between every spacer i and target j at position k, expressing an effective free-energy Z.sub.ij for each spacer/target-pair as the sum
.times..times..alpha..times..DELTA..times..times..function. ##EQU00002## wherein .DELTA.G.sub.ij(k) is local thermodynamic contributions, estimated from a biochemical prediction algorithm and .alpha..sub.k is position-dependent weights, and estimating the effective free-energy Z through the relationship p.sub.ij.varies.e.sup.-.beta.Z.sup.ij wherein p.sub.ij is the measured cutting frequency by spacer i on target j and .beta. is a positive constant fit across the entire data-set, and estimating the position-dependent weights .alpha..sub.k by fitting G{right arrow over (.alpha.)}={right arrow over (Z)} such that each spacer-target pair (i,j) corresponds to a row in the matrix G and each position k in the spacer-target pairing corresponds to a column in the same matrix, and estimating the effective free-energy {right arrow over (Z.sub.est)}=G{right arrow over (.alpha.)} between each spacer and every potential target in the genome by using the fitted values .alpha..sub.k, and selecting, based on calculated effective free-energy values, the candidate spacer/target pair ij according to their specificity and/or the efficiency, given the estimated spacer-target cutting frequencies p.sub.est.varies.e.sup.-.beta.Z.sup.est. The cell can be from a eukaryotic organism as herein discussed.
The invention includes a computer-readable medium comprising codes that, upon execution by one or more processors, implements a method for selecting a CRISPR complex for targeting and/or cleavage of a candidate target nucleic acid, e.g., sequence within a cell, comprising the steps of: (a) determining amount, location and nature of mismatch(es) of potential CRISPR complex(es) and the candidate target nucleic acid sequence, (b) determining the contribution of the mismatch(es) based on the amount and location of the mismatch(es), (c) based on the contribution analysis of step (b), predicting cleavage at the location(s) of the mismatch(es), and (d) selecting the CRISPR complex from potential CRISPR complex(es) based on whether the prediction of step (c) indicates that it is more likely than not that cleavage will occur at location(s) of mismatch(es) by the CRISPR complex. The cell can be from a eukaryotic organism as herein discussed.
Also, the invention involves computer systems for selecting a CRISPR complex for targeting and/or cleavage of a candidate target nucleic acid sequence, e.g., within a cell, the system comprising: a. a memory unit configured to receive and/or store sequence information of the candidate target nucleic acid sequence; and b. one or more processors alone or in combination programmed to (a) determine amount, location and nature of mismatch(es) of potential CRISPR complex(es) and the candidate target nucleic acid sequence, (b) determine the contribution of the mismatch(es) based on the amount and location of the mismatch(es), (c) based on the contribution analysis of step (b), predicting cleavage at the location(s) of the mismatch(es), and (d) select the CRISPR complex from potential CRISPR complex(es) based on whether the prediction of step (c) indicates that it is more likely than not that cleavage will occur at location(s) of mismatch(es) by the CRISPR complex. The cell can be from a eukaryotic organism as herein discussed. The system can display or transmit the selection.
In aspects of the invention mentioned herein, the amount of mismatches indicates the number of mismatches in DNA: RNA base pairing between the DNA of the target sequence and the RNA of the guide sequence. In aspects of the invention the location of mismatches indicates the specific location along the sequence occupied by the mismatch and if more than one mismatch is present if the mismatches are concatenated or occur consecutively or if they are separated by at least one of more residues. In aspects of the invention the nature of mismatches indicates the nucleotide type involved in the mismatched base pairing. Base pairs are matched according to G-C and A-U Watson-Crick base pairing.
Accordingly, aspects of the invention relate to methods and compositions used to determine the specificity of Cas9. In one aspect the position and number of mismatches in the guide RNA is tested against cleavage efficiency. This information enables the design of target sequences that have minimal off-target effects.
The invention also comprehends a method of identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises a) determining average cutting frequency of guide-RNA/target mismatches at a particular position for a particular Cas from a training data set as to that Cas, and if more than one mismatch is present then step a) is repeated so as to determine cutting frequency for each mismatch after which frequencies of mismatches are multiplied to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. The invention further comprehends a method of identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises a) creating a training data set as to a particular Cas, b) determining average cutting frequency of guide-RNA/target mismatches at a particular position for a particular Cas from the training data set, if more than one mismatch exists, repeat step b) so as to determine cutting frequency for each mismatch, then multiply frequencies of mismatches to thereby obtain a ranking, which allows for the identification of one or more unique target sequences. The invention also relates to computer systems and computer readable media that executes these methods.
In various aspects, the invention involves a computer system for selecting a candidate target sequence within a nucleic acid sequence or for selecting a Cas for a candidate target sequence, e.g., selecting a target in a eukaryotic cell for targeting by a CRISPR complex.
The computer system may comprise: (a) a memory unit configured to receive and/or store said nucleic acid sequence; and (b) one or more processors alone or in combination programmed to perform as herein discussed. For example, programmed to: (i) locate a CRISPR motif sequence (e.g., PAM) within said nucleic acid sequence, and (ii) select a sequence adjacent to said located CRISPR motif sequence (e.g. PAM) as the candidate target sequence to which the CRISPR complex binds. In some embodiments, said locating step may comprise identifying a CRISPR motif sequence (e.g. PAM) located less than about 10000 nucleotides away from said target sequence, such as less than about 5000, 2500, 1000, 500, 250, 100, 50, 25, or fewer nucleotides away from the target sequence. In some embodiments, the candidate target sequence is at least 10, 15, 20, 25, 30, or more nucleotides in length. In some embodiments the candidate target sequence is 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39 or 40 nucleotides in length. In some embodiments, the nucleotide at the 3' end of the candidate target sequence is located no more than about 10 nucleotides upstream of the CRISPR motif sequence (e.g. PAM), such as no more than 5, 4, 3, 2, or 1 nucleotides. In some embodiments, the nucleic acid sequence in the eukaryotic cell is endogenous to the cell or organism, e.g., eukaryotic genome. In some embodiments, the nucleic acid sequence in the eukaryotic cell is exogenous to the cell or organism, e.g., eukaryotic genome.
In various aspects, the invention provides a computer-readable medium comprising codes that, upon execution by one or more processors, implements a method described herein, e.g., of selecting a candidate target sequence within a nucleic acid sequence or selecting a CRISPR candidate for a target sequence; for instance, a target sequence in a cell such as in a eukaryotic cell for targeting by a CRISPR complex. The method can comprise: (i) locate a CRISPR motif sequence (e.g., PAM) within said nucleic acid sequence, and (ii) select a sequence adjacent to said located CRISPR motif sequence (e.g. PAM) as the candidate target sequence to which the CRISPR complex binds. In some embodiments, said locating step may comprise identifying a CRISPR motif sequence (e.g. PAM) located less than about 10000 nucleotides away from said target sequence, such as less than about 5000, 2500, 1000, 500, 250, 100, 50, 25, or fewer nucleotides away from the target sequence. In some embodiments, the candidate target sequence is at least 10, 15, 20, 25, 30, or more nucleotides in length. In some embodiments the candidate target sequence is 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39 or 40 nucleotides in length. In some embodiments, the nucleotide at the 3' end of the candidate target sequence is located no more than about 10 nucleotides upstream of the CRISPR motif sequence (e.g. PAM), such as no more than 5, 4, 3, 2, or 1 nucleotides. In some embodiments, the nucleic acid sequence in the eukaryotic cell is endogenous to the cell or organism, e.g., eukaryotic genome. In some embodiments, the nucleic acid sequence in the eukaryotic cell is exogenous to the cell or organism, e.g., eukaryotic genome.
A computer system (or digital device) may be used to receive, transmit, display and/or store results, analyze the results, and/or produce a report of the results and analysis. A computer system may be understood as a logical apparatus that can read instructions from media (e.g. software) and/or network port (e.g. from the internet), which can optionally be connected to a server having fixed media. A computer system may comprise one or more of a CPU, disk drives, input devices such as keyboard and/or mouse, and a display (e.g. a monitor). Data communication, such as transmission of instructions or reports, can be achieved through a communication medium to a server at a local or a remote location. The communication medium can include any means of transmitting and/or receiving data. For example, the communication medium can be a network connection, a wireless connection, or an internet connection. Such a connection can provide for communication over the World Wide Web. It is envisioned that data relating to the present invention can be transmitted over such networks or connections (or any other suitable means for transmitting information, including but not limited to mailing a physical report, such as a print-out) for reception and/or for review by a receiver. The receiver can be but is not limited to an individual, or electronic system (e.g. one or more computers, and/or one or more servers).
In some embodiments, the computer system comprises one or more processors. Processors may be associated with one or more controllers, calculation units, and/or other units of a computer system, or implanted in firmware as desired. If implemented in software, the routines may be stored in any computer readable memory such as in RAM, ROM, flash memory, a magnetic disk, a laser disk, or other suitable storage medium. Likewise, this software may be delivered to a computing device via any known delivery method including, for example, over a communication channel such as a telephone line, the internet, a wireless connection, etc., or via a transportable medium, such as a computer readable disk, flash drive, etc. The various steps may be implemented as various blocks, operations, tools, modules and techniques which, in turn, may be implemented in hardware, firmware, software, or any combination of hardware, firmware, and/or software. When implemented in hardware, some or all of the blocks, operations, techniques, etc. may be implemented in, for example, a custom integrated circuit (IC), an application specific integrated circuit (ASIC), a field programmable logic array (FPGA), a programmable logic array (PLA), etc.
A client-server, relational database architecture can be used in embodiments of the invention. A client-server architecture is a network architecture in which each computer or process on the network is either a client or a server. Server computers are typically powerful computers dedicated to managing disk drives (file servers), printers (print servers), or network traffic (network servers). Client computers include PCs (personal computers) or workstations on which users run applications, as well as example output devices as disclosed herein. Client computers rely on server computers for resources, such as files, devices, and even processing power. In some embodiments of the invention, the server computer handles all of the database functionality. The client computer can have software that handles all the front-end data management and can also receive data input from users.
A machine readable medium comprising computer-executable code may take many forms, including but not limited to, a tangible storage medium, a carrier wave medium or physical transmission medium. Non-volatile storage media include, for example, optical or magnetic disks, such as any of the storage devices in any computer(s) or the like, such as may be used to implement the databases, etc. shown in the drawings. Volatile storage media include dynamic memory, such as main memory of such a computer platform. Tangible transmission media include coaxial cables; copper wire and fiber optics, including the wires that comprise a bus within a computer system. Carrier-wave transmission media may take the form of electric or electromagnetic signals, or acoustic or light waves such as those generated during radio frequency (RF) and infrared (IR) data communications. Common forms of computer-readable media therefore include for example: a floppy disk, a flexible disk, hard disk, magnetic tape, any other magnetic medium, a CD-ROM, DVD or DVD-ROM, any other optical medium, punch cards paper tape, any other physical storage medium with patterns of holes, a RAM, a ROM, a PROM and EPROM, a FLASH-EPROM, any other memory chip or cartridge, a carrier wave transporting data or instructions, cables or links transporting such a carrier wave, or any other medium from which a computer may read programming code and/or data. Many of these forms of computer readable media may be involved in carrying one or more sequences of one or more instructions to a processor for execution.
The subject computer-executable code can be executed on any suitable device comprising a processor, including a server, a PC, or a mobile device such as a smartphone or tablet. Any controller or computer optionally includes a monitor, which can be a cathode ray tube ("CRT") display, a flat panel display (e.g., active matrix liquid crystal display, liquid crystal display, etc.), or others. Computer circuitry is often placed in a box, which includes numerous integrated circuit chips, such as a microprocessor, memory, interface circuits, and others. The box also optionally includes a hard disk drive, a floppy disk drive, a high capacity removable drive such as a writeable CD-ROM, and other common peripheral elements. Inputting devices such as a keyboard, mouse, or touch-sensitive screen, optionally provide for input from a user. The computer can include appropriate software for receiving user instructions, either in the form of user input into a set of parameter fields, e.g., in a GUI, or in the form of preprogrammed instructions, e.g., preprogrammed for a variety of different specific operations.
Accordingly, it is an object of the invention to not encompass within the invention any previously known product, process of making the product, or method of using the product such that Applicants reserve the right and hereby disclose a disclaimer of any previously known product, process, or method. It is further noted that the invention does not intend to encompass within the scope of the invention any product, process, or making of the product or method of using the product, which does not meet the written description and enablement requirements of the USPTO (35 U.S.C. .sctn. 112, first paragraph) or the EPO (Article 83 of the EPC), such that Applicants reserve the right and hereby disclose a disclaimer of any previously described product, process of making the product, or method of using the product.
It is noted that in this disclosure and particularly in the claims and/or paragraphs, terms such as "comprises", "comprised", "comprising" and the like can have the meaning attributed to it in U.S. Patent law; e.g., they can mean "includes", "included", "including", and the like; and that terms such as "consisting essentially of" and "consists essentially of" have the meaning ascribed to them in U.S. Patent law, e.g., they allow for elements not explicitly recited, but exclude elements that are found in the prior art or that affect a basic or novel characteristic of the invention.
These and other embodiments are disclosed or are obvious from and encompassed by, the following Detailed Description.
BRIEF DESCRIPTION OF THE DRAWINGS
The novel features of the invention are set forth with particularity in the appended claims. A better understanding of the features and advantages of the present invention will be obtained by reference to the following detailed description that sets forth illustrative embodiments, in which the principles of the invention are utilized, and the accompanying drawings of which:
FIG. 1 shows a schematic of RNA-guided Cas9 nuclease. The Cas9 nuclease from Streptococcus pyogenes is targeted to genomic DNA by a synthetic guide RNA (sgRNA) consisting of a 20-nt guide sequence and a scaffold. The guide sequence base-pairs with the DNA target, directly upstream of a requisite 5'-NGG protospacer adjacent motif (PAM; magenta), and Cas9 mediates a double-stranded break (DSB) .about.3 bp upstream of the PAM (indicated by triangle).
FIG. 2A-F shows an exemplary CRISPR system and a possible mechanism of action (A), an example adaptation for expression in eukaryotic cells, and results of tests assessing nuclear localization and CRISPR activity (B-F).
FIG. 3 shows a schematic representation assay carried out to evaluate the cleavage specificity of Cas9 form Streptococcus pyogenes. Single base pair mismatches between the guide RNA sequence and the target DNA are mapped against cleavage efficiency in %.
FIG. 4 shows a mapping of mutations in the PAM sequence to cleavage efficiency in %.
FIG. 5A-C shows histograms of distances between adjacent S. pyogenes SF370 locus 1 PAM (NGG) (FIG. 5A) and S. thermophilus LMD9 locus 2 PAM (NNAGAAW) (FIG. 5B) in the human genome; and distances for each PAM by chromosome (Chr) (FIG. 5C).
FIG. 6A-C shows the graphing of distribution of distances between NGG and NRG motifs in the human genome in an "overlapping" fashion.
FIG. 7A-D shows a circular depiction of the phylogenetic analysis revealing five families of Cas9s, including three groups of large Cas9s (.about.1400 amino acids) and two of small Cas9s (.about.1100 amino acids).
FIG. 8A-F shows a linear depiction of the phylogenetic analysis revealing five families of Cas9s, including three groups of large Cas9s (.about.1400 amino acids) and two of small Cas9s (.about.1100 amino acids).
FIG. 9A-G shows the optimization of guide RNA architecture for SpCas9-mediated mammalian genome editing. (a) Schematic of bicistronic expression vector (PX330) for U6 promoter-driven single guide RNA (sgRNA) and CBh promoter-driven human codon-optimized Streptococcus pyogenes Cas9 (hSpCas9) used for all subsequent experiments. The sgRNA consists of a 20-nt guide sequence (blue) and scaffold (red), truncated at various positions as indicated. (b) SURVEYOR assay for SpCas9-mediated indels at the human EMX1 and PVALB loci. Arrows indicate the expected SURVEYOR fragments (n=3). (c) Northern blot analysis for the four sgRNA truncation architectures, with UI as loading control. (d) Both wildtype (wt) or nickase mutant (D10A) of SpCas9 promoted insertion of a HindIII site into the human EMX1 gene. Single stranded oligonucleotides (ssODNs), oriented in either the sense or antisense direction relative to genome sequence, were used as homologous recombination templates (FIG. 68). (e) Schematic of the human SERPINB5 locus. sgRNAs and PAMs are indicated by colored bars above sequence; methylcytosine (Me) are highlighted (pink) and numbered relative to the transcriptional start site (TSS, +1). (f) Methylation status of SERPINB5 assayed by bisulfite sequencing of 16 clones. Filled circles, methylated CpG; open circles, unmethylated CpG. (g) Modification efficiency by three sgRNAs targeting the methylated region of SERPINB5, assayed by deep sequencing (n=2). Error bars indicate Wilson intervals.
FIG. 10A-C shows position, distribution, number and mismatch-identity of some mismatch guide RNAs that can be used in generating the data training set (study on off target Cas9 activity).
FIG. 11A-B shows further positions, distributions, numbers and mismatch-identities of some mismatch guide RNAs that can be used in generating the data training set (study on off target Cas9 activity).
FIG. 12A-E shows guide RNA single mismatch cleavage efficiency. a, Multiple target sites were selected from the human EMX1 locus. Individual bases at positions 1-19 along the guide RNA sequence, which complementary to the target DNA sequence, were mutated to every ribonucleotide mismatch from the original guide RNA (blue `N`). b, On-target Cas9 cleavage activity for guide RNAs containing single base mutations (light blue: high cutting, dark blue: low cutting) relative to the on-target guide RNA (grey). c, Base transition heat map representing relative Cas9 cleavage activity for each possible RNA:DNA base pair. Rows were sorted based on cleavage activity in the PAM-proximal 10 bases of the guide RNA (high to low). Mean cleavage levels were calculated across base transitions in the PAM-proximal 10 bases (right bar) and across all transitions at each position (bottom bar). Heat map represents aggregate single-base mutation data from 15 EMX1 targets. d, Mean Cas9 locus modification efficiency at targets with all possible PAM sequences. e, Histogram of distances between 5'-NRG PAM occurrences within the human genome. Putative targets were identified using both the plus and minus strand of human chromosomal sequences.
FIG. 13A-C shows Cas9 on-target cleavage efficiency with multiple guide RNA mismatches and genome-wide specificity. a, Cas9 targeting efficiency with guide RNAs containing concatenated mismatches of 2 (top), 3 (middle), or 5 (bottom) consecutive bases for EMX1 targets 1 and 6. Rows represent different mutated guide RNAs and show the identity of each nucleotide mutation (white cells; grey cells denote unmutated bases). b, Cas9 was targeted with guide RNAs containing 3 (top, middle) or 4 (bottom) mismatches (white cells) separated by different numbers of unmutated bases (gray cells). c, Cleavage activity at targeted EMX1 target loci (top bar) as well as at candidate off-target genomic sites. Putative off-target loci contained 1-3 individual base differences (white cells) compared to the on-target loci.
FIG. 14A-B shows SpCas9 cleaves methylated targets in vitro. a, Plasmid targets containing CpG dinucleotides are either left unmethylated or methylated in vitro by M.SssI. Methyl-CpG in either the target sequence or PAM are indicated. b, Cleavage of either unmethylated or methylated targets 1 and 2 by SpCas9 cell lysate.
FIG. 15 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the human genome. A list of unique sites for the human, mouse, rat, zebrafish, fruit fly, and C. elegans genomes have been computationally identified and converted into tracks that can be visualized using the UCSC genome browser. Unique sites are defined as those sites with seed sequences (3'-most 12 nucleotides of the spacer sequence plus the NGG PAM sequence) that are unique in the entire genome.
FIG. 16 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the mouse genome.
FIG. 17 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the rat genome.
FIG. 18 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the zebra fish genome.
FIG. 19 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the D. melanogaster genome.
FIG. 20 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the C. elegans genome.
FIG. 21 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the pig genome.
FIG. 22 shows a UCSC Genome Browser track for identifying unique S. pyogenes Cas9 target sites in the cow genome.
FIG. 23 shows CRISPR Designer, a web app for the identification of Cas9 target sites. Most target regions (such as exons) contain multiple possible CRISPR sgRNA+PAM sequences. To minimize predicted off-targeted cleavage across the genome, a web-based computational pipeline ranks all possible sgRNA sites by their predicted genome-wide specificity and generates primers and oligos required for construction of each possible CRISPR as well as primers (via Primer3) for high-throughput assay of potential off-target cleavage in a next-generation sequencing experiment. Optimization of the choice of sgRNA within a user's target sequence: The goal is to minimize total off-target activity across the human genome. For each possible sgRNA choice, there is identification of off-target sequences (preceding either NAG or NGG PAMs) across the human genome that contain up to 5 mismatched base-pairs. The cleavage efficiency at each off-target sequence is predicted using an experimentally-derived weighting scheme. Each possible sgRNA is then ranked according to its total predicted off-target cleavage; the top-ranked sgRNAs represent those that are likely to have the greatest on-target and the least off-target cleavage. In addition, automated reagent design for CRISPR construction, primer design for the on-target SURVEYOR assay, and primer design for high-throughput detection and quantification of off-target cleavage via next-gen sequencing are advantageously facilitated.
FIG. 24A-C shows Target selection and reagent preparation. (a) For S. pyogenes Cas9, 20-bp targets (highlighted in blue) must be followed by 5'-NGG, which can occur in either strand on genomic DNA. (b) Schematic for co-transfection of Cas9 expression plasmid (PX165) and PCR-amplified U6-driven sgRNA expression cassette. Using a U6 promoter-containing PCR template and a fixed forward primer (U6 Fwd), sgRNA-encoding DNA can appended onto the U6 reverse primer (U6 Rev) and synthesized as an extended DNA oligo (Ultramer oligos from IDT). Note the guide sequence (blue N's) in U6 Rev is the reverse complement of the 5'-NGG flanking target sequence. (c) Schematic for scarless cloning of the guide sequence oligos into a plasmid containing Cas9 and sgRNA scaffold (PX330). The guide oligos (blue N's) contain overhangs for ligation into the pair of BbsI sites on PS330, with the top and bottom strand orientations matching those of the genomic target (i.e. top oligo is the 20-bp sequence preceding 5'-NGG in genomic DNA). Digestion of PX330 with BbsI allows the replacement of the Type IIs restriction sites (blue outline) with direct insertion of annealed oligos. It is worth noting that an extra G was placed before the first base of the guide sequence. Applicants have found that an extra G in front of the guide sequence does not adversely affect targeting efficiency. In cases when the 20-nt guide sequence of choice does not begin with guanine, the extra guanine will ensure the sgRNA is efficiently transcribed by the U6 promoter, which prefers a guanine in the first base of the transcript.
FIG. 25A-E shows the single nucleotide specificity of SpCas9. (a) Schematic of the experimental design. sgRNAs carrying all possible single base-pair mismatches (blue Ns) throughout the guide sequence were tested for each EMX1 target site (target site 1 shown as example). (b) Heatmap representation of relative SpCas9 cleavage efficiency by 57 single-mutated and 1 non-mutated sgRNA s each for four EMX1 target sites. For each EMX1 target, the identities of single base-pair substitutions are indicated on the left; original guide sequence is shown above and highlighted in the heatmap (grey squares). Modification efficiencies (increasing from white to dark blue) are normalized to the original guide sequence. (c) Heatmap for relative SpCas9 cleavage efficiency for each possible RNA:DNA base pair, compiled from aggregate data from single-mismatch guide RNAs for 15 EMX1 targets. Mean cleavage levels were calculated for the 10 PAM-proximal bases (right bar) and across all substitutions at each position (bottom bar); positions in grey were not covered by the 469 single-mutated and 15 non-mutated sgRNAs tested. (d) SpCas9-mediated indel frequencies at targets with all possible PAM sequences, determined using the SURVEYOR nuclease assay. Two target sites from the EMX1 locus were tested for each PAM (Table 4). (e) Histogram of distances between 5'-NRG PAM occurrences within the human genome. Putative targets were identified using both strands of human chromosomal sequences (GRCh37/hg19).
FIG. 26A-C shows the multiple mismatch specificity of SpCas9. (a) SpCas9 cleavage efficiency with guide RNAs containing a, consecutive mismatches of 2, 3, or 5 bases, or (b, c) multiple mismatches separated by different numbers of unmutated bases for EMX1 targets 1, 2, 3, and 6. Rows represent each mutated guide RNA; nucleotide substitutions are shown in white cells; grey cells denote unmutated bases. All indel frequencies are absolute and analyzed by deep sequencing from 2 biological replicas. Error bars indicate Wilson intervals (Example 7, Methods and Materials)
FIG. 27A-D shows SpCas9-mediated indel frequencies at predicted genomic off-target loci. (a and b) Cleavage levels at putative genomic off-target loci containing 2 or 3 individual mismatches (white cells) for EMX1 target 1 and target 3 are analyzed by deep sequencing. List of off-target sites are ordered by median position of mutations. Putative off-target sites with additional mutations did not exhibit detectable indels (Table 4). The Cas9 dosage was 3.times.10-10 nmol/cell, with equimolar sgRNA delivery. Error bars indicate Wilson intervals. (c and d) Indel frequencies for EMX1 targets 1 and 3 and selected off target loci (OT) as a function of SpCas9 and sgRNA dosage, normalized to on-target cleavage at highest transfection dosage (n=2). 400 ng to 10 ng of Cas9-sgRNA plasmid corresponds to 7.1.times.10-10 to 1.8.times.10-11 nmol/cell. Cleavage specificity is measured as a ratio of on- to off-target cleavage.
FIG. 28A-B shows the human EMX1 locus with target sites. Schematic of the human EMX1 locus showing the location of 15 target DNA sites, indicated by blue lines with corresponding PAM in magenta.
FIG. 29A-B shows additional genomic off-target site analysis. Cleavage levels at candidate genomic off-target loci (white cells) for a, EMX1 target 2 and b, EMX1 target 6 were analyzed by deep sequencing. All indel frequencies are absolute and analyzed by deep sequencing from 2 biological replicates. Error bars indicate Wilson confidence intervals
FIG. 30 shows predicted and observed cutting frequency-ranks among genome-wide targets.
FIG. 31 shows that the PAM for Staphylococcus aureus sp. Aureus Cas9 is NNGRR.
FIG. 32 shows a flow diagram as to locational methods of the invention.
FIG. 33A-B shows a flow diagram as to thermodynamic methods of the invention.
FIG. 34 shows a flow diagram as to multiplication methods of the invention.
FIG. 35 shows a schematic block diagram of a computer system which can be used to implement the methods described herein.
FIG. 36 shows a scatter plot of unfitted training data and testing data.
FIG. 37 shows a scatter plot of fitted training data and testing data.
The figures herein are for illustrative purposes only and are not necessarily drawn to scale.
DETAILED DESCRIPTION OF THE INVENTION
The invention relates to the engineering and optimization of systems, methods and compositions used for the control of gene expression involving sequence targeting, such as genome perturbation or gene-editing, that relate to the CRISPR/Cas system and components thereof (FIGS. 1 and 2). In advantageous embodiments, the Cas enzyme is Cas9.
The terms "polynucleotide", "nucleotide", "nucleotide sequence", "nucleic acid" and "oligonucleotide" are used interchangeably. They refer to a polymeric form of nucleotides of any length, either deoxyribonucleotides or ribonucleotides, or analogs thereof. Polynucleotides may have any three dimensional structure, and may perform any function, known or unknown. The following are non-limiting examples of polynucleotides: coding or non-coding regions of a gene or gene fragment, loci (locus) defined from linkage analysis, exons, introns, messenger RNA (mRNA), transfer RNA, ribosomal RNA, short interfering RNA (siRNA), short-hairpin RNA (shRNA), micro-RNA (miRNA), ribozymes, cDNA, recombinant polynucleotides, branched polynucleotides, plasmids, vectors, isolated DNA of any sequence, isolated RNA of any sequence, nucleic acid probes, and primers. The term also encompasses nucleic-acid-like structures with synthetic backbones, see, e.g., Eckstein, 1991; Baserga et al., 1992; Milligan, 1993; WO 97/03211; WO 96/39154; Mata, 1997; Strauss-Soukup, 1997; and Samstag, 1996. A polynucleotide may comprise one or more modified nucleotides, such as methylated nucleotides and nucleotide analogs. If present, modifications to the nucleotide structure may be imparted before or after assembly of the polymer. The sequence of nucleotides may be interrupted by non-nucleotide components. A polynucleotide may be further modified after polymerization, such as by conjugation with a labeling component.
As used herein the term "wild type" is a term of the art understood by skilled persons and means the typical form of an organism, strain, gene or characteristic as it occurs in nature as distinguished from mutant or variant forms.
As used herein the term "variant" should be taken to mean the exhibition of qualities that have a pattern that deviates from what occurs in nature.
The terms "non-naturally occurring" or "engineered" are used interchangeably and indicate the involvement of the hand of man. The terms, when referring to nucleic acid molecules or polypeptides mean that the nucleic acid molecule or the polypeptide is at least substantially free from at least one other component with which they are naturally associated in nature and as found in nature.
"Complementarity" refers to the ability of a nucleic acid to form hydrogen bond(s) with another nucleic acid sequence by either traditional Watson-Crick or other non-traditional types. A percent complementarity indicates the percentage of residues in a nucleic acid molecule which can form hydrogen bonds (e.g., Watson-Crick base pairing) with a second nucleic acid sequence (e.g., 5, 6, 7, 8, 9, 10 out of 10 being 50%, 60%, 70%, 80%, 90%, and 100% complementary). "Perfectly complementary" means that all the contiguous residues of a nucleic acid sequence will hydrogen bond with the same number of contiguous residues in a second nucleic acid sequence. "Substantially complementary" as used herein refers to a degree of complementarity that is at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, 98%, 99%, or 100% over a region of 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 30, 35, 40, 45, 50, or more nucleotides, or refers to two nucleic acids that hybridize under stringent conditions.
As used herein, "stringent conditions" for hybridization refer to conditions under which a nucleic acid having complementarity to a target sequence predominantly hybridizes with the target sequence, and substantially does not hybridize to non-target sequences. Stringent conditions are generally sequence-dependent, and vary depending on a number of factors. In general, the longer the sequence, the higher the temperature at which the sequence specifically hybridizes to its target sequence. Non-limiting examples of stringent conditions are described in detail in Tijssen (1993), Laboratory Techniques In Biochemistry And Molecular Biology-Hybridization With Nucleic Acid Probes Part I, Second Chapter "Overview of principles of hybridization and the strategy of nucleic acid probe assay", Elsevier, N.Y.
"Hybridization" refers to a reaction in which one or more polynucleotides react to form a complex that is stabilized via hydrogen bonding between the bases of the nucleotide residues. The hydrogen bonding may occur by Watson Crick base pairing, Hoogstein binding, or in any other sequence specific manner. The complex may comprise two strands forming a duplex structure, three or more strands forming a multi stranded complex, a single self-hybridizing strand, or any combination of these. A hybridization reaction may constitute a step in a more extensive process, such as the initiation of PCR, or the cleavage of a polynucleotide by an enzyme. A sequence capable of hybridizing with a given sequence is referred to as the "complement" of the given sequence.
As used herein, the term "genomic locus" or "locus" (plural loci) is the specific location of a gene or DNA sequence on a chromosome. A "gene" refers to stretches of DNA or RNA that encode a polypeptide or an RNA chain that has functional role to play in an organism and hence is the molecular unit of heredity in living organisms. For the purpose of this invention it may be considered that genes include regions which regulate the production of the gene product, whether or not such regulatory sequences are adjacent to coding and/or transcribed sequences. Accordingly, a gene includes, but is not necessarily limited to, promoter sequences, terminators, translational regulatory sequences such as ribosome binding sites and internal ribosome entry sites, enhancers, silencers, insulators, boundary elements, replication origins, matrix attachment sites and locus control regions.
As used herein, "expression of a genomic locus" or "gene expression" is the process by which information from a gene is used in the synthesis of a functional gene product. The products of gene expression are often proteins, but in non-protein coding genes such as rRNA genes or tRNA genes, the product is functional RNA. The process of gene expression is used by all known life--eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea) and viruses to generate functional products to survive. As used herein "expression" of a gene or nucleic acid encompasses not only cellular gene expression, but also the transcription and translation of nucleic acid(s) in cloning systems and in any other context. As used herein, "expression" also refers to the process by which a polynucleotide is transcribed from a DNA template (such as into and mRNA or other RNA transcript) and/or the process by which a transcribed mRNA is subsequently translated into peptides, polypeptides, or proteins. Transcripts and encoded polypeptides may be collectively referred to as "gene product." If the polynucleotide is derived from genomic DNA, expression may include splicing of the mRNA in a eukaryotic cell.
The terms "polypeptide", "peptide" and "protein" are used interchangeably herein to refer to polymers of amino acids of any length. The polymer may be linear or branched, it may comprise modified amino acids, and it may be interrupted by non amino acids. The terms also encompass an amino acid polymer that has been modified; for example, disulfide bond formation, glycosylation, lipidation, acetylation, phosphorylation, or any other manipulation, such as conjugation with a labeling component. As used herein the term "amino acid" includes natural and/or unnatural or synthetic amino acids, including glycine and both the D or L optical isomers, and amino acid analogs and peptidomimetics.
As used herein, the term "domain" or "protein domain" refers to a part of a protein sequence that may exist and function independently of the rest of the protein chain.
As described in aspects of the invention, sequence identity is related to sequence homology. Homology comparisons may be conducted by eye, or more usually, with the aid of readily available sequence comparison programs. These commercially available computer programs may calculate percent (%) homology between two or more sequences and may also calculate the sequence identity shared by two or more amino acid or nucleic acid sequences. In some preferred embodiments, the capping region of the dTALEs described herein have sequences that are at least 95% identical or share identity to the capping region amino acid sequences provided herein.
Sequence homologies may be generated by any of a number of computer programs known in the art, for example BLAST or FASTA, etc. A suitable computer program for carrying out such an alignment is the GCG Wisconsin Bestfit package (University of Wisconsin, U.S.A; Devereux et al., 1984, Nucleic Acids Research 12:387). Examples of other software than may perform sequence comparisons include, but are not limited to, the BLAST package (see Ausubel et al., 1999 ibid--Chapter 18), FASTA (Atschul et al., 1990, J. Mol. Biol., 403-410) and the GENEWORKS suite of comparison tools. Both BLAST and FASTA are available for offline and online searching (see Ausubel et al., 1999 ibid, pages 7-58 to 7-60). However it is preferred to use the GCG Bestfit program. % homology may be calculated over contiguous sequences, i.e., one sequence is aligned with the other sequence and each amino acid or nucleotide in one sequence is directly compared with the corresponding amino acid or nucleotide in the other sequence, one residue at a time. This is called an "ungapped" alignment. Typically, such ungapped alignments are performed only over a relatively short number of residues. Although this is a very simple and consistent method, it fails to take into consideration that, for example, in an otherwise identical pair of sequences, one insertion or deletion may cause the following amino acid residues to be put out of alignment, thus potentially resulting in a large reduction in % homology when a global alignment is performed. Consequently, most sequence comparison methods are designed to produce optimal alignments that take into consideration possible insertions and deletions without unduly penalizing the overall homology or identity score. This is achieved by inserting "gaps" in the sequence alignment to try to maximize local homology or identity. However, these more complex methods assign "gap penalties" to each gap that occurs in the alignment so that, for the same number of identical amino acids, a sequence alignment with as few gaps as possible--reflecting higher relatedness between the two compared sequences--may achieve a higher score than one with many gaps. "Affinity gap costs" are typically used that charge a relatively high cost for the existence of a gap and a smaller penalty for each subsequent residue in the gap. This is the most commonly used gap scoring system. High gap penalties may, of course, produce optimized alignments with fewer gaps. Most alignment programs allow the gap penalties to be modified. However, it is preferred to use the default values when using such software for sequence comparisons. For example, when using the GCG Wisconsin Bestfit package the default gap penalty for amino acid sequences is -12 for a gap and -4 for each extension. Calculation of maximum % homology therefore first requires the production of an optimal alignment, taking into consideration gap penalties. A suitable computer program for carrying out such an alignment is the GCG Wisconsin Bestfit package (Devereux et al., 1984 Nuc. Acids Research 12 p 387). Examples of other software than may perform sequence comparisons include, but are not limited to, the BLAST package (see Ausubel et al., 1999 Short Protocols in Molecular Biology, 4th Ed.--Chapter 18), FASTA (Altschul et al., 1990 J. Mol. Biol. 403-410) and the GENEWORKS suite of comparison tools. Both BLAST and FASTA are available for offline and online searching (see Ausubel et al., 1999, Short Protocols in Molecular Biology, pages 7-58 to 7-60). However, for some applications, it is preferred to use the GCG Bestfit program. A new tool, called BLAST 2 Sequences is also available for comparing protein and nucleotide sequences (see FEMS Microbiol Lett. 1999 174(2): 247-50; FEMS Microbiol Lett. 1999 177(1): 187-8 and the website of the National Center for Biotechnology information at the website of the National Institutes for Health). Although the final % homology may be measured in terms of identity, the alignment process itself is typically not based on an all-or-nothing pair comparison. Instead, a scaled similarity score matrix is generally used that assigns scores to each pair-wise comparison based on chemical similarity or evolutionary distance. An example of such a matrix commonly used is the BLOSUM62 matrix--the default matrix for the BLAST suite of programs. GCG Wisconsin programs generally use either the public default values or a custom symbol comparison table, if supplied (see user manual for further details). For some applications, it is preferred to use the public default values for the GCG package, or in the case of other software, the default matrix, such as BLOSUM62.
Alternatively, percentage homologies may be calculated using the multiple alignment feature in DNASIS.TM. (Hitachi Software), based on an algorithm, analogous to CLUSTAL (Higgins D G & Sharp P M (1988), Gene 73(1), 237-244). Once the software has produced an optimal alignment, it is possible to calculate % homology, preferably % sequence identity. The software typically does this as part of the sequence comparison and generates a numerical result.
The sequences may also have deletions, insertions or substitutions of amino acid residues which produce a silent change and result in a functionally equivalent substance. Deliberate amino acid substitutions may be made on the basis of similarity in amino acid properties (such as polarity, charge, solubility, hydrophobicity, hydrophilicity, and/or the amphipathic nature of the residues) and it is therefore useful to group amino acids together in functional groups. Amino acids may be grouped together based on the properties of their side chains alone. However, it is more useful to include mutation data as well. The sets of amino acids thus derived are likely to be conserved for structural reasons. These sets may be described in the form of a Venn diagram (Livingstone C. D. and Barton G. J. (1993) "Protein sequence alignments: a strategy for the hierarchical analysis of residue conservation" Comput. Appl. Biosci. 9: 745-756) (Taylor W. R. (1986) "The classification of amino acid conservation" J. Theor. Biol. 119; 205-218). Conservative substitutions may be made, for example according to the table below which describes a generally accepted Venn diagram grouping of amino acids.
TABLE-US-00001 Set Sub-set Hydrophobic F W Y H K M Aromatic F W Y H I L V A G C Aliphatic I L V Polar W Y H K R E Charged H K R E D D C S T N Q Positively H K R charged Negatively E D charged Small V C A G S P Tiny A G S T N D
Embodiments of the invention include sequences (both polynucleotide or polypeptide) which may comprise homologous substitution (substitution and replacement are both used herein to mean the interchange of an existing amino acid residue or nucleotide, with an alternative residue or nucleotide) that may occur i.e., like-for-like substitution in the case of amino acids such as basic for basic, acidic for acidic, polar for polar, etc. Non-homologous substitution may also occur i.e., from one class of residue to another or alternatively involving the inclusion of unnatural amino acids such as ornithine (hereinafter referred to as Z), diaminobutyric acid ornithine (hereinafter referred to as B), norleucine ornithine (hereinafter referred to as O), pyriylalanine, thienylalanine, naphthylalanine and phenylglycine.
Variant amino acid sequences may include suitable spacer groups that may be inserted between any two amino acid residues of the sequence including alkyl groups such as methyl, ethyl or propyl groups in addition to amino acid spacers such as glycine or .beta.-alanine residues. A further form of variation, which involves the presence of one or more amino acid residues in peptoid form, may be well understood by those skilled in the art. For the avoidance of doubt, "the peptoid form" is used to refer to variant amino acid residues wherein the .alpha.-carbon substituent group is on the residue's nitrogen atom rather than the .alpha.-carbon. Processes for preparing peptides in the peptoid form are known in the art, for example Simon R J et al., PNAS (1992) 89(20), 9367-9371 and Horwell D C, Trends Biotechnol. (1995) 13(4), 132-134.
The practice of the present invention employs, unless otherwise indicated, conventional techniques of immunology, biochemistry, chemistry, molecular biology, microbiology, cell biology, genomics and recombinant DNA, which are within the skill of the art. See Sambrook, Fritsch and Maniatis, MOLECULAR CLONING: A LABORATORY MANUAL, 2nd edition (1989); CURRENT PROTOCOLS IN MOLECULAR BIOLOGY (F. M. Ausubel, et al. eds., (1987)); the series METHODS IN ENZYMOLOGY (Academic Press, Inc.): PCR 2: A PRACTICAL APPROACH (M. J. MacPherson, B. D. Hames and G. R. Taylor eds. (1995)), Harlow and Lane, eds. (1988) ANTIBODIES, A LABORATORY MANUAL, and ANIMAL CELL CULTURE (R. I. Freshney, ed. (1987)).
In one aspect, the invention provides for vectors that are used in the engineering and optimization of CRISPR/Cas systems. A used herein, a "vector" is a tool that allows or facilitates the transfer of an entity from one environment to another. It is a replicon, such as a plasmid, phage, or cosmid, into which another DNA segment may be inserted so as to bring about the replication of the inserted segment. Generally, a vector is capable of replication when associated with the proper control elements. In general, the term "vector" refers to a nucleic acid molecule capable of transporting another nucleic acid to which it has been linked. Vectors include, but are not limited to, nucleic acid molecules that are single-stranded, double-stranded, or partially double-stranded; nucleic acid molecules that comprise one or more free ends, no free ends (e.g. circular); nucleic acid molecules that comprise DNA, RNA, or both; and other varieties of polynucleotides known in the art. One type of vector is a "plasmid," which refers to a circular double stranded DNA loop into which additional DNA segments can be inserted, such as by standard molecular cloning techniques. Another type of vector is a viral vector, wherein virally-derived DNA or RNA sequences are present in the vector for packaging into a virus (e.g. retroviruses, replication defective retroviruses, adenoviruses, replication defective adenoviruses, and adeno-associated viruses). Viral vectors also include polynucleotides carried by a virus for transfection into a host cell. Certain vectors are capable of autonomous replication in a host cell into which they are introduced (e.g. bacterial vectors having a bacterial origin of replication and episomal mammalian vectors). Other vectors (e.g., non-episomal mammalian vectors) are integrated into the genome of a host cell upon introduction into the host cell, and thereby are replicated along with the host genome. Moreover, certain vectors are capable of directing the expression of genes to which they are operatively-linked. Such vectors are referred to herein as "expression vectors." Common expression vectors of utility in recombinant DNA techniques are often in the form of plasmids. Recombinant expression vectors can comprise a nucleic acid of the invention in a form suitable for expression of the nucleic acid in a host cell, which means that the recombinant expression vectors include one or more regulatory elements, which may be selected on the basis of the host cells to be used for expression, that is operatively-linked to the nucleic acid sequence to be expressed. Within a recombinant expression vector, "operably linked" is intended to mean that the nucleotide sequence of interest is linked to the regulatory element(s) in a manner that allows for expression of the nucleotide sequence (e.g. in an in vitro transcription/translation system or in a host cell when the vector is introduced into the host cell). With regards to recombination and cloning methods, mention is made of U.S. patent application Ser. No. 10/815,730, the contents of which are herein incorporated by reference in their entirety.
Aspects of the invention can relate to bicistronic vectors for chimeric RNA and Cas9. Cas9 is driven by the CBh promoter and the chimeric RNA is driven by a U6 promoter. The chimeric guide RNA consists of a 20 bp guide sequence (Ns) joined to the tracr sequence (running from the first "U" of the lower strand to the end of the transcript), which is truncated at various positions as indicated. The guide and tracr sequences are separated by the tracr-mate sequence GUUUUAGAGCUA followed by the loop sequence GAAA. Results of SURVEYOR assays for Cas9-mediated indels at the human EMX1 and PVALB loci are illustrated in FIGS. 16b and 16c, respectively. Arrows indicate the expected SURVEYOR fragments. ChiRNAs are indicated by their "+n" designation, and crRNA refers to a hybrid RNA where guide and tracr sequences are expressed as separate transcripts. Throughout this application, chimeric RNA (chiRNA) may also be called single guide, or synthetic guide RNA (sgRNA).
The term "regulatory element" is intended to include promoters, enhancers, internal ribosomal entry sites (IRES), and other expression control elements (e.g. transcription termination signals, such as polyadenylation signals and poly-U sequences). Such regulatory elements are described, for example, in Goeddel, GENE EXPRESSION TECHNOLOGY: METHODS IN ENZYMOLOGY 185, Academic Press, San Diego, Calif. (1990). Regulatory elements include those that direct constitutive expression of a nucleotide sequence in many types of host cell and those that direct expression of the nucleotide sequence only in certain host cells (e.g., tissue-specific regulatory sequences). A tissue-specific promoter may direct expression primarily in a desired tissue of interest, such as muscle, neuron, bone, skin, blood, specific organs (e.g. liver, pancreas), or particular cell types (e.g. lymphocytes). Regulatory elements may also direct expression in a temporal-dependent manner, such as in a cell-cycle dependent or developmental stage-dependent manner, which may or may not also be tissue or cell-type specific. In some embodiments, a vector comprises one or more pol III promoter (e.g. 1, 2, 3, 4, 5, or more pol I promoters), one or more pol II promoters (e.g. 1, 2, 3, 4, 5, or more pol II promoters), one or more pol I promoters (e.g. 1, 2, 3, 4, 5, or more pol I promoters), or combinations thereof. Examples of pol III promoters include, but are not limited to, U6 and H1 promoters. Examples of pol II promoters include, but are not limited to, the retroviral Rous sarcoma virus (RSV) LTR promoter (optionally with the RSV enhancer), the cytomegalovirus (CMV) promoter (optionally with the CMV enhancer) [see, e.g., Boshart et al, Cell, 41:521-530 (1985)], the SV40 promoter, the dihydrofolate reductase promoter, the .beta.-actin promoter, the phosphoglycerol kinase (PGK) promoter, and the EF1.alpha. promoter. Also encompassed by the term "regulatory element" are enhancer elements, such as WPRE; CMV enhancers; the R-U5' segment in LTR of HTLV-I (Mol. Cell. Biol., Vol. 8(1), p. 466-472, 1988); SV40 enhancer; and the intron sequence between exons 2 and 3 of rabbit .beta.-globin (Proc. Natl. Acad. Sci. USA., Vol. 78(3), p. 1527-31, 1981). It will be appreciated by those skilled in the art that the design of the expression vector can depend on such factors as the choice of the host cell to be transformed, the level of expression desired, etc. A vector can be introduced into host cells to thereby produce transcripts, proteins, or peptides, including fusion proteins or peptides, encoded by nucleic acids as described herein (e.g., clustered regularly interspersed short palindromic repeats (CRISPR) transcripts, proteins, enzymes, mutant forms thereof, fusion proteins thereof, etc.). With regards to regulatory sequences, mention is made of U.S. patent application Ser. No. 10/491,026, the contents of which are incorporated by reference herein in their entirety. With regards to promoters, mention is made of PCT publication WO 2011/028929 and U.S. application Ser. No. 12/511,940, the contents of which are incorporated by reference herein in their entirety.
Vectors can be designed for expression of CRISPR transcripts (e.g. nucleic acid transcripts, proteins, or enzymes) in prokaryotic or eukaryotic cells. For example, CRISPR transcripts can be expressed in bacterial cells such as Escherichia coli, insect cells (using baculovirus expression vectors), yeast cells, or mammalian cells. Suitable host cells are discussed further in Goeddel, GENE EXPRESSION TECHNOLOGY: METHODS IN ENZYMOLOGY 185, Academic Press, San Diego, Calif. (1990). Alternatively, the recombinant expression vector can be transcribed and translated in vitro, for example using T7 promoter regulatory sequences and T7 polymerase. Vectors may be introduced and propagated in a prokaryote or prokaryotic cell. In some embodiments, a prokaryote is used to amplify copies of a vector to be introduced into a eukaryotic cell or as an intermediate vector in the production of a vector to be introduced into a eukaryotic cell (e.g. amplifying a plasmid as part of a viral vector packaging system). In some embodiments, a prokaryote is used to amplify copies of a vector and express one or more nucleic acids, such as to provide a source of one or more proteins for delivery to a host cell or host organism. Expression of proteins in prokaryotes is most often carried out in Escherichia coli with vectors containing constitutive or inducible promoters directing the expression of either fusion or non-fusion proteins. Fusion vectors add a number of amino acids to a protein encoded therein, such as to the amino terminus of the recombinant protein. Such fusion vectors may serve one or more purposes, such as: (i) to increase expression of recombinant protein; (ii) to increase the solubility of the recombinant protein; and (iii) to aid in the purification of the recombinant protein by acting as a ligand in affinity purification. Often, in fusion expression vectors, a proteolytic cleavage site is introduced at the junction of the fusion moiety and the recombinant protein to enable separation of the recombinant protein from the fusion moiety subsequent to purification of the fusion protein. Such enzymes, and their cognate recognition sequences, include Factor Xa, thrombin and enterokinase. Example fusion expression vectors include pGEX (Pharmacia Biotech Inc; Smith and Johnson, 1988. Gene 67: 31-40), pMAL (New England Biolabs, Beverly, Mass.) and pRIT5 (Pharmacia, Piscataway, N.J.) that fuse glutathione S-transferase (GST), maltose E binding protein, or protein A, respectively, to the target recombinant protein. Examples of suitable inducible non-fusion E. coli expression vectors include pTrc (Amrann et al., (1988) Gene 69:301-315) and pET 11d (Studier et al., GENE EXPRESSION TECHNOLOGY: METHODS IN ENZYMOLOGY 185, Academic Press, San Diego, Calif. (1990) 60-89). In some embodiments, a vector is a yeast expression vector. Examples of vectors for expression in yeast Saccharomyces cerivisae include pYepSec1 (Baldari, et al., 1987. EMBO J. 6: 229-234), pMFa (Kuijan and Herskowitz, 1982. Cell 30: 933-943), pJRY88 (Schultz et al., 1987. Gene 54: 113-123), pYES2 (Invitrogen Corporation, San Diego, Calif.), and picZ (InVitrogen Corp, San Diego, Calif.). In some embodiments, a vector drives protein expression in insect cells using baculovirus expression vectors. Baculovirus vectors available for expression of proteins in cultured insect cells (e.g., SF9 cells) include the pAc series (Smith, et al., 1983. Mol. Cell. Biol. 3: 2156-2165) and the pVL series (Lucklow and Summers, 1989. Virology 170: 31-39). In some embodiments, a vector is capable of driving expression of one or more sequences in mammalian cells using a mammalian expression vector. Examples of mammalian expression vectors include pCDM8 (Seed, 1987. Nature 329: 840) and pMT2PC (Kaufman, et al., 1987. EMBO J. 6: 187-195). When used in mammalian cells, the expression vector's control functions are typically provided by one or more regulatory elements. For example, commonly used promoters are derived from polyoma, adenovirus 2, cytomegalovirus, simian virus 40, and others disclosed herein and known in the art. For other suitable expression systems for both prokaryotic and eukaryotic cells see, e.g., Chapters 16 and 17 of Sambrook, et al., MOLECULAR CLONING: A LABORATORY MANUAL. 2nd ed., Cold Spring Harbor Laboratory, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 1989.
In some embodiments, the recombinant mammalian expression vector is capable of directing expression of the nucleic acid preferentially in a particular cell type (e.g., tissue-specific regulatory elements are used to express the nucleic acid). Tissue-specific regulatory elements are known in the art. Non-limiting examples of suitable tissue-specific promoters include the albumin promoter (liver-specific; Pinkert, et al., 1987. Genes Dev. 1: 268-277), lymphoid-specific promoters (Calame and Eaton, 1988. Adv. Immunol. 43: 235-275), in particular promoters of T cell receptors (Winoto and Baltimore, 1989. EMBO J. 8: 729-733) and immunoglobulins (Baneiji, et al., 1983. Cell 33: 729-740; Queen and Baltimore, 1983. Cell 33: 741-748), neuron-specific promoters (e.g., the neurofilament promoter; Byrne and Ruddle, 1989. Proc. Natl. Acad. Sci. USA 86: 5473-5477), pancreas-specific promoters (Edlund, et al., 1985. Science 230: 912-916), and mammary gland-specific promoters (e.g., milk whey promoter; U.S. Pat. No. 4,873,316 and European Application Publication No. 264,166). Developmentally-regulated promoters are also encompassed, e.g., the murine hox promoters (Kessel and Gruss, 1990. Science 249: 374-379) and the .alpha.-fetoprotein promoter (Campes and Tilghman, 1989. Genes Dev. 3: 537-546). With regard to these prokaryotic and eukaryotic vectors, mention is made of U.S. Pat. No. 6,750,059, the contents of which are incorporated by reference herein in their entirety. Other embodiments of the invention may relate to the use of viral vectors, with regards to which mention is made of U.S. patent application Ser. No. 13/092,085, the contents of which are incorporated by reference herein in their entirety. Tissue-specific regulatory elements are known in the art and in this regard, mention is made of U.S. Pat. No. 7,776,321, the contents of which are incorporated by reference herein in their entirety.
In some embodiments, a regulatory element is operably linked to one or more elements of a CRISPR system so as to drive expression of the one or more elements of the CRISPR system. In general, CRISPRs (Clustered Regularly Interspaced Short Palindromic Repeats), also known as SPIDRs (SPacer Interspersed Direct Repeats), constitute a family of DNA loci that are usually specific to a particular bacterial species. The CRISPR locus comprises a distinct class of interspersed short sequence repeats (SSRs) that were recognized in E. coli (Ishino et al., J. Bacteriol., 169:5429-5433 [1987]; and Nakata et al., J. Bacteriol., 171:3553-3556 [1989]), and associated genes. Similar interspersed SSRs have been identified in Haloferax mediterranei, Streptococcus pyogenes, Anabaena, and Mycobacterium tuberculosis (See, Groenen et al., Mol. Microbiol., 10:1057-1065 [1993]; Hoe et al., Emerg. Infect. Dis., 5:254-263 [1999]; Masepohl et al., Biochim. Biophys. Acta 1307:26-30 [1996]; and Mojica et al., Mol. Microbiol., 17:85-93 [1995]). The CRISPR loci typically differ from other SSRs by the structure of the repeats, which have been termed short regularly spaced repeats (SRSRs) (Janssen et al., OMICS J. Integ. Biol., 6:23-33 [2002]; and Mojica et al., Mol. Microbiol., 36:244-246 [2000]). In general, the repeats are short elements that occur in clusters that are regularly spaced by unique intervening sequences with a substantially constant length (Mojica et al., [2000], supra). Although the repeat sequences are highly conserved between strains, the number of interspersed repeats and the sequences of the spacer regions typically differ from strain to strain (van Embden et al., J. Bacteriol., 182:2393-2401 [2000]). CRISPR loci have been identified in more than 40 prokaryotes (See e.g., Jansen et al., Mol. Microbiol., 43:1565-1575 [2002]; and Mojica et al., [2005]) including, but not limited to Aeropyrum, Pyrobaculum, Sulfolobus, Archaeoglobus, Halocarcula, Methanobacterium, Methanococcus, Methanosarcina, Methanopyrus, Pyrococcus, Picrophilus, Thermoplasma, Corynebacterium, Mycobacterium, Streptomyces, Aquifex, Porphyromonas, Chlorobium, Thermus, Bacillus, Listeria, Staphylococcus, Clostridium, Thermoanaerobacter, Mycoplasma, Fusobacterium, Azarcus, Chromobacterium, Neisseria, Nitrosomonas, Desulfovibrio, Geobacter, Myxococcus, Campylobacter, Wolinella, Acinetobacter, Erwinia, Escherichia, Legionella, Methylococcus, Pasteurella, Photobacterium, Salmonella, Xanthomonas, Yersinia, Treponema, and Thermotoga.
In general, "CRISPR system" refers collectively to transcripts and other elements involved in the expression of or directing the activity of CRISPR-associated ("Cas") genes, including sequences encoding a Cas gene, a tracr (trans-activating CRISPR) sequence (e.g. tracrRNA or an active partial tracrRNA), a tracr-mate sequence (encompassing a "direct repeat" and a tracrRNA-processed partial direct repeat in the context of an endogenous CRISPR system), a guide sequence (also referred to as a "spacer" in the context of an endogenous CRISPR system), or other sequences and transcripts from a CRISPR locus. In embodiments of the invention the terms guide sequence and guide RNA are used interchangeably. In some embodiments, one or more elements of a CRISPR system is derived from a type I, type II, or type III CRISPR system. In some embodiments, one or more elements of a CRISPR system is derived from a particular organism comprising an endogenous CRISPR system, such as Streptococcus pyogenes. In general, a CRISPR system is characterized by elements that promote the formation of a CRISPR complex at the site of a target sequence (also referred to as a protospacer in the context of an endogenous CRISPR system). In the context of formation of a CRISPR complex, "target sequence" refers to a sequence to which a guide sequence is designed to have complementarity, where hybridization between a target sequence and a guide sequence promotes the formation of a CRISPR complex. A target sequence may comprise any polynucleotide, such as DNA or RNA polynucleotides. In some embodiments, a target sequence is located in the nucleus or cytoplasm of a cell.
In preferred embodiments of the invention, the CRISPR system is a type II CRISPR system and the Cas enzyme is Cas9, which catalyzes DNA cleavage. Enzymatic action by Cas9 derived from Streptococcus pyogenes or any closely related Cas9 generates double stranded breaks at target site sequences which hybridize to 20 nucleotides of the guide sequence and that have a protospacer-adjacent motif (PAM) sequence NGG following the 20 nucleotides of the target sequence. CRISPR activity through Cas9 for site-specific DNA recognition and cleavage is defined by the guide sequence, the tracr sequence that hybridizes in part to the guide sequence and the PAM sequence. More aspects of the CRISPR system are described in Karginov and Hannon, The CRISPR system: small RNA-guided defense in bacteria and archae, Mole Cell 2010, Jan. 15; 37(1): 7.
The type II CRISPR locus from Streptococcus pyogenes SF370, which contains a cluster of four genes Cas9, Cas1, Cas2, and Csn1, as well as two non-coding RNA elements, tracrRNA and a characteristic array of repetitive sequences (direct repeats) interspaced by short stretches of non-repetitive sequences (spacers, about 30 bp each). In this system, targeted DNA double-strand break (DSB) is generated in four sequential steps. First, two non-coding RNAs, the pre-crRNA array and tracrRNA, are transcribed from the CRISPR locus. Second, tracrRNA hybridizes to the direct repeats of pre-crRNA, which is then processed into mature crRNAs containing individual spacer sequences. Third, the mature crRNA:tracrRNA complex directs Cas9 to the DNA target consisting of the protospacer and the corresponding PAM via heteroduplex formation between the spacer region of the crRNA and the protospacer DNA. Finally, Cas9 mediates cleavage of target DNA upstream of PAM to create a DSB within the protospacer. Several aspects of the CRISPR system can be further improved to increase the efficiency and versatility of CRISPR targeting. Optimal Cas9 activity may depend on the availability of free Mg2+ at levels higher than that present in the mammalian nucleus (see e.g. Jinek et al., 2012, Science, 337:816), and the preference for an NGG motif immediately downstream of the protospacer restricts the ability to target on average every 12-bp in the human genome.
Typically, in the context of an endogenous CRISPR system, formation of a CRISPR complex (comprising a guide sequence hybridized to a target sequence and complexed with one or more Cas proteins) results in cleavage of one or both strands in or near (e.g. within 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 50, or more base pairs from) the target sequence. Without wishing to be bound by theory, the tracr sequence, which may comprise or consist of all or a portion of a wild-type tracr sequence (e.g. about or more than about 20, 26, 32, 45, 48, 54, 63, 67, 85, or more nucleotides of a wild-type tracr sequence), may also form part of a CRISPR complex, such as by hybridization along at least a portion of the tracr sequence to all or a portion of a tracr mate sequence that is operably linked to the guide sequence. In some embodiments, one or more vectors driving expression of one or more elements of a CRISPR system are introduced into a host cell such that expression of the elements of the CRISPR system direct formation of a CRISPR complex at one or more target sites. For example, a Cas enzyme, a guide sequence linked to a tracr-mate sequence, and a tracr sequence could each be operably linked to separate regulatory elements on separate vectors. Alternatively, two or more of the elements expressed from the same or different regulatory elements, may be combined in a single vector, with one or more additional vectors providing any components of the CRISPR system not included in the first vector. CRISPR system elements that are combined in a single vector may be arranged in any suitable orientation, such as one element located 5' with respect to ("upstream" of) or 3' with respect to ("downstream" of) a second element. The coding sequence of one element may be located on the same or opposite strand of the coding sequence of a second element, and oriented in the same or opposite direction. In some embodiments, a single promoter drives expression of a transcript encoding a CRISPR enzyme and one or more of the guide sequence, tracr mate sequence (optionally operably linked to the guide sequence), and a tracr sequence embedded within one or more intron sequences (e.g. each in a different intron, two or more in at least one intron, or all in a single intron). In some embodiments, the CRISPR enzyme, guide sequence, tracr mate sequence, and tracr sequence are operably linked to and expressed from the same promoter.
In some embodiments, a vector comprises one or more insertion sites, such as a restriction endonuclease recognition sequence (also referred to as a "cloning site"). In some embodiments, one or more insertion sites (e.g. about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more insertion sites) are located upstream and/or downstream of one or more sequence elements of one or more vectors. In some embodiments, a vector comprises an insertion site upstream of a tracr mate sequence, and optionally downstream of a regulatory element operably linked to the tracr mate sequence, such that following insertion of a guide sequence into the insertion site and upon expression the guide sequence directs sequence-specific binding of a CRISPR complex to a target sequence in a eukaryotic cell. In some embodiments, a vector comprises two or more insertion sites, each insertion site being located between two tracr mate sequences so as to allow insertion of a guide sequence at each site. In such an arrangement, the two or more guide sequences may comprise two or more copies of a single guide sequence, two or more different guide sequences, or combinations of these. When multiple different guide sequences are used, a single expression construct may be used to target CRISPR activity to multiple different, corresponding target sequences within a cell. For example, a single vector may comprise about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, or more guide sequences. In some embodiments, about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more such guide-sequence-containing vectors may be provided, and optionally delivered to a cell.
In some embodiments, a vector comprises a regulatory element operably linked to an enzyme-coding sequence encoding a CRISPR enzyme, such as a Cas protein. Non-limiting examples of Cas proteins include Cas1, Cas1B, Cas2, Cas3, Cas4, Cas5, Cas6, Cas7, Cas8, Cas9 (also known as Csn1 and Csx12), Cas10, Csy1, Csy2, Csy3, Cse1, Cse2, Csc1, Csc2, Csa5, Csn2, Csm2, Csm3, Csm4, Csm5, Csm6, Cmr1, Cmr3, Cmr4, Cmr5, Cmr6, Csb1, Csb2, Csb3, Csx17, Csx14, Csx10, Csx16, CsaX, Csx3, Csx1, Csx15, Csf1, Csf2, Csf3, Csf4, homologues thereof, or modified versions thereof. In some embodiments, the unmodified CRISPR enzyme has DNA cleavage activity, such as Cas9. In some embodiments, the CRISPR enzyme directs cleavage of one or both strands at the location of a target sequence, such as within the target sequence and/or within the complement of the target sequence. In some embodiments, the CRISPR enzyme directs cleavage of one or both strands within about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, 25, 50, 100, 200, 500, or more base pairs from the first or last nucleotide of a target sequence. In some embodiments, a vector encodes a CRISPR enzyme that is mutated to with respect to a corresponding wild-type enzyme such that the mutated CRISPR enzyme lacks the ability to cleave one or both strands of a target polynucleotide containing a target sequence. For example, an aspartate-to-alanine substitution (D10A) in the RuvC I catalytic domain of Cas9 from S. pyogenes converts Cas9 from a nuclease that cleaves both strands to a nickase (cleaves a single strand). Other examples of mutations that render Cas9 a nickase include, without limitation, H840A, N854A, and N863A. As a further example, two or more catalytic domains of Cas9 (RuvC I, RuvC II, and RuvC III or the HNH domain) may be mutated to produce a mutated Cas9 substantially lacking all DNA cleavage activity. In some embodiments, a D10A mutation is combined with one or more of H840A, N854A, or N863A mutations to produce a Cas9 enzyme substantially lacking all DNA cleavage activity. In some embodiments, a CRISPR enzyme is considered to substantially lack all DNA cleavage activity when the DNA cleavage activity of the mutated enzyme is less than about 25%, 10%, 5%, 1%, 0.1%, 0.01%, or lower with respect to its non-mutated form. An aspartate-to-alanine substitution (D10A) in the RuvC I catalytic domain of SpCas9 converts the nuclease into a nickase (see e.g. Sapranauskas et al., 2011, Nucleic Acis Research, 39: 9275; Gasiunas et al., 2012, Proc. Natl. Acad. Sci. USA, 109:E2579), such that nicked genomic DNA undergoes the high-fidelity homology-directed repair (HDR). In some embodiments, an enzyme coding sequence encoding a CRISPR enzyme is codon optimized for expression in particular cells, such as eukaryotic cells. The eukaryotic cells may be those of or derived from a particular organism, such as a mammal, including but not limited to human, mouse, rat, rabbit, dog, or non-human primate. In general, codon optimization refers to a process of modifying a nucleic acid sequence for enhanced expression in the host cells of interest by replacing at least one codon (e.g. about or more than about 1, 2, 3, 4, 5, 10, 15, 20, 25, 50, or more codons) of the native sequence with codons that are more frequently or most frequently used in the genes of that host cell while maintaining the native amino acid sequence. Various species exhibit particular bias for certain codons of a particular amino acid. Codon bias (differences in codon usage between organisms) often correlates with the efficiency of translation of messenger RNA (mRNA), which is in turn believed to be dependent on, among other things, the properties of the codons being translated and the availability of particular transfer RNA (tRNA) molecules. The predominance of selected tRNAs in a cell is generally a reflection of the codons used most frequently in peptide synthesis. Accordingly, genes can be tailored for optimal gene expression in a given organism based on codon optimization. Codon usage tables are readily available, See Nakamura. Y., et al. "Codon usage tabulated from the international DNA sequence databases: status for the year 2000" Nucl. Acids Res. 28:292 (2000). Computer algorithms for codon optimizing a particular sequence for expression in a particular host cell are also available, such as Gene Forge (Aptagen; Jacobus, Pa.), are also available. In some embodiments, one or more codons (e.g. 1, 2, 3, 4, 5, 10, 15, 20, 25, 50, or more, or all codons) in a sequence encoding a CRISPR enzyme correspond to the most frequently used codon for a particular amino acid.
In some embodiments, a vector encodes a CRISPR enzyme comprising one or more nuclear localization sequences (NLSs), such as about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more NLSs. In some embodiments, the CRISPR enzyme comprises about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more NLSs at or near the amino-terminus, about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more NLSs at or near the carboxy-terminus, or a combination of these (e.g. one or more NLS at the amino-terminus and one or more NLS at the carboxy terminus). When more than one NLS is present, each may be selected independently of the others, such that a single NLS may be present in more than one copy and/or in combination with one or more other NLSs present in one or more copies. In a preferred embodiment of the invention, the CRISPR enzyme comprises at most 6 NLSs. In some embodiments, an NLS is considered near the N- or C-terminus when the nearest amino acid of the NLS is within about 1, 2, 3, 4, 5, 10, 15, 20, 25, 30, 40, 50, or more amino acids along the polypeptide chain from the N- or C-terminus. Non-limiting examples of NLSs include an NLS sequence derived from: the NLS of the SV40 virus large T-antigen, having the amino acid sequence PKKKRKV; the NLS from nucleoplasmin (e.g. the nucleoplasmin bipartite NLS with the sequence KRPAATKKAGQAKKKK); the c-myc NLS having the amino acid sequence PAAKRVKLD or RQRRNELKRSP; the hRNPA1 M9 NLS having the sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY; the sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV of the IBB domain from importin-alpha; the sequences VSRKRPRP and PPKKARED of the myoma T protein; the sequence POPKKKPL of human p53; the sequence SALIKKKKKMAP of mouse c-abl IV; the sequences DRLRR and PKQKKRK of the influenza virus NS1; the sequence RKLKKKIKKL of the Hepatitis virus delta antigen; the sequence REKKKFLKRR of the mouse Mx1 protein; the sequence KRKGDEVDGVDEVAKKKSKK of the human poly(ADP-ribose) polymerase; and the sequence RKCLQAGMNLEARKTKK of the steroid hormone receptors (human) glucocorticoid.
In general, the one or more NLSs are of sufficient strength to drive accumulation of the CRISPR enzyme in a detectable amount in the nucleus of a eukaryotic cell. In general, strength of nuclear localization activity may derive from the number of nuclear localization sequence(s) (NLS(s)) in the CRISPR enzyme, the particular NLS(s) used, or a combination of these factors. Detection of accumulation in the nucleus may be performed by any suitable technique. For example, a detectable marker may be fused to the CRISPR enzyme, such that location within a cell may be visualized, such as in combination with a means for detecting the location of the nucleus (e.g. a stain specific for the nucleus such as DAPI). Cell nuclei may also be isolated from cells, the contents of which may then be analyzed by any suitable process for detecting protein, such as immunohistochemistry, Western blot, or enzyme activity assay. Accumulation in the nucleus may also be determined indirectly, such as by an assay for the effect of CRISPR complex formation (e.g. assay for DNA cleavage or mutation at the target sequence, or assay for altered gene expression activity affected by CRISPR complex formation and/or CRISPR enzyme activity), as compared to a control no exposed to the CRISPR enzyme or complex, or exposed to a CRISPR enzyme lacking the one or more NLSs.
In general, a guide sequence is any polynucleotide sequence having sufficient complementarity with a target polynucleotide sequence to hybridize with the target sequence and direct sequence-specific binding of a CRISPR complex to the target sequence. Throughout this application the guide sequence may be interchangeably referred to as a guide or a spacer. In some embodiments, the degree of complementarity between a guide sequence and its corresponding target sequence, when optimally aligned using a suitable alignment algorithm, is about or more than about 50%, 60%, 75%, 80%, 85%, 90%, 95%, 97.5%, 99%, or more. Optimal alignment may be determined with the use of any suitable algorithm for aligning sequences, non-limiting example of which include the Smith-Waterman algorithm, the Needleman-Wunsch algorithm, algorithms based on the Burrows-Wheeler Transform (e.g. the Burrows Wheeler Aligner), ClustalW, Clustal X, BLAT, Novoalign (Novocraft Technologies; available at www.novocraft.com), ELAND (Illumina, San Diego, Calif.), SOAP (available at soap.genomics.org.cn), and Maq (available at maq.sourceforge.net). In some embodiments, a guide sequence is about or more than about 5, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 35, 40, 45, 50, 75, or more nucleotides in length. In some embodiments, a guide sequence is less than about 75, 50, 45, 40, 35, 30, 25, 20, 15, 12, or fewer nucleotides in length. The ability of a guide sequence to direct sequence-specific binding of a CRISPR complex to a target sequence may be assessed by any suitable assay. For example, the components of a CRISPR system sufficient to form a CRISPR complex, including the guide sequence to be tested, may be provided to a host cell having the corresponding target sequence, such as by transfection with vectors encoding the components of the CRISPR sequence, followed by an assessment of preferential cleavage within the target sequence, such as by SURVEYOR assay as described herein. Similarly, cleavage of a target polynucleotide sequence may be evaluated in a test tube by providing the target sequence, components of a CRISPR complex, including the guide sequence to be tested and a control guide sequence different from the test guide sequence, and comparing binding or rate of cleavage at the target sequence between the test and control guide sequence reactions. Other assays are possible, and will occur to those skilled in the art.
A guide sequence may be selected to target any target sequence. In some embodiments, the target sequence is a sequence within a genome of a cell. Exemplary target sequences include those that are unique in the target genome. For example, for the S. pyogenes Cas9, a unique target sequence in a genome may include a Cas9 target site of the form MMMMMMMMNNNNNNNNNNNNXGG where NNNNNNNNNNNNXGG (N is A, G, T, or C; and X can be anything) has a single occurrence in the genome. A unique target sequence in a genome may include an S. pyogenes Cas9 target site of the form MMMMMMMMMNNNNNNNNNNNXGG where NNNNNNNNNNNXGG (N is A, G, T, or C; and X can be anything) has a single occurrence in the genome. For the S. thermophilus CRISPR1 Cas9, a unique target sequence in a genome may include a Cas9 target site of the form MMMMMMMMNNNNNNNNNNNNXXAGAAW where NNNNNNNNNNNNXXAGAAW (N is A, G, T, or C; X can be anything; and W is A or T) has a single occurrence in the genome. A unique target sequence in a genome may include an S. thermophilus CRISPR1 Cas9 target site of the form MMMMMMMMMNNNNNNNNNNNXXAGAAW where NNNNNNNNNNNXXAGAAW (N is A, G, T, or C; X can be anything; and W is A or T) has a single occurrence in the genome. For the S. pyogenes Cas9, a unique target sequence in a genome may include a Cas9 target site of the form MMMMMMMMNNNNNNNNNNNNXGGXG where NNNNNNNNNNNNXGGXG (N is A, G, T, or C; and X can be anything) has a single occurrence in the genome. A unique target sequence in a genome may include an S. pyogenes Cas9 target site of the form MMMMMMMMMNNNNNNNNNNNXGGXG where NNNNNNNNNNNXGGXG (N is A, G, T, or C; and X can be anything) has a single occurrence in the genome. In each of these sequences "M" may be A, G, T, or C, and need not be considered in identifying a sequence as unique.
In some embodiments, a guide sequence is selected to reduce the degree secondary structure within the guide sequence. In some embodiments, about or less than about 75%, 50%, 40%, 30%, 25%, 20%, 15%, 10%, 5%, 1%, or fewer of the nucleotides of the guide sequence participate in self-complementary base pairing when optimally folded. Optimal folding may be determined by any suitable polynucleotide folding algorithm. Some programs are based on calculating the minimal Gibbs free energy. An example of one such algorithm is mFold, as described by Zuker and Stiegler (Nucleic Acids Res. 9 (1981), 133-148). Another example folding algorithm is the online webserver RNAfold, developed at Institute for Theoretical Chemistry at the University of Vienna, using the centroid structure prediction algorithm (see e.g. A. R. Gruber et al., 2008, Cell 106(1): 23-24; and P A Carr and G M Church, 2009, Nature Biotechnology 27(12): 1151-62).
In general, a tracr mate sequence includes any sequence that has sufficient complementarity with a tracr sequence to promote one or more of: (1) excision of a guide sequence flanked by tracr mate sequences in a cell containing the corresponding tracr sequence; and (2) formation of a CRISPR complex at a target sequence, wherein the CRISPR complex comprises the tracr mate sequence hybridized to the tracr sequence. In general, degree of complementarity is with reference to the optimal alignment of the tracr mate sequence and tracr sequence, along the length of the shorter of the two sequences. Optimal alignment may be determined by any suitable alignment algorithm, and may further account for secondary structures, such as self-complementarity within either the tracr sequence or tracr mate sequence. In some embodiments, the degree of complementarity between the tracr sequence and tracr mate sequence along the length of the shorter of the two when optimally aligned is about or more than about 25%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, 97.5%, 99%, or higher. In some embodiments, the tracr sequence is about or more than about 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, 30, 40, 50, or more nucleotides in length. In some embodiments, the tracr sequence and tracr mate sequence are contained within a single transcript, such that hybridization between the two produces a transcript having a secondary structure, such as a hairpin. In an embodiment of the invention, the transcript or transcribed polynucleotide sequence has at least two or more hairpins. In preferred embodiments, the transcript has two, three, four or five hairpins. In a further embodiment of the invention, the transcript has at most five hairpins. In a hairpin structure the portion of the sequence 5' of the final "N" and upstream of the loop corresponds to the tracr mate sequence, and the portion of the sequence 3' of the loop corresponds to the tracr sequence An example illustration of such a hairpin structure is provided in the lower portion of FIG. 15B. Further non-limiting examples of single polynucleotides comprising a guide sequence, a tracr mate sequence, and a tracr sequence are as follows (listed 5' to 3'), where "N" represents a base of a guide sequence, the first block of lower case letters represent the tracr mate sequence, and the second block of lower case letters represent the tracr sequence, and the final poly-T sequence represents the transcription terminator: (1) NNNNNNNNNNNNNNNNNNNNgtttttgtactctcaagatttaGAAAtaaatcttgcagaagctacaaagataa- ggctt catgccgaaatcaacaccctgtcattttatggcagggtgttttcgttatttaaTTTTTT; (2) NNNNNNNNNNNNNNgtttttgtacttcaGAAAtgcagaagctacaaagataaggcttcatgccgaaatca acaccctgtcattttatggcagggtgttttcgttatttaaTTTTTT; (3) NNNNNNNNNNNNNNNNNNgtttttgtactctcaGAAAtgcagaagctacaaagataaggcttcatgccgaaat- ca acaccctgtcattttatggcagggtgtTTTTTT; (4) NNNNNNNNNNNNNNNNNNNNgttttagagctaGAAAtagcaagttaaaataaggctagtccgttatcaacttg- aaaa agtggcaccgagtcggtgcTTTTTT; (5) NNNNNNNNNNNNNNNNNNNNgttttagagctaGAAATAGcaagttaaaataaggctagtccgttatcaacttg- aa aaagtgTTTTTT; and (6) NNNNNNNNNNNNNNNNNNNNgttttagagctagAAATAGcaagttaaaataaggctagtccgttatcaTTTTT TTT. In some embodiments, sequences (1) to (3) are used in combination with Cas9 from S. thermophilus CRISPR1. In some embodiments, sequences (4) to (6) are used in combination with Cas9 from S. pyogenes. In some embodiments, the tracr sequence is a separate transcript from a transcript comprising the tracr mate sequence.
In some embodiments, a recombination template is also provided. A recombination template may be a component of another vector as described herein, contained in a separate vector, or provided as a separate polynucleotide. In some embodiments, a recombination template is designed to serve as a template in homologous recombination, such as within or near a target sequence nicked or cleaved by a CRISPR enzyme as a part of a CRISPR complex. A template polynucleotide may be of any suitable length, such as about or more than about 10, 15, 20, 25, 50, 75, 100, 150, 200, 500, 1000, or more nucleotides in length. In some embodiments, the template polynucleotide is complementary to a portion of a polynucleotide comprising the target sequence. When optimally aligned, a template polynucleotide might overlap with one or more nucleotides of a target sequences (e.g. about or more than about 1, 5, 10, 15, 20, or more nucleotides). In some embodiments, when a template sequence and a polynucleotide comprising a target sequence are optimally aligned, the nearest nucleotide of the template polynucleotide is within about 1, 5, 10, 15, 20, 25, 50, 75, 100, 200, 300, 400, 500, 1000, 5000, 10000, or more nucleotides from the target sequence.
In some embodiments, the CRISPR enzyme is part of a fusion protein comprising one or more heterologous protein domains (e.g. about or more than about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, or more domains in addition to the CRISPR enzyme). A CRISPR enzyme fusion protein may comprise any additional protein sequence, and optionally a linker sequence between any two domains. Examples of protein domains that may be fused to a CRISPR enzyme include, without limitation, epitope tags, reporter gene sequences, and protein domains having one or more of the following activities: methylase activity, demethylase activity, transcription activation activity, transcription repression activity, transcription release factor activity, histone modification activity, RNA cleavage activity and nucleic acid binding activity. Non-limiting examples of epitope tags include histidine (His) tags, V5 tags, FLAG tags, influenza hemagglutinin (HA) tags, Myc tags, VSV-G tags, and thioredoxin (Trx) tags. Examples of reporter genes include, but are not limited to, glutathione-S-transferase (GST), horseradish peroxidase (HRP), chloramphenicol acetyltransferase (CAT) beta-galactosidase, beta-glucuronidase, luciferase, green fluorescent protein (GFP), HcRed, DsRed, cyan fluorescent protein (CFP), yellow fluorescent protein (YFP), and autofluorescent proteins including blue fluorescent protein (BFP). A CRISPR enzyme may be fused to a gene sequence encoding a protein or a fragment of a protein that bind DNA molecules or bind other cellular molecules, including but not limited to maltose binding protein (MBP), S-tag, Lex A DNA binding domain (DBD) fusions, GALA DNA binding domain fusions, and herpes simplex virus (HSV) BP16 protein fusions. Additional domains that may form part of a fusion protein comprising a CRISPR enzyme are described in US20110059502, incorporated herein by reference. In Chalasani, US2011/0059502, it is stated, "In certain embodiments, the activity of a second domain in a fusion protein can be about 25% to about 90% more specific than the activity of a second domain not in a fusion protein. In some embodiments, a second domain can comprise an endonuclease activity. In certain embodiments, a second domain can comprise a type II endonuclease activity . . . type IIs endonuclease activity (e.g., Fok I . . . )." Accordingly, a functional domain as herein can be a FokI domain. In some embodiments, a tagged CRISPR enzyme is used to identify the location of a target sequence.
In some embodiments, a CRISPR enzyme may form a component of an inducible system. The inducible nature of the system would allow for spatiotemporal control of gene editing or gene expression using a form of energy. The form of energy may include but is not limited to electromagnetic radiation, sound energy, chemical energy and thermal energy. Examples of inducible system include tetracycline inducible promoters (Tet-On or Tet-Off), small molecule two-hybrid transcription activations systems (FKBP, ABA, etc), or light inducible systems (Phytochrome, LOV domains, or cryptochorome). In one embodiment, the CRISPR enzyme may be a part of a Light Inducible Transcriptional Effector (LITE) to direct changes in transcriptional activity in a sequence-specific manner. The components of a light may include a CRISPR enzyme, a light-responsive cytochrome heterodimer (e.g. from Arabidopsis thaliana), and a transcriptional activation/repression domain. Further examples of inducible DNA binding proteins and methods for their use are provided in U.S. 61/736,465 and U.S. 61/721,283, which is hereby incorporated by reference in its entirety.
In some aspects, the invention comprehends delivering one or more polynucleotides, such as or one or more vectors as described herein, one or more transcripts thereof, and/or one or proteins transcribed therefrom, to a host cell. In some aspects, the invention comprehends cells produced by such methods, and animals comprising or produced from such cells. In some embodiments, a CRISPR enzyme in combination with (and optionally complexed with) a guide sequence is delivered to a cell. Conventional viral and non-viral based gene transfer methods can be used to introduce nucleic acids in mammalian cells or target tissues. Such methods can be used to administer nucleic acids encoding components of a CRISPR system to cells in culture, or in a host organism. Non-viral vector delivery systems include DNA plasmids, RNA (e.g. a transcript of a vector described herein), naked nucleic acid, and nucleic acid complexed with a delivery vehicle, such as a liposome. Viral vector delivery systems include DNA and RNA viruses, which have either episomal or integrated genomes after delivery to the cell. For a review of gene therapy procedures, see Anderson, Science 256:808-813 (1992); Nabel & Felgner, TIBTECH 11:211-217 (1993); Mitani & Caskey, TIBTECH 11:162-166 (1993); Dillon, TIBTECH 11:167-175 (1993); Miller, Nature 357:455-460 (1992); Van Brunt, Biotechnology 6(10):1149-1154 (1988); Vigne, Restorative Neurology and Neuroscience 8:35-36 (1995); Kremer & Perricaudet, British Medical Bulletin 51(1):31-44 (1995); Haddada et al., in Current Topics in Microbiology and Immunology Doerfler and Bohm (eds) (1995); and Yu et al., Gene Therapy 1:13-26 (1994).
In some embodiments, a host cell contains the target sequence, and the cell can be derived from cells taken from a subject, such as a cell line. A wide variety of cell lines for tissue culture are known in the art. Examples of cell lines include, but are not limited to, C8161, CCRF-CEM, MOLT, mIMCD-3, NHDF, HeLa-S3, Huh1, Huh4, Huh7, HUVEC, HASMC, HEKn, HEKa, MiaPaCell, Panc1, PC-3, TF1, CTLL-2, C1R, Rat6, CV1, RPTE, A10, T24, J82, A375, ARH-77, Calu1, SW480, SW620, SKOV3, SK-UT, CaCo2, P388D1, SEM-K2, WEHI-231, HB56, TIB55, Jurkat, J45.01, LRMB, Bcl-1, BC-3, IC21, DLD2, Raw264.7, NRK, NRK-52E, MRC5, MEF, Hep G2, HeLa B, HeLa T4, COS, COS-1, COS-6, COS-M6A, BS-C-1 monkey kidney epithelial, BALB/3T3 mouse embryo fibroblast, 3T3 Swiss, 3T3-L1, 132-d5 human fetal fibroblasts; 10.1 mouse fibroblasts, 293-T, 3T3, 721, 9L, A2780, A2780ADR, A2780cis, A172, A20, A253, A431, A-549, ALC, B16, B35, BCP-1 cells, BEAS-2B, bEnd.3, BHK-21, BR 293, BxPC3, C3H-10T1/2, C6/36, Cal-27, CHO, CHO-7, CHO-IR, CHO-K1, CHO-K2, CHO-T, CHO Dhfr-/-, COR-L23, COR-L23/CPR, COR-L23/5010, COR-L23/R23, COS-7, COV-434, CML T1, CMT, CT26, D17, DH82, DU145, DuCaP, EL4, EM2, EM3, EMT6/AR1, EMT6/AR10.0, FM3, H1299, H69, HB54, HB55, HCA2, HEK-293, HeLa, Hepa1c1c7, HL-60, HMEC, HT-29, Jurkat, JY cells, K562 cells, Ku812, KCL22, KG1, KYO1, LNCap, Ma-Mel 1-48, MC-38, MCF-7, MCF-10A, MDA-MB-231, MDA-MB-468, MDA-MB-435, MDCK II, MDCK II, MOR/0.2R, MONO-MAC 6, MTD-1A, MyEnd, NCI-H69/CPR, NCI-H69/LX10, NCI-H69/LX20, NCI-H69/LX4, NIH-3T3, NALM-1, NW-145, OPCN/OPCT cell lines, Peer, PNT-1A/PNT 2, RenCa, RIN-5F, RMA/RMAS, Saos-2 cells, Sf-9, SkBr3, T2, T-47D, T84, THP1 cell line, U373, U87, U937, VCaP, Vero cells, WM39, WT-49, X63, YAC-1, YAR, and transgenic varieties thereof. Cell lines are available from a variety of sources known to those with skill in the art (see, e.g., the American Type Culture Collection (ATCC) (Manassas, Va.)). In some embodiments, a cell transfected with one or more vectors described herein is used to establish a new cell line comprising one or more vector-derived sequences. In some embodiments, a cell transiently transfected with the components of a CRISPR system as described herein (such as by transient transfection of one or more vectors, or transfection with RNA), and modified through the activity of a CRISPR complex, is used to establish a new cell line comprising cells containing the modification but lacking any other exogenous sequence. In some embodiments, cells transiently or non-transiently transfected with one or more vectors described herein, or cell lines derived from such cells are used in assessing one or more test compounds. Target sequence(s) can be in such cells.
With recent advances in crop genomics, the ability to use CRISPR-Cas9 systems to perform efficient and cost effective gene editing and manipulation will allow the rapid selection and comparison of single and multiplexed genetic manipulations to transform such genomes for improved production and enhanced traits. In this regard reference is made to US patents and publications: U.S. Pat. No. 6,603,061--Agrobacterium-Mediated Plant Transformation Method; U.S. Pat. No. 7,868,149--Plant Genome Sequences and Uses Thereof and US 2009/0100536--Transgenic Plants with Enhanced Agronomic Traits, all the contents and disclosure of each of which are herein incorporated by reference in their entirety. In the practice of the invention, the contents and disclosure of Morrell et al "Crop genomics: advances and applications" Nat Rev Genet. 2011 Dec. 29; 13(2):85-96 are also herein incorporated by reference in their entirety. In an advantageous embodiment of the invention, the CRISPR/Cas9 system is used to engineer microalgae. Thus, target polynucleotides in the invention can be plant, algae, prokaryotic or eukaryotic.
CRISPR systems can be useful for creating an animal or cell that may be used as a disease model. Thus, identification of target sequences for CRISPR systems can be useful for creating an animal or cell that may be used as a disease model. As used herein, "disease" refers to a disease, disorder, or indication in a subject. For example, a method of the invention may be used to create an animal or cell that comprises a modification in one or more nucleic acid sequences associated with a disease, or an animal or cell in which the expression of one or more nucleic acid sequences associated with a disease are altered. Such a nucleic acid sequence may encode a disease associated protein sequence or may be a disease associated control sequence.
In some methods, the disease model can be used to study the effects of mutations on the animal or cell and development and/or progression of the disease using measures commonly used in the study of the disease. Alternatively, such a disease model is useful for studying the effect of a pharmaceutically active compound on the disease.
In some methods, the disease model can be used to assess the efficacy of a potential gene therapy strategy. That is, a disease-associated gene or polynucleotide can be modified such that the disease development and/or progression is inhibited or reduced. In particular, the method comprises modifying a disease-associated gene or polynucleotide such that an altered protein is produced and, as a result, the animal or cell has an altered response. Accordingly, in some methods, a genetically modified animal may be compared with an animal predisposed to development of the disease such that the effect of the gene therapy event may be assessed.
CRISPR systems can be used to develop a biologically active agent that modulates a cell signaling event associated with a disease gene; and hence, identifying target sequences can be so used.
CRISPR systems can be used to develop a cell model or animal model can be constructed in combination with the method of the invention for screening a cellular function change; and hence, identifying target sequences can be so used. Such a model may be used to study the effects of a genome sequence modified by the CRISPR complex of the invention on a cellular function of interest. For example, a cellular function model may be used to study the effect of a modified genome sequence on intracellular signaling or extracellular signaling. Alternatively, a cellular function model may be used to study the effects of a modified genome sequence on sensory perception. In some such models, one or more genome sequences associated with a signaling biochemical pathway in the model are modified.
An altered expression of one or more genome sequences associated with a signaling biochemical pathway can be determined by assaying for a difference in the mRNA levels of the corresponding genes between the test model cell and a control cell, when they are contacted with a candidate agent. Alternatively, the differential expression of the sequences associated with a signaling biochemical pathway is determined by detecting a difference in the level of the encoded polypeptide or gene product. To assay for an agent-induced alteration in the level of mRNA transcripts or corresponding polynucleotides, nucleic acid contained in a sample is first extracted according to standard methods in the art. For instance, mRNA can be isolated using various lytic enzymes or chemical solutions according to the procedures set forth in Sambrook et al. (1989), or extracted by nucleic-acid-binding resins following the accompanying instructions provided by the manufacturers. The mRNA contained in the extracted nucleic acid sample is then detected by amplification procedures or conventional hybridization assays (e.g. Northern blot analysis) according to methods widely known in the art or based on the methods exemplified herein.
For purpose of this invention, amplification means any method employing a primer and a polymerase capable of replicating a target sequence with reasonable fidelity. Amplification may be carried out by natural or recombinant DNA polymerases such as TaqGold.TM., T7 DNA polymerase, Klenow fragment of E. coli DNA polymerase, and reverse transcriptase. A preferred amplification method is PCR. In particular, the isolated RNA can be subjected to a reverse transcription assay that is coupled with a quantitative polymerase chain reaction (RT-PCR) in order to quantify the expression level of a sequence associated with a signaling biochemical pathway.
Detection of the gene expression level can be conducted in real time in an amplification assay. In one aspect, the amplified products can be directly visualized with fluorescent DNA-binding agents including but not limited to DNA intercalators and DNA groove binders. Because the amount of the intercalators incorporated into the double-stranded DNA molecules is typically proportional to the amount of the amplified DNA products, one can conveniently determine the amount of the amplified products by quantifying the fluorescence of the intercalated dye using conventional optical systems in the art. DNA-binding dye suitable for this application include SYBR green, SYBR blue, DAPI, propidium iodine, Hoeste, SYBR gold, ethidium bromide, acridines, proflavine, acridine orange, acriflavine, fluorcoumanin, ellipticine, daunomycin, chloroquine, distamycin D, chromomycin, homidium, mithramycin, ruthenium polypyridyls, anthramycin, and the like.
In another aspect, other fluorescent labels such as sequence specific probes can be employed in the amplification reaction to facilitate the detection and quantification of the amplified products. Probe-based quantitative amplification relies on the sequence-specific detection of a desired amplified product. It utilizes fluorescent, target-specific probes (e.g., TaqMan.RTM. probes) resulting in increased specificity and sensitivity. Methods for performing probe-based quantitative amplification are well established in the art and are taught in U.S. Pat. No. 5,210,015.
In yet another aspect, conventional hybridization assays using hybridization probes that share sequence homology with sequences associated with a signaling biochemical pathway can be performed. Typically, probes are allowed to form stable complexes with the sequences associated with a signaling biochemical pathway contained within the biological sample derived from the test subject in a hybridization reaction. It will be appreciated by one of skill in the art that where antisense is used as the probe nucleic acid, the target polynucleotides provided in the sample are chosen to be complementary to sequences of the antisense nucleic acids. Conversely, where the nucleotide probe is a sense nucleic acid, the target polynucleotide is selected to be complementary to sequences of the sense nucleic acid.
Hybridization can be performed under conditions of various stringency. Suitable hybridization conditions for the practice of the present invention are such that the recognition interaction between the probe and sequences associated with a signaling biochemical pathway is both sufficiently specific and sufficiently stable. Conditions that increase the stringency of a hybridization reaction are widely known and published in the art. See, for example, (Sambrook, et al., (1989); Nonradioactive In Situ Hybridization Application Manual, Boehringer Mannheim, second edition). The hybridization assay can be formed using probes immobilized on any solid support, including but are not limited to nitrocellulose, glass, silicon, and a variety of gene arrays. A preferred hybridization assay is conducted on high-density gene chips as described in U.S. Pat. No. 5,445,934.
For a convenient detection of the probe-target complexes formed during the hybridization assay, the nucleotide probes are conjugated to a detectable label. Detectable labels suitable for use in the present invention include any composition detectable by photochemical, biochemical, spectroscopic, immunochemical, electrical, optical or chemical means. A wide variety of appropriate detectable labels are known in the art, which include fluorescent or chemiluminescent labels, radioactive isotope labels, enzymatic or other ligands. In preferred embodiments, one will likely desire to employ a fluorescent label or an enzyme tag, such as digoxigenin, .beta.-galactosidase, urease, alkaline phosphatase or peroxidase, avidin/biotin complex.
The detection methods used to detect or quantify the hybridization intensity will typically depend upon the label selected above. For example, radiolabels may be detected using photographic film or a phosphoimager. Fluorescent markers may be detected and quantified using a photodetector to detect emitted light. Enzymatic labels are typically detected by providing the enzyme with a substrate and measuring the reaction product produced by the action of the enzyme on the substrate; and finally colorimetric labels are detected by simply visualizing the colored label.
An agent-induced change in expression of sequences associated with a signaling biochemical pathway can also be determined by examining the corresponding gene products. Determining the protein level typically involves a) contacting the protein contained in a biological sample with an agent that specifically bind to a protein associated with a signaling biochemical pathway; and (b) identifying any agent:protein complex so formed. In one aspect of this embodiment, the agent that specifically binds a protein associated with a signaling biochemical pathway is an antibody, preferably a monoclonal antibody. The reaction is performed by contacting the agent with a sample of the proteins associated with a signaling biochemical pathway derived from the test samples under conditions that will allow a complex to form between the agent and the proteins associated with a signaling biochemical pathway. The formation of the complex can be detected directly or indirectly according to standard procedures in the art. In the direct detection method, the agents are supplied with a detectable label and unreacted agents may be removed from the complex; the amount of remaining label thereby indicating the amount of complex formed. For such method, it is preferable to select labels that remain attached to the agents even during stringent washing conditions. It is preferable that the label does not interfere with the binding reaction. In the alternative, an indirect detection procedure may use an agent that contains a label introduced either chemically or enzymatically. A desirable label generally does not interfere with binding or the stability of the resulting agent:polypeptide complex. However, the label is typically designed to be accessible to an antibody for an effective binding and hence generating a detectable signal. A wide variety of labels suitable for detecting protein levels are known in the art. Non-limiting examples include radioisotopes, enzymes, colloidal metals, fluorescent compounds, bioluminescent compounds, and chemiluminescent compounds.
The amount of agent:polypeptide complexes formed during the binding reaction can be quantified by standard quantitative assays. As illustrated above, the formation of agent:polypeptide complex can be measured directly by the amount of label remained at the site of binding. In an alternative, the protein associated with a signaling biochemical pathway is tested for its ability to compete with a labeled analog for binding sites on the specific agent. In this competitive assay, the amount of label captured is inversely proportional to the amount of protein sequences associated with a signaling biochemical pathway present in a test sample.
A number of techniques for protein analysis based on the general principles outlined above are available in the art. They include but are not limited to radioimmunoassays, ELISA (enzyme linked immunoradiometric assays), "sandwich" immunoassays, immunoradiometric assays, in situ immunoassays (using e.g., colloidal gold, enzyme or radioisotope labels), western blot analysis, immunoprecipitation assays, immunofluorescent assays, and SDS-PAGE.
Antibodies that specifically recognize or bind to proteins associated with a signaling biochemical pathway are preferable for conducting the aforementioned protein analyses. Where desired, antibodies that recognize a specific type of post-translational modifications (e.g., signaling biochemical pathway inducible modifications) can be used. Post-translational modifications include but are not limited to glycosylation, lipidation, acetylation, and phosphorylation. These antibodies may be purchased from commercial vendors. For example, anti-phosphotyrosine antibodies that specifically recognize tyrosine-phosphorylated proteins are available from a number of vendors including Invitrogen and Perkin Elmer. Anti-phosphotyrosine antibodies are particularly useful in detecting proteins that are differentially phosphorylated on their tyrosine residues in response to an ER stress. Such proteins include but are not limited to eukaryotic translation initiation factor 2 alpha (eIF-2.alpha.). Alternatively, these antibodies can be generated using conventional polyclonal or monoclonal antibody technologies by immunizing a host animal or an antibody-producing cell with a target protein that exhibits the desired post-translational modification.
It may be desirable to discern the expression pattern of an protein associated with a signaling biochemical pathway in different bodily tissue, in different cell types, and/or in different subcellular structures. These studies can be performed with the use of tissue-specific, cell-specific or subcellular structure specific antibodies capable of binding to protein markers that are preferentially expressed in certain tissues, cell types, or subcellular structures.
An altered expression of a gene associated with a signaling biochemical pathway can also be determined by examining a change in activity of the gene product relative to a control cell. The assay for an agent-induced change in the activity of a protein associated with a signaling biochemical pathway will dependent on the biological activity and/or the signal transduction pathway that is under investigation. For example, where the protein is a kinase, a change in its ability to phosphorylate the downstream substrate(s) can be determined by a variety of assays known in the art. Representative assays include but are not limited to immunoblotting and immunoprecipitation with antibodies such as anti-phosphotyrosine antibodies that recognize phosphorylated proteins. In addition, kinase activity can be detected by high throughput chemiluminescent assays such as AlphaScreen.TM. (available from Perkin Elmer) and eTag.TM. assay (Chan-Hui, et al. (2003) Clinical Immunology 111: 162-174).
Where the protein associated with a signaling biochemical pathway is part of a signaling cascade leading to a fluctuation of intracellular pH condition, pH sensitive molecules such as fluorescent pH dyes can be used as the reporter molecules. In another example where the protein associated with a signaling biochemical pathway is an ion channel, fluctuations in membrane potential and/or intracellular ion concentration can be monitored. A number of commercial kits and high-throughput devices are particularly suited for a rapid and robust screening for modulators of ion channels. Representative instruments include FLIPR.TM. (Molecular Devices, Inc.) and VIPR (Aurora Biosciences). These instruments are capable of detecting reactions in over 1000 sample wells of a microplate simultaneously, and providing real-time measurement and functional data within a second or even a minisecond.
In practicing any of the methods disclosed herein, a suitable vector can be introduced to a cell or an embryo via one or more methods known in the art, including without limitation, microinjection, electroporation, sonoporation, biolistics, calcium phosphate-mediated transfection, cationic transfection, liposome transfection, dendrimer transfection, heat shock transfection, nucleofection transfection, magnetofection, lipofection, impalefection, optical transfection, proprietary agent-enhanced uptake of nucleic acids, and delivery via liposomes, immunoliposomes, virosomes, or artificial virions. In some methods, the vector is introduced into an embryo by microinjection. The vector or vectors may be microinjected into the nucleus or the cytoplasm of the embryo. In some methods, the vector or vectors may be introduced into a cell by nucleofection.
The target polynucleotide of a CRISPR complex can be any polynucleotide endogenous or exogenous to the eukaryotic cell. For example, the target polynucleotide can be a polynucleotide residing in the nucleus of the eukaryotic cell. The target polynucleotide can be a sequence coding a gene product (e.g., a protein) or a non-coding sequence (e.g., a regulatory polynucleotide or a junk DNA).
Examples of target polynucleotides include a sequence associated with a signaling biochemical pathway, e.g., a signaling biochemical pathway-associated gene or polynucleotide. Examples of target polynucleotides include a disease associated gene or polynucleotide. A "disease-associated" gene or polynucleotide refers to any gene or polynucleotide which is yielding transcription or translation products at an abnormal level or in an abnormal form in cells derived from a disease-affected tissues compared with tissues or cells of a non disease control. It may be a gene that becomes expressed at an abnormally high level; it may be a gene that becomes expressed at an abnormally low level, where the altered expression correlates with the occurrence and/or progression of the disease. A disease-associated gene also refers to a gene possessing mutation(s) or genetic variation that is directly responsible or is in linkage disequilibrium with a gene(s) that is responsible for the etiology of a disease. The transcribed or translated products may be known or unknown, and may be at a normal or abnormal level.
The target polynucleotide of a CRISPR complex can be any polynucleotide endogenous or exogenous to the eukaryotic cell. For example, the target polynucleotide can be a polynucleotide residing in the nucleus of the eukaryotic cell. The target polynucleotide can be a sequence coding a gene product (e.g., a protein) or a non-coding sequence (e.g., a regulatory polynucleotide or a junk DNA).
The target polynucleotide of a CRISPR complex may include a number of disease-associated genes and polynucleotides as well as signaling biochemical pathway-associated genes and polynucleotides as listed in US provisional patent applications 61/736,527 and 61/748,427 having Broad reference BI-2011/008/WSGR Docket No. 44063-701.101 and BI-2011/008/WSGR Docket No. 44063-701.102 respectively, both entitled SYSTEMS METHODS AND COMPOSITIONS FOR SEQUENCE MANIPULATION filed on Dec. 12, 2012 and Jan. 2, 2013, respectively, the contents of all of which are herein incorporated by reference in their entirety.
Examples of target polynucleotides include a sequence associated with a signaling biochemical pathway, e.g., a signaling biochemical pathway-associated gene or polynucleotide. Examples of target polynucleotides include a disease associated gene or polynucleotide. A "disease-associated" gene or polynucleotide refers to any gene or polynucleotide which is yielding transcription or translation products at an abnormal level or in an abnormal form in cells derived from a disease-affected tissues compared with tissues or cells of a non disease control. It may be a gene that becomes expressed at an abnormally high level; it may be a gene that becomes expressed at an abnormally low level, where the altered expression correlates with the occurrence and/or progression of the disease. A disease-associated gene also refers to a gene possessing mutation(s) or genetic variation that is directly responsible or is in linkage disequilibrium with a gene(s) that is responsible for the etiology of a disease. The transcribed or translated products may be known or unknown, and may be at a normal or abnormal level.
Embodiments of the invention also relate to methods and compositions related to knocking out genes, amplifying genes and repairing particular mutations associated with DNA repeat instability and neurological disorders (Robert D. Wells, Tetsuo Ashizawa, Genetic Instabilities and Neurological Diseases, Second Edition, Academic Press, Oct. 13, 2011--Medical). Specific aspects of tandem repeat sequences have been found to be responsible for more than twenty human diseases (New insights into repeat instability: role of RNA DNA hybrids. Mclvor El, Polak U. Napierala M. RNA Biol. 2010 September-October; 7(5):551-8). The CRISPR-Cas system may be harnessed to correct these defects of genomic instability. And thus, target sequences can be found in these defects of genomic instability.
Further embodiments of the invention relate to algorithms that lay the foundation of methods relating to CRISPR enzyme, e.g. Cas, specificity or off-target activity. In general, algorithms refer to an effective method expressed as a finite list of well defined instructions for calculating one or more functions of interest. Algorithms may be expressed in several kinds of notation, including but not limited to programming languages, flow charts, control tables, natural languages, mathematical formula and pseudocode. In a preferred embodiment, the algorithm may be expressed in a programming language that expresses the algorithm in a form that may be executed by a computer or a computer system.
Methods relating to CRISPR enzyme, e.g. Cas, specificity or off-target activity are based on algorithms that include but are not limited to the thermodynamic algorithm, multiplicative algorithm and positional algorithm. These algorithms take in an input of a sequence of interest and identify candidate target sequences to then provide an output of a ranking of candidate target sequences or a score associated with a particular target sequence based on predicted off-target sites. Candidate target sites may be selected by an end user or a customer based on considerations which include but are not limited to modification efficiency, number, or location of predicted off-target cleavage. In a more preferred embodiment, a candidate target site is unique or has minimal predicted off-target cleavage given the previous parameters. However, the functional relevance of potential off-target modification should also be considered when choosing a target site. In particular, an end user or a customer may consider whether the off-target sites occur within loci of known genetic function, i.e. protein-coding exons, enhancer regions, or intergenic regulatory elements. There may also be cell-type specific considerations, i.e. if an off-target site occurs in a locus that is not functionally relevant in the target cell type. Taken together, a end user or customer may then make an informed, application-specific selection of a candidate target site with minimal off-target modification.
The thermodynamic algorithm may be applied in selecting a CRISPR complex for targeting and/or cleavage of a candidate target nucleic acid sequence within a cell. The first step is to input the target sequence (Step S400) which may have been determined using the positional algorithm. A CRISPR complex is also input (Step S402). The next step is to compare the target sequence with the guide sequence for the CRISPR complex (Step S404) to identify any mismatches. Furthermore, the amount, location and nature of the mismatch(es) between the guide sequence of the potential CRISPR complex and the candidate target nucleic acid sequence may be determined. The hybridization free energy of binding between the target sequence and the guide sequence is then calculated (Step S406). For example, this may be calculated by determining a contribution of each of the amount, location and nature of mismatch(es) to the hybridization free energy of binding between the target nucleic acid sequence and the guide sequence of potential CRISPR complex(es). Furthermore, this may be calculated by applying a model calculated using a training data set as explained in more detail below. Based on the hybridization free energy (i.e. based on the contribution analysis) a prediction of the likelihood of cleavage at the location(s) of the mismatch(es) of the target nucleic acid sequence by the potential CRISPR complex(es) is generated (Step S408). The system then determines whether or not there are any additional CRISPR complexes to consider and if so repeats the comparing, calculating and predicting steps. Each CRISPR complex is selected from the potential CRISPR complex(es) based on whether the prediction indicates that it is more likely than not that cleavage will occur at location(s) of mismatch(es) by the CRISPR complex (Step S410). Optionally, the probabilities of cleavage may be ranked so that a unique CRISPR complex is selected. Determining the contribution of each of the amount, location and nature of mismatch(es) to hybridization free energy includes but is not limited to determining the relative contribution of these factors. The term "location" as used in the term "location of mismatch(es)" may refer to the actual location of the one or more base pair mismatch(es) but may also include the location of a stretch of base pairs that flank the base pair mismatch(es) or a range of locations/positions. The stretch of base pairs that flank the base pair mismatch(es) may include but are not limited to at least one, at least two, at least three base pairs, at least four or at least five or more base pairs on either side of the one or more mismatch(es). As used herein, the "hybridization free energy" may be an estimation of the free energy of binding, e.g. DNA:RNA free energy of binding which may be estimated from data on DNA:DNA free energy of binding and RNA:RNA free energy of binding.
In methods relating to the multiplicative algorithm applied in identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises: a) creating a data training set as to a particular Cas, b) determining average cutting frequency at a particular position for the particular Cas from the data training set, c) determining average cutting frequency of a particular mismatch for the particular Cas from the data training set, d) multiplying the average cutting frequency at a particular position by the average cutting frequency of a particular mismatch to obtain a first product, e) repeating steps b) to d) to obtain second and further products for any further particular position (s) of mismatches and particular mismatches and multiplying those second and further products by the first product, for an ultimate product, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position (or optionally e) repeating steps b) to d) to obtain second and further products for any further particular position (s) of mismatches and particular mismatches and multiplying those second and further products by the first product, for an ultimate product, and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position), and f) multiplying the ultimate product by the result of dividing the minimum distance between consecutive mismatches by 18 and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position (or optionally f) multiplying the ultimate product by the result of dividing the minimum distance between consecutive mismatches by 18 and omitting this step if there is no mismatch at any position or if there is only one particular mismatch at one particular position), to thereby obtain a ranking, which allows for the identification of one or more unique target sequences, the predicted cutting frequencies for genome-wide targets may be calculated by multiplying, in series: f.sub.est=f(1)g(N.sub.1,N.sub.1').times.f(2)g(N.sub.2N.sub.2').times. . . . f(19)g(N.sub.19,N.sub.19').times.h with values f(i) and g(N.sub.i,N.sub.i') at position i corresponding, respectively, to the aggregate position- and base-mismatch cutting frequencies for positions and pairings indicated in a generalized base transition matrix or an aggregate matrix, e.g. a matrix as indicated in FIG. 12c. Each frequency was normalized to range from 0 to 1, such that f.fwdarw.(f-f.sub.min)/(f.sub.max-f.sub.min). In case of a match, both were set equal to 1. The value h meanwhile re-weighted the estimated frequency by the minimum pairwise distance between consecutive mismatches in the target sequence. This value distance, in base-pairs, was divided by 18 to give a maximum value of 1 (in cases where fewer than 2 mismatches existed, or where mismatches occurred on opposite ends of the 19 bp target-window). Samples having a read-count of at least 10,000 (n=43) were plotted. Those tied in rank were given a rank-average. The Spearman correlation coefficient, 0.58, indicated that the estimated frequencies recapitulated 58% of the rank-variance for the observed cutting frequencies. Comparing f.sub.est with the cutting frequencies directly yielded a Pearson correlation of 0.89. While dominated by the highest-frequency gRNA/target pairs, this value indicated that nearly 90% of all cutting-frequency variance was explained by the predictions above. In further aspects of the invention, the multiplicative algorithm or the methods mentioned herein may also include thermodynamic factors, e.g. hybridization energies, or other factors of interest being multiplied in series to arrive at the ultimate product.
In embodiments of the invention, determining the off-target activity of a CRISPR enzyme may allow an end user or a customer to predict the best cutting sites in a genomic locus of interest. In a further embodiment of the invention, one may obtain a ranking of cutting frequencies at various putative off-target sites to verify in vitro, in vivo or ex vivo if one or more of the worst case scenario of non-specific cutting does or does not occur. In another embodiment of the invention, the determination of off-target activity may assist with selection of specific sites if an end user or customer is interested in maximizing the difference between on-target cutting frequency and the highest cutting frequency obtained in the ranking of off-target sites. Another aspect of selection includes reviewing the ranking of sites and identifying the genetic loci of the non-specific targets to ensure that a specific target site selected has the appropriate difference in cutting frequency from say targets that may encode for oncogenes or other genetic loci of interest. Aspects of the invention may include methods of minimizing therapeutic risk by verifying the off-target activity of the CRISPR-Cas complex. Further aspects of the invention may include utilizing information on off-target activity of the CRISPR-Cas complex to create specific model systems (e.g. mouse) and cell lines. The methods of the invention allow for rapid analysis of non-specific effects and may increase the efficiency of a laboratory.
In methods relating to the positional algorithm applied in identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system, wherein the method comprises: a) determining average cutting frequency of guide-RNA/target mismatches at a particular position for a particular Cas from a training data set as to that Cas, if more than one mismatch, repeat step a) so as to determine cutting frequency for each mismatch, multiply frequencies of mismatches to thereby obtain a ranking, which allows for the identification of one or more unique target sequences, an example of an application of this algorithm may be seen in FIG. 23.
FIGS. 32, 33A, 33B and 34, respectively, each show a flow diagram of methods of the invention. FIG. 32 provides a flow diagram as to locational or positional methods of the invention, i.e., with respect to computational identification of unique CRISPR target sites: To identify unique target sites for a Cas, e.g., a Cas9, e.g., the S. pyogenes SF370 Cas9 (SpCas9) enzyme, in nucleic acid molecules, e.g., of cells, e.g., of organisms, which include but are not limited to human, mouse, rat, zebrafish, fruit fly, and C. elegans genome, Applicants developed a software package to scan both strands of a DNA sequence and identify all possible SpCas9 target sites. The method is shown in FIG. 32 which shows that the first step is to input the genome sequence (Step S100). The CRISPR motif(s) which are suitable for this genome sequence are then selected (Step S102). For this example, the CRISPR motif is an NGG protospacer adjacent motif (PAM) sequence. A fragment of fixed length which needs to occur in the overall sequence before the selected motif (i.e. upstream in the sequence) is then selected (Step S102). In this case, the fragment is a 20 bp sequence. Thus, each SpCas9 target site was is operationally defined as a 20 bp sequence followed by an NGG protospacer adjacent motif (PAM) sequence, and all sequences satisfying this 5'-N20-NGG-3' definition on all chromosomes were identified (Step S106). To prevent non-specific genome editing, after identifying all potential sites, all target sites were filtered based on the number of times they appear in the relevant reference genome (Step S108). (Essentially, all the 20-bp fragments (candidate target sites) upstream of the NGG PAM motif are aggregated. If a particular 20-bp fragment occurs more than once in your genome-wide search, it is considered not unique and `strikes out`, aka filtered. The 20-bp fragments that REMAIN therefore occur only once in the target genome, making it unique; and, instead of taking a 20-bp fragment (the full Cas9 target site), this algorithm takes the first, for example, 11-12 bp upstream of the PAM motif and requires that to be unique.) Finally, a unique target site is selected (Step S110), e.g. To take advantage of sequence specificity of Cas, e.g., Cas9 activity conferred by a `seed` sequence, which can be, for example, approximately 11-12 bp sequence 5' from the PAM sequence, 5'-NNNNNNNNNN-NGG-3' sequences were selected to be unique in the relevant genome. Genomic sequences are available on the UCSC Genome Browser and sample visualizations of the information for the Human genome hg, Mouse genome mm, Rat genome rn, Zebrafish genome danRer, D. melanogaster genome dm, C. elegans genome cc, the pig genome and cow genome are shown in FIGS. 15 through 22 respectively.
FIGS. 33A and 33B each provides a flow diagram as to thermodynamic methods of the invention. FIG. 34 provides a flow diagram as to multiplication methods of the invention. Referring to FIGS. 33A and 33B, and considering the least squares thermodynamic model of CRISPR-Cas cutting efficiency, for arbitrary Cas9 target sites, Applicants generated a numerical thermodynamic model that predicts Cas9 cutting efficiency. Applicants propose 1) that the Cas9 guide RNA has specific free energies of hybridization to its target and any off-target DNA sequences and 2) that Cas9 modifies RNA:DNA hybridization free-energies locally in a position-dependent but sequence-independent way. Applicants trained a model for predicting CRISPR-Cas cutting efficiency based on their CRISPR-Cas guide RNA mutation data and RNA:DNA thermodynamic free energy calculations using a machine learning algorithm. Applicants then validated their resulting models by comparing their predictions of CRISPR-Cas off-target cutting at multiple genomic loci with experimental data assessing locus modification at the same sites. The methodology adopted in developing this algorithm is as follows: The problem summary states that for arbitrary spacers and targets of constant length, a numerical model that makes thermodynamic sense and predicts Cas9 cutting efficiency is to be found. Suppose Cas9 modifies DNA:RNA hybridization free-energies locally in a position-dependent but sequence-independent way. The first step is to define a model having a set a weights which links the free energy of hybridization Z with the local free energies G (Step S200). Then for DNA:RNA hybridization free energies .DELTA.G.sub.ij(k) (for position k between 1 and N) of spacer i and target j
.times..times..alpha..times..DELTA..times..times..function. ##EQU00003##
Z.sub.ij can be treated as an "effective" free-energy modified by the multiplicative position-weights .alpha..sub.k. The "effective" free-energy Z.sub.ij corresponds to an associated cutting-probability .about.e.sup.-.beta.Z.sup.ij (for some constant .beta.) in the same way that an equilibrium model of hybridization (without position-weighting) would have predicted a hybridization-probability .about.e.sup.-.beta..DELTA.G.sup.ij. Since cutting-efficiency has been measured, the values Z.sub.ij can be treated as their observables. Meanwhile, .DELTA.G.sub.ij(k) can be calculated for any experiment's spacer-target pairing. Applicants task was to find the values .alpha..sub.k, since this would allow them to estimate Z.sub.ij for any spacer-target pair. The weights are determined by inputting known values for Z and G from a training set of sequences with the known values being determined by experimentation as necessary. Thus, Applicants need to define a training set of sequences (Step S202) and calculate a value of Z for each sequence in the training set (Step S204). Writing the above equation for Z.sub.ij in matrix form Applicants get: {right arrow over (Z)}=G{right arrow over (.alpha.)} (1)
The least-squares estimate is then {right arrow over (.alpha.)}.sub.est=(G.sup.TG).sub.-1G.sup.T{right arrow over (Z)}
where G.sup.T is the matrix-transpose of the G and (G.sup.TG).sup.-1 is the inverse of their matrix-product. In the above G is a matrix of local DNA:RNA free-energy values whose rth row corresponds to experimental trial r and whose kth column corresponds to the kth position in the DNA:RNA hybrid tested in that experimental trial. These values of G are thus input into the training system (Step S204). {right arrow over (Z)} is meanwhile a column-vector whose rth row corresponds to observables from the same experimental trial as G's rth row. Because of the relation described above wherein the CRISPR cutting frequencies are estimated to vary as .about.e.sup.-.beta.Z.sup.ij, these observables, Z.sub.ij, were calculated as the natural logarithm of the observed cutting frequency. The observable is the cleavage efficiency of Cas, e.g., Cas9, at a target DNA for a particular guide RNA and target DNA pair. The experiment is Cas, e.g., Cas9, with a particular sgRNA/DNA target pairing, and the observable is the cleavage percentage (whether measured as indel formation percentage from cells or simply cleavage percentage in vitro) (see herein discussion on generating training data set). More in particular, every unique PCR reaction that was sequenced should be treated as a unique experimental trial to encompass replicability within the vector. This means that experimental replicates each go into separate rows of equation 1 (and because of this, some rows of G will be identical). The advantage of this is that when {right arrow over (.alpha.)} is fit, all relevant information--including replicability--is taken into account in the final estimate. Observable {right arrow over (Z)}, values were calculated as log (observed frequency of cutting) (Step S206). Cutting frequencies were optionally normalized identically (so that they all have the same "units") (Step S208). For plugging in sequencing indel-frequency values, it may be best, however, to standardize sequencing depth. The preferred way to do this would be to set a standard sequencing-depth D for which all experiments included in {right arrow over (Z)} have at least that number of reads. Since cutting frequencies below 1/D cannot be consistently detected, this should be set as the minimum frequency for the data-set, and the values in {right arrow over (Z)} should range from log(1/D) to log(1). One could vary the value of D later on to ensure that the {right arrow over (.alpha.)} estimate isn't too dependent on the value chosen. Thus, values of Z could be filtered out if they do not meet the minimum sequencing depth (Step S210). Once the values of G and Z are input to the machine learning system, the weights can be determined (Step S212) and output (Step S214). These weights can then be used to estimate the free energy Z and the cutting frequency for any sequence. In a further aspect, there are different methods of graphing NGG and NNAGAAW sequences. One is with the `non-overlapping` method. NGG and NRG may be regraphed in an "overlapping" fashion, as indicated in FIGS. 6 A-C. Applicants also performed a study on off target Cas9 activity as indicated in FIGS. 10, 11 and 12. Aspects of the invention also relate to predictive models that may not involve hybridization energies but instead simply use the cutting frequency information as a prediction.
FIG. 34 shows the steps in one method relating to the multiplicative algorithm which may be applied in identifying one or more unique target sequences in a genome of a eukaryotic organism, whereby the target sequence is susceptible to being recognized by a CRISPR-Cas system. The method comprises: a) creating a data training set as to a particular Cas. The data training set may be created as described in more detail later by determining the weights associated with a model. Once a data training set has been established, it can be used to predict the behavior of an input sequence and to identify one or more unique target sequences therein. At step S300, the genome sequence is input to the system. For a particular Cas, the next step is to locate a mismatch between a target sequence within the input sequence and guide RNA for the particular Cas (Step S302). For the identified mismatch, two average cutting frequencies are determined using the data training set. These are the average cutting frequency at the position of the mismatch (step S304) and the average cutting frequency associated with that type of mismatch (Step S306). These average cutting frequencies are determined from the data training set which is particular to that Cas. The next step S308 is to create a product by multiplying the average cutting frequency at a particular position by the average cutting frequency of a particular mismatch to obtain a first product. It is then determined at step S310 whether or not there are any other mismatches. If there are none, the target sequence is output as the unique target sequence. However, if there are other mismatches, steps 304 to 308 are repeated to obtain second and further products for any further particular position (s) of mismatches and particular mismatches. Where second and further products are created and all products are multiplied together to create an ultimate product. The ultimate product is then multiplied by the result of dividing the minimum distance between consecutive mismatches by the length of the target sequence (e.g. 18) (step S314) which effectively scales each ultimate product. It will be appreciated that steps 312 and 314 are omitted if there is no mismatch at any position or if there is only one particular mismatch at one particular position. The process is then repeated for any other target sequences. The "scaled" ultimate products for each target sequence are each ranked to thereby obtain a ranking (Step S316), which allows for the identification of one or more unique target sequences by selecting the highest ranked one (Step S318). Thus the "scaled" ultimate product which represents the predicted cutting frequencies for genome-wide targets may be calculated by: f.sub.est=f(1)g(N.sub.1,N.sub.1').times.f(2)g(N.sub.2,N.sub.2').times. . . . f(19)g(N.sub.19,N.sub.19').times.h.
with values f(i) and g(N.sub.i, N.sub.i') at position i corresponding, respectively, to the aggregate position- and base-mismatch cutting frequencies for positions and pairings indicated in a generalized base transition matrix or an aggregate matrix, e.g. a matrix as indicated in FIG. 12c. In other words, f(i) is the average cutting frequency at the particular position for the mismatch and g(N.sub.i, N'.sub.i) is the average cutting frequency for the particular mismatch type for the mismatch. Each frequency was normalized to range from 0 to 1, such that f.fwdarw.(f-f.sub.min)/(f.sub.max-f.sub.min). In case of a match, both were set equal to 1. The value h meanwhile re-weighted the estimated frequency by the minimum pairwise distance between consecutive mismatches in the target sequence. This value distance, in base-pairs, was divided by a constant which was indicative of the length of the target sequence (e.g. 18) to give a maximum value of 1 (in cases where fewer than 2 mismatches existed, or where mismatches occurred on opposite ends of the 19 bp target-window). Samples having a read-count of at least 10,000 (n=43) were plotted. Those tied in rank were given a rank-average. The Spearman correlation coefficient, 0.58, indicated that the estimated frequencies recapitulated 58% of the rank-variance for the observed cutting frequencies. Comparing fir with the cutting frequencies directly yielded a Pearson correlation of 0.89. While dominated by the highest-frequency gRNA/target pairs, this value indicated that nearly 90% of all cutting-frequency variance was explained by the predictions above. In further aspects of the invention, the multiplicative algorithm or the methods mentioned herein may also include thermodynamic factors, e.g. hybridization energies, or other factors of interest being multiplied in series to arrive at the ultimate product.
FIG. 35 shows a schematic block diagram of a computer system which can be used to implement the methods described herein. The computer system 50 comprises a processor 52 coupled to code and data memory 54 and an input/output system 56 (for example comprising interfaces for a network and/or storage media and/or other communications). The code and/or data stored in memory 54 may be provided on a removable storage medium 60. There may also be a user interface 58 for example comprising a keyboard and/or mouse and a user display 62. The computer system is connected to a database 78. The database 78 comprises the data associated with the data training sets. The computer system is shown as a single computing device with multiple internal components which may be implemented from a single or multiple central processing units, e.g. microprocessors. It will be appreciated that the functionality of the device may be distributed across several computing devices. It will also be appreciated that the individual components may be combined into one or more components providing the combined functionality. Moreover, any of the modules, databases or devices shown may be implemented in a general purpose computer modified (e.g. programmed or configured) by software to be a special-purpose computer to perform the functions described herein. The processor may be configured to carry out the steps shown in the various flowcharts. The user interface may be used to input the genome sequence, the CRISPR motif and/or Cas for which a target sequence is to be identified. The output unique target sequence(s) may be displayed on the user display.
EXAMPLES
The following examples are given for the purpose of illustrating various embodiments of the invention and are not meant to limit the present invention in any fashion. The present examples, along with the methods described herein are presently representative of preferred embodiments, are exemplary, and are not intended as limitations on the scope of the invention. Changes therein and other uses which are encompassed within the spirit of the invention as defined by the scope of the claims will occur to those skilled in the art.
Example 1
Evaluation of the Specificity of Cas9-Mediated Genome Cleavage
Applicants carried out an initial test to evaluate the cleavage specificity of Cas9 from Streptococcus pyogenes. The assay was designed to test the effect of single basepair mismatches between the guide RNA sequence and the target DNA. The results from the initial round of testing are depicted in FIG. 3.
Applicants carried out the assay using 293FT cells in 96 well plates. Cells were transfected with 65 ng of a plasmid carrying Cas9 and 10 ng of a PCR amplicon carrying the pol3 promoter U6 and the guide RNA. The experiment was conducted using a high amount of Cas9 and guide RNA, which probably explains the seemingly low specificity (i.e. single base mismatches is not sufficient to abolish cleavage). Applicants also evaluate the effect of different concentration of Cas9 and RNA on cleavage specificity. Additionally, Applicants carry out a comprehensive evaluation of every possible mismatch in each position of the guide RNA. The end goal is to generate a model to inform the design of guide RNAs having high cleavage specificity.
Additional experiments test position and number of mismatches in the guide RNA on cleavage efficiency. The following table shows a list of 48 mismatch possibilities. In the table 0 means no mutation and 1 means with mutation.
TABLE-US-00002 20 19 18 17 16 15 14 13 12 11 10 9 8 7 6 5 4 3 2 1 NGG Test Rule 1: More mismatches = bigger effect on cutting Test Rule 2: Mismatches on 5' end have less effect than mismatches on 3' end 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 2 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 0 0 3 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 0 0 0 0 4 0 0 0 0 0 0 0 0 0 0 0 0 1 1 0 0 0 0 0 0 5 0 0 0 0 0 0 0 0 0 0 1 1 0 0 0 0 0 0 0 0 6 0 0 0 0 0 0 0 0 1 1 0 0 0 0 0 0 0 0 0 0 7 0 0 0 0 0 0 1 1 0 0 0 0 0 0 0 0 0 0 0 0 8 0 0 0 0 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 9 0 0 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 10 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 11 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 1 0 12 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 1 1 1 13 0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 1 0 0 0 0 14 0 0 0 0 0 0 0 0 0 0 0 0 1 1 1 1 1 0 0 0 15 0 0 0 0 0 0 0 0 0 0 1 1 1 0 0 0 0 0 0 0 16 0 0 0 0 0 0 0 0 0 1 1 1 1 1 0 0 0 0 0 0 17 0 0 0 0 0 0 0 1 1 1 0 0 0 0 0 0 0 0 0 0 18 0 0 0 0 0 0 1 1 1 1 1 0 0 0 0 0 0 0 0 0 19 0 0 0 0 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 20 0 0 0 1 1 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0 21 0 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 22 1 1 1 1 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 Test Rule 3: Mismatches more spreadout have less effect than mismatches more concentrated 23 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1 24 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 1 25 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 1 26 0 0 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 1 27 1 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 28 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1 0 0 29 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 1 0 0 30 0 0 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 1 0 0 31 0 0 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 32 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1 0 0 0 0 33 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 1 0 0 0 0 34 0 0 0 0 0 0 1 0 0 0 0 0 0 0 0 1 0 0 0 0 35 0 1 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 0 36 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1 0 1 37 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 0 1 0 0 1 38 0 0 0 0 0 0 0 0 0 0 0 1 0 0 0 1 0 0 0 1 39 0 0 0 0 0 0 0 0 0 1 0 0 0 0 1 0 0 0 0 1 40 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1 0 1 0 0 41 0 0 0 0 0 0 0 0 0 0 0 1 0 0 1 0 0 1 0 0 42 0 0 0 0 0 0 0 0 0 1 0 0 0 1 0 0 0 1 0 0 43 0 0 0 0 0 0 0 1 0 0 0 0 1 0 0 0 0 1 0 0 44 0 0 0 0 0 0 0 0 0 0 0 0 0 1 0 1 0 1 0 1 45 0 0 0 0 0 0 0 0 0 0 1 0 0 1 0 0 1 0 0 1 46 0 0 0 0 0 0 0 1 0 0 0 1 0 0 0 1 0 0 0 1 47 0 0 0 0 1 0 0 0 0 1 0 0 0 0 1 0 0 0 0 1 48 0 1 0 0 0 0 0 1 0 0 0 0 0 1 0 0 0 0 0 1
Example 2
Evaluation of Mutations in the PAM Sequence, and its Effect on Cleavage Efficiency
Applicants tested mutations in the PAM sequence and its effect on cleavage. The PAM sequence for Streptococcus pyogenes Cas9 is NGG, where the GG is thought to be required for cleavage. To test whether Cas9 can cleavage sequences with PAMs that are different than NGG, Applicants chose the following 30 target sites from the Emx1 locus of the human genome--2 for each of the 15 PAM possibilities: NAA, NAC, NAT, NAG, NCA, NCC, NCG, NCT, NTA, NTC, NTG, NTT, NGA, NGC, and NGT; NGG is not selected because it can be targeted efficiently.
The cleavage efficiency data is shown in FIG. 4. The data shows that other than NGG, only sequences with NAG PAMs can be targeted.
TABLE-US-00003 PAM Target 1 Target 2 NAA AGGCCCCAGTGGCTGCTCT TCATCTGTGCCCCTCCCTC NAT ACATCAACCGGTGGCGCAT GGGAGGACATCGATGTCAC NAC AAGGTGTGGTTCCAGAACC CAAACGGCAGAAGCTGGAG NAG CCATCACATCAACCGGTGG GGGTGGGCAACCACAAACC NTA AAACGGGAGAAGCTGGAGG GGTGGGCAACCACAAACCC NTT GGCAGAAGCTGGAGGAGGA GGCTCCCATCACATCAACC NTC GGTGTGGTTCCAGAACCGG GAAGGGCCTGAGTCCGAGC NTG AACCGGAGGACAAAGTACA CAACCGGTGGCGCATTGCC NCA TTCCAGAACCGGAGGACAA AGGAGGAAGGGCCTGAGTC NCT GTGTGGTTCCAGAACCGGA AGCTGGAGGAGGAAGGGCC NCC TCCAGAACCGGAGGACAAA GCATTGCCACGAAGCAGGC NCG CAGAAGCTGGAGGAGGAAG ATTGCCACGAAGCAGGCCA NGA CATCAACCGGTGGCGCATT AGAACCGGAGGACAAAGTA NGT GCAGAAGCTGGAGGAGGAA TCAACCGGTGGCGCATTGC NGC CCTCCCTCCCTGGCCCAGG GAAGCTGGAGGAGGAAGGG
Example 3
Cas9 Diversity and RNAs, PAMS, Targets
The CRISPR-Cas system is an adaptive immune mechanism against invading exogenous DNA employed by diverse species across bacteria and archaea. The type II CRISPR-Cas9 system consists of a set of genes encoding proteins responsible for the "acquisition" of foreign DNA into the CRISPR locus, as well as a set of genes encoding the "execution" of the DNA cleavage mechanism; these include the DNA nuclease (Cas9), a non-coding transactivating cr-RNA (tracrRNA), and an array of foreign DNA-derived spacers flanked by direct repeats (crRNAs). Upon maturation by Cas9, the tracrRNA and crRNA duplex guide the Cas9 nuclease to a target DNA sequence specified by the spacer guide sequences, and mediates double-stranded breaks in the DNA near a short sequence motif in the target DNA that is required for cleavage and specific to each CRISPR-Cas system. The type II CRISPR-Cas systems are found throughout the bacterial kingdom (FIGS. 7 and 8A-F) and highly diverse in Cas9 protein sequence and size, tracrRNA and crRNA direct repeat sequence, genome organization of these elements, and the motif requirement for target cleavage. One species may have multiple distinct CRISPR-Cas systems.
Applicants evaluated 207 putative Cas9s from bacterial species (FIG. 8A-F) identified based on sequence homology to known Cas9s and structures orthologous to known subdomains. Using the method of Example 1, Applicants will carry out a comprehensive evaluation of every possible mismatch in each position of the guide RNA for these different Cas9s to generate a model to inform the design of guide RNAs having high cleavage specificity for each based on the impact of the test position and number of mismatches in the guide RNA on cleavage efficiency for each Cas9.
The CRISPR-Cas system is amenable for achieving tissue-specific and temporally controlled targeted deletion of candidate disease genes. Examples include but are not limited to genes involved in cholesterol and fatty acid metabolism, amyloid diseases, dominant negative diseases, latent viral infections, among other disorders. Accordingly, target sequences can be in candidate disease genes, e.g.:
TABLE-US-00004 Disease GENE SPACER PAM Mechanism References Hypercholes- HMG- GCCAAATTG CGG Knockout Fluvastatin: a review of its terolemia CR GACGACCCT pharmacology and use in the CG management of hypercholesterolaemia. (Plosker GL et al. Drugs 1996, 51(3):433- 459) Hypercholes- SQLE CGAGGAGAC TGG Knockout Potential role of nonstatin terolemia CCCCGTTTC cholesterol lowering agents GG (Trapani et al. IUBMB Life, Volume 63, Issue 11, pages 964- 971, November 2011) Hyper- DGAT CCCGCCGCC AGG Knockout DGAT1 inhibitors as anti-obesity lipidemia 1 GCCGTGGCT and anti-diabetic agents. (Birch CG AM et al. Current Opinion in Drug Discovery & Development [2010, 13(4):489-496) Leukemia BCR- TGAGCTCTA AGG Knockout Killing of leukemic cells with a ABL CGAGATCCA BCR/ABL fusion gene by RNA CA interference (RNAi). (Fuchs et al. Oncogene 2002, 21(37):5716- 5724)
Examples of a pair of guide-RNA to introduce chromosomal microdeletion at a gene locus
TABLE-US-00005 Disease GENE SPACER PAM Mechanism References Hyperlip- PLIN2 CTCAAAATT TGG Micro- Perilipin-2 Null Mice are idemia guide1 CATACCGGT deletion Protected Against Diet-Induced TG Obesity, Adipose Inflammation and Fatty Liver Disease (McManaman JL et al. The Journal of Lipid Research, jlr.M035063. First Published on February 12, 2013) Hyperlip- PLIN2 CGTTAAACA TGG Micro- idemia guide2 ACAACCGGA deletion CT Hyperlip- SREBP TTCACCCCG ggg Micro- Inhibition of SREBP by a Small idemia guide1 CGGCGCTGA deletion Molecule, Betulin, Improves AT Hyperlipidemia and Insulin Resistance and Reduces Atherosclerotic Plaques (Tang J et al. Cell Metabolism, Volume 13, Issue 1, 44-56, 5 January 2011) Hyperlip- SREBP ACCACTACC agg Micro- idemia guide2 AGTCCGTCC deletion AC Examples of potential HIV-1 targeted spacers adapted from Mcintyre et al, which generated shRNAs against HIV-1 optimized for maximal coverage of HIV-1 variants. CACTGCTTAAGCCTCGCTCGAGG TCACCAGCAATATTCGCTCGAGG CACCAGCAATATTCCGCTCGAGG TAGCAACAGACATACGCTCGAGG GGGCAGTAGTAATACGCTCGAGG CCAATTCCCATACATTATTGTAC
Identification of Cas9 target site: Applicants analyzed the human CFTR genomic locus and identified the Cas9 target site (PAM may contain a NGG or a NNAGAAW motif). The frequency of these PAM sequences in the human genome are shown in FIG. 5.
Protospacer IDs and their corresponding genomic target, protospacer sequence, PAM sequence, and strand location are provided in the below Table. Guide sequences were designed to be complementary to the entire protospacer sequence in the case of separate transcripts in the hybrid system, or only to the underlined portion in the case of chimeric RNAs.
TABLE-US-00006 TABLE Protospacer IDs and their corresponding genomic target, protospacer sequence, PAM sequence, and strand location proto- protospacer spacer genomic sequence ID target (5' to 3') PAM strand 1 EMX1 GGACATCGATGTCAC TGG + CTCCAATGACTAGGG 2 EMX1 CATTGGAGGTGACAT TGG - CGATGTCCTCCCCAT 3 EMX1 GGAAGGGCCTGAGTC GGG + CGAGCAGAAGAAGAA 4 PVALB GGTGGCGAGAGGGGC AGG + CGAGATTGGGTGTTC 5 PVALB ATGCAGGAGGGTGGC TGG + GAGAGGGGCCGAGAT
Computational Identification of Unique CRISPR Target Sites:
To identify unique target sites for a Cas, e.g., a Cas9, e.g., the S. pyogenes SF370 Cas9 (SpCas9) enzyme, in nucleic acid molecules, e.g., of cells, e.g., of organisms, which include but are not limited to human, mouse, rat, zebrafish, fruit fly, and C. elegans genome, Applicants developed a software package to scan both strands of a DNA sequence and identify all possible SpCas9 target sites. For this example, each SpCas9 target site was operationally defined as a 20 bp sequence followed by an NGG protospacer adjacent motif (PAM) sequence, and all sequences satisfying this 5'-N.sub.20-NGG-3' definition on all chromosomes were identified. To prevent non-specific genome editing, after identifying all potential sites, all target sites were filtered based on the number of times they appear in the relevant reference genome. To take advantage of sequence specificity of Cas, e.g., Cas9 activity conferred by a `seed` sequence, which can be, for example, approximately 11-12 bp sequence 5' from the PAM sequence, 5'-NNNNNNNNNN-NGG-3' sequences were selected to be unique in the relevant genome. Genomic sequences are available on the UCSC Genome Browser and sample visualizations of the information for the Human genome hg, Mouse genome mm, Rat genome rn, Zebrafish genome danRer, D. melanogaster genome dm, C. elegans genome ce, the pig genome and cow genome are shown in FIGS. 15 through 22 respectively.
A similar analysis may be carried out for other Cas enzymes utilizing their respective PAM sequences, for e.g. Staphylococcus aureus sp. Aureus Cas9 and its PAM sequence NNGRR (FIG. 31).
Example 4
Experimental Architecture for Evaluating CRISPR-Cas Target Activity and Specificity
Targeted nucleases such as the CRISPR-Cas systems for gene editing applications allow for highly precise modification of the genome. However, the specificity of gene editing tools is a crucial consideration for avoiding adverse off-target activity. Here, Applicants describe a Cas9 guide RNA selection algorithm that predicts off-target sites for any desired target site within mammalian genomes.
Applicants constructed large oligo libraries of guide RNAs carrying combinations of mutations to study the sequence dependence of Cas9 programming. Using next-generation deep sequencing, Applicants studied the ability of single mutations and multiple combinations of mismatches within different Cas9 guide RNAs to mediate target DNA locus modification. Applicants evaluated candidate off-target sites with sequence homology to the target site of interest to assess any off-target cleavage.
Algorithm for Predicting CRISPR-Cas Target Activity and Specificity:
Data from these studies were used to develop algorithms for the prediction of CRISPR-Cas off-target activity across the human genome. The Applicants' resulting computational platform supports the prediction of all CRISPR-Cas system target activity and specificity in any genome. Applicants evaluate CRISPR-Cas activity and specificity by predicting the Cas9 cutting efficiency for any CRISPR-Cas target against all other genomic CRISPR-Cas targets, excluding constraining factors, i.e., some epigenetic modifications like repressive chromatin/heterochromatin.
The algorithms Applicants describe 1) evaluate any target site and give potential off-targets and 2) generate candidate target sites for any locus of interest with minimal predicted off-target activity.
Least Squares Thermodynamic Model of CRISPR-Cas Cutting Efficiency:
For arbitrary Cas9 target sites, Applicants generated a numerical thermodynamic model that predicts Cas9 cutting efficiency. Applicants propose 1) that the Cas9 guide RNA has specific free energies of hybridization to its target and any off-target DNA sequences and 2) that Cas9 modifies RNA:DNA hybridization free-energies locally in a position-dependent but sequence-independent way. Applicants trained a model for predicting CRISPR-Cas cutting efficiency based on their CRISPR-Cas guide RNA mutation data and RNA:DNA thermodynamic free energy calculations using a machine learning algorithm. Applicants then validated their resulting models by comparing their predictions of CRISPR-Cas off-target cutting at multiple genomic loci with experimental data assessing locus modification at the same sites.
The methodology adopted in developing this algorithm is as follows: The problem summary states that for arbitrary spacers and targets of constant length, a numerical model that makes thermodynamic sense and predicts Cas9 cutting efficiency is to be found.
Suppose Cas9 modifies DNA:RNA hybridization free-energies locally in a position-dependent but sequence-independent way. Then for DNA:RNA hybridization free energies .DELTA.G.sub.ij(k) (for position k between 1 and N) of spacer i and target j
.times..times..alpha..times..DELTA..times..times..function. ##EQU00004## Z.sub.ij can be treated as an "effective" free-energy modified by the multiplicative position-weights .alpha..sub.k.
The "effective" free-energy Z.sub.ij corresponds to an associated cutting-probability e.sup.-.beta.Z.sup.ij (for some constant .beta.) in the same way that an equilibrium model of hybridization (without position-weighting) would have predicted a hybridization-probability .about.e.sup.-.beta..DELTA.G.sup.ij. Since cutting-efficiency has been measured, the values Z.sub.ij can be treated as their observables. Meanwhile, .DELTA.G.sub.ij(k) can be calculated for any experiment's spacer-target pairing. Applicants task was to find the values .alpha..sub.k, since this would allow them to estimate Z.sub.ij for any spacer-target pair.
Writing the above equation for Z.sub.ij in matrix form Applicants get: {right arrow over (Z)}=G{right arrow over (.alpha.)} (1) The least-squares estimate is then {right arrow over (.alpha.)}.sub.est=(G.sup.TG).sup.-1G.sup.T{right arrow over (Z)} where G.sup.T is the matrix-transpose of the G and (G.sup.TG).sup.-1 is the inverse of their matrix-product.
In the above G is a matrix of local DNA:RNA free-energy values whose rth row corresponds to experimental trial r and whose kth column corresponds to the kth position in the DNA:RNA hybrid tested in that experimental trial. {right arrow over (Z)} is meanwhile a column-vector whose rth row corresponds to observables from the same experimental trial as G's rth row. Because of the relation described above wherein the CRISPR cutting frequencies are estimated to vary as e.sup.-.beta.Z.sup.ij, these observables, Z.sub.ij, were calculated as the natural logarithm of the observed cutting frequency. The observable is the cleavage efficiency of Cas, e.g., Cas9, at a target DNA for a particular guide RNA and target DNA pair. The experiment is Cas, e.g., Cas9, with a particular sgRNA/DNA target pairing, and the observable is the cleavage percentage (whether measured as indel formation percentage from cells or simply cleavage percentage in vitro) (see herein discussion on generating training data set). More in particular, every unique PCR reaction that was sequenced should be treated as a unique experimental trial to encompass replicability within the vector. This means that experimental replicates each go into separate rows of equation 1 (and because of this, some rows of G will be identical). The advantage of this is that when a is fit, all relevant information--including replicability--is taken into account in the final estimate.
Observable {right arrow over (Z)}, values were calculated as log (observed frequency of cutting). Cutting frequencies were normalized identically (so that they all have the same "units"). For plugging in sequencing indel-frequency values, it may be best, however, to standardize sequencing depth.
The preferred way to do this would be to set a standard sequencing-depth D for which all experiments included in {right arrow over (Z)} have at least that number of reads. Since cutting frequencies below 1/D cannot be consistently detected, this should be set as the minimum frequency for the data-set, and the values in {right arrow over (Z)} should range from log(1/D) to log(1). One could vary the value of D later on to ensure that the {right arrow over (.alpha.)} estimate isn't too dependent on the value chosen.
In a further aspect, there are different methods of graphing NGG and NNAGAAW sequences. One is with the `non-overlapping` method. NGG and NRG may be regraphed in an "overlapping" fashion, as indicated in FIGS. 6 A-C.
Applicants also performed a study on off target Cas9 activity as indicated in FIGS. 10, 11 and 12. Aspects of the invention also relate to predictive models that may not involve hybridization energies but instead simply use the cutting frequency information as a prediction (See FIG. 29).
Example 5
DNA Targeting Specificity of the RNA-Guided Cas9 Nuclease
Here, Applicants report optimization of various applications of SpCas9 for mammalian genome editing and demonstrate that SpCas9-mediated cleavage is unaffected by DNA methylation (FIG. 14). Applicants further characterize SpCas9 targeting specificity using over 700 guide RNA variants and evaluate SpCas9-induced indel mutation levels at over 100 predicted genomic off-target loci. Contrary to previous models, Applicants found that SpCas9 tolerates mismatches between guide RNA and target DNA at different positions in a sequence-context dependent manner, sensitive to the number, position and distribution of mismatches. Finally, Applicants demonstrate that the dosage of SpCas9 and sgRNA can be titrated to minimize off-target modification. To facilitate mammalian genome engineering applications, Applicants used these results to establish a computational platform to guide the selection and validation of target sequences as well as off-target analyses.
The bacterial type II CRISPR system from S. pyogenes may be reconstituted in mammalian cells using three minimal components: the Cas9 nuclease (SpCas9), a specificity-determining CRISPR RNA (crRNA), and an auxiliary trans-activating crRNA (tracrRNA). Following crRNA and tracrRNA hybridization, SpCas9 is localized to the genomic target matching a 20-nt guide sequence within the crRNA, immediately upstream of a required 5'-NGG protospacer adjacent motif (PAM). Each crRNA and tracrRNA duplex may also be fused to generate a chimeric single guide RNA (sgRNA) that mimics the natural crRNA-tracrRNA hybrid. Both crRNA-tracrRNA duplexes and sgRNAs can be used to target SpCas9 for multiplexed genome editing in eukaryotic cells.
Although an sgRNA design consisting of a truncated crRNA and tracrRNA had been previously shown to mediate efficient cleavage in vitro, it failed to achieve detectable cleavage at several loci that were efficiently modified by crRNA-tracrRNA duplexes bearing identical guide sequences. Because the major difference between this sgRNA design and the native crRNA-tracrRNA duplex is the length of the tracrRNA sequence, Applicants tested whether extension of the tracrRNA tail was able to improve SpCas9 activity.
Applicants generated a set of sgRNAs targeting multiple sites within the human EMX1 and PVALB loci with different tracrRNA 3' truncations. Using the SURVEYOR nuclease assay, Applicants assessed the ability of each Cas9 sgRNA complex to generate indels in HEK 293FT cells through the induction of DNA double-stranded breaks (DSBs) and subsequent non-homologous end joining (NHEJ) DNA damage repair (Methods and Materials). sgRNAs with +67 or +85 nucleotide (nt) tracrRNA tails mediated DNA cleavage at all target sites tested, with up to 5-fold higher levels of indels than the corresponding crRNA-tracrRNA duplexes. Furthermore, both sgRNA designs efficiently modified PVALB loci that were previously not targetable using crRNA-tracrRNA duplexes. For all five tested targets, Applicants observed a consistent increase in modification efficiency with increasing tracrRNA length. Applicants performed Northern blots for the guide RNA truncations and found increased levels expression for the longer tracrRNA sequences, suggesting that improved target cleavage was due to higher sgRNA expression or stability. Taken together, these data indicate that the tracrRNA tail is important for optimal SpCas9 expression and activity in vivo.
Applicants further investigated the sgRNA architecture by extending the duplex length from 12 to the 22 nt found in the native crRNA-tracrRNA duplex. Applicants also mutated the sequence encoding sgRNA to abolish any poly-T tracts that could serve as premature transcriptional terminators for U6-driven transcription. Applicants tested these new sgRNA scaffolds on 3 targets within the human EMX1 gene and observed only modest changes in modification efficiency. Thus, Applicants established sgRNA(+85), identical to some sgRNAs previously used, as an effective SpCas9 guide RNA architecture and used it in all subsequent studies.
Applicants have previously shown that a catalytic mutant of SpCas9 (D10A nickase) can mediate gene editing by homology-directed repair (HR) without detectable indel formation. Given its higher cleavage efficiency, Applicants tested whether sgRNA(+85), in complex with the Cas9 nickase, can likewise facilitate HR without incurring on-target NHEJ. Using single-stranded oligonucleotides (ssODNs) as repair templates, Applicants observed that both the wild-type and the D10A SpCas9 mediate HR in HEK 293FT cells, while only the former is able to do so in human embryonic stem cells. Applicants further confirmed using SURVEYOR assay that no target indel mutations are induced by the SpCas9 D10A nickase.
To explore whether the genome targeting ability of sgRNA(+85) is influenced by epigenetic factors that constrain the alternative transcription activator-like effector nuclease (TALENs) and potentially also zinc finger nuclease (ZFNs) technologies, Applicants further tested the ability of SpCas9 to cleave methylated DNA. Using either unmethylated or M. SssI-methylated pUC19 as DNA targets (FIG. 14a,b) in a cell-free cleavage assay, Applicants showed that SpCas9 efficiently cleaves pUC19 regardless of CpG methylation status in either the 20-bp target sequence or the PAM (FIG. 14c). To test whether this is also true in vivo, Applicants designed sgRNAs to target a highly methylated region of the human SERPINB5 locus. All three sgRNAs tested were able to mediate indel mutations in endogenously methylated targets.
Having established the optimal guide RNA architecture for SpCas9 and demonstrated its insensitivity to genomic CpG methylation, Applicants sought to conduct a comprehensive characterization of the DNA targeting specificity of SpCas9. Previous studies on SpCas9 cleavage specificity were limited to a small set of single-nucleotide mismatches between the guide sequence and DNA target, suggesting that perfect base-pairing within 10-12 bp directly 5' of PAM determines Cas9 specificity, whereas PAM-distal multiple mismatches can be tolerated. In addition, a recent study using catalytically inactive SpCas9 as a transcriptional repressor found no significant off-target effects throughout the E. coli transcriptome. However, a systematic analysis of Cas9 specificity within the context of a larger mammalian genome has not yet been reported.
To address this, Applicants first evaluated the effect of imperfect guide RNA identity for targeting genomic DNA on SpCas9 activity, and then assessed the cleavage activity resulting from a single sgRNA on multiple genomic off-target loci with sequence similarity. To facilitate large scale testing of mismatched guide sequences, Applicants developed a simple sgRNA testing assay by generating expression cassettes encoding U6-driven sgRNAs by PCR and transfecting the resulting amplicons. Applicants then performed deep sequencing of the region flanking each target site for two independent biological replicates. From these data, Applicants applied a binomial model to detect true indel events resulting from SpCas9 cleavage and NHEJ misrepair and calculated 95% confidence intervals for all reported NHEJ frequencies.
Applicants used a linear model of free energy position-dependence to investigate the combined contribution of DNA:RNA sequence and mismatch-location on Cas9 cutting efficiency. While sequence composition and mismatch location alone generated Spearman correlations between estimated and observed cutting efficiencies for EMX1 target site 1 and 0.78, respectively, integration of the two parameters greatly improved this agreement, with Spearman correlation 0.86 (p<0.001). Furthermore, the incorporation of nupac RNA:RNA hybridization energies into Applicants' free energy model resulted in a 10% increase in the Spearman correlation coefficient. Taken together, the data suggests an effect of SpCas9-specific perturbations on the Watson-Crick base-pairing free energies. Meanwhile, sequence composition did not substantially improve agreement between estimated and observed cutting efficiencies for EMX1 target site 6 (Spearman correlation 0.91, p<0.001). This suggested that single mismatches in EMX1 target site 6 contributed minimally to the thermodynamic binding free energy itself.
Potential genomic off-target sites with sequence similarity to a target site of interest may often have multiple base mismatches. Applicants designed a set of guide RNAs for EMX1 targets 1 and 6 that contains different combinations of mismatches to investigate the effect of mismatch number, position, and spacing on Cas9 target cleavage activity (FIG. 13a,b).
By concatenating blocks of mismatches, Applicants found that two consecutive mismatches within the PAM-proximal sequence reduced Cas9 cutting for both targets to <1% (FIG. 13a; top panels). Target site 1 cutting increased as the double mismatches shifted distally from the PAM, whereas observed cleavage for target site 6 consistently remained <0.5%. Blocks of three or five consecutive mismatches for both targets diminished Cas9 cutting to levels <0.5% regardless of position (FIG. 13, lower panels).
To investigate the effect of mismatch spacing, Applicants anchored a single PAM-proximal mutation while systematically increasing the separation between subsequent mismatches. Groups of 3 or 4 mutations each separated by 3 or fewer bases diminished Cas9 nuclease activity to levels <0.5%. However, Cas9 cutting at target site 1 increased to 3-4% when the mutations were separated by 4 or more unmutated bases (FIG. 13b). Similarly, groups of 4 mutations separated by 4 or more bases led to indel efficiencies from 0.5-1%. However, cleavage at target site 6 consistently remained below 0.5% regardless of the number or spacing of the guide RNA mismatches.
The multiple guide RNA mismatch data indicate that increasing the number of mutations diminishes and eventually abolishes cleavage. Unexpectedly, isolated mutations are tolerated as separation increased between each mismatch. Consistent with the single mismatch data, multiple mutations within the PAM-distal region are generally tolerated by Cas9 while clusters of PAM-proximal mutations are not. Finally, although the mismatch combinations represent a limited subset of base mutations, there appears to be target-specific susceptibility to guide RNA mismatches. For example, target site 6 generally showed lower cleavage with multiple mismatches, a property also reflected in its longer 12-14 bp PAM-proximal region of mutation intolerance (FIG. 12). Further investigation of Cas9 sequence-specificity may reveal design guidelines for choosing more specific DNA targets.
To determine if Applicants' findings from the guide RNA mutation data generalize to target DNA mismatches and allow the prediction of off-target cleavage within the genome, Applicants transfected cells with Cas9 and guide RNAs targeting either target 3 or target 6, and performed deep sequencing of candidate off-target sites with sequence similarity. No genomic loci with only 1 mismatch to either targets was identified. Genomic loci containing 2 or 3 mismatches relative to target 3 or target 6 revealed cleavage at some of the off-targets assessed (FIG. 13c). Targets 3 and 6 exhibited cleavage efficiencies of 7.5% and 8.0%, whereas off-target sites 3-1, 3-2, 3-4, and 3-5 were modified at 0.19%, 0.42%, 0.97%, and 0.50%, respectively. All other off-target sites cleaved at under 0.1% or were modified at levels indistinguishable from sequencing error. The off-target cutting rates were consistent with the collective results from the guide RNA mutation data: cleavage was observed at a small subset of target 3 off-targets that contained either very PAM-distal mismatches or had single mismatches separated by 4 or more bases.
Given that the genome targeting efficiencies of TALENs and ZFNs may be sensitive to confounding effects such as chromatin state or DNA methylation, Applicants sought to test whether RNA-guided SpCas9 cleavage activity would be affected by the epigenetic state of a target locus. To test this, Applicants methylated a plasmid in vitro and performed an in vitro cleavage assay on two pairs of targets containing either unmethylated or methylated CpGs. SpCas9 mediated efficient cleavage of the plasmid whether methylation occurred in the target proper or within the PAM, suggesting that SpCas9 may not be susceptible to DNA methylation effects.
The ability to program Cas9 to target specific sites in the genome by simply designing a short sgRNA has enormous potential for a variety of applications. Applicants' results demonstrate that the specificity of Cas9-mediated DNA cleavage is sequence-dependent and is governed not only by the location of mismatching bases, but also by their spacing. Importantly, while the PAM-proximal 9-12 nt of the guide sequence generally defines specificity, the PAM-distal sequences also contribute to the overall specificity of Cas9-mediated DNA cleavage. Although there are off-target cleavage sites for a given guide sequence, expected off-target sites are likely predictable based on their mismatch locations. Further work looking at the thermodynamics of sgRNA-DNA interaction will likely yield additional predictive power for off-target activity, and exploration of alternative Cas9 orthologs may also yield novel variants of Cas9s with improved specificity. Taken together, the high efficiency of Cas9 as well as its low off-target activity make CRISPR-Cas an attractive genome engineering technology.
Example 6
Use of Cas9 to Target a Variety of Disease Types
The specificity of Cas9 orthologs can be evaluated by testing the ability of each Cas9 to tolerate mismatches between the guide RNA and its DNA target. For example, the specificity of SpCas9 has been characterized by testing the effect of mutations in the guide RNA on cleavage efficiency. Libraries of guide RNAs were made with single or multiple mismatches between the guide sequence and the target DNA. Based on these findings, target sites for SpCas9 can be selected based on the following guidelines:
To maximize SpCas9 specificity for editing a particular gene, one should choose a target site within the locus of interest such that potential `off-target` genomic sequences abide by the following four constraints: First and foremost, they should not be followed by a PAM with either 5'-NGG or NAG sequences. Second, their global sequence similarity to the target sequence should be minimized. Third, a maximal number of mismatches should lie within the PAM-proximal region of the off-target site. Finally, a maximal number of mismatches should be consecutive or spaced less than four bases apart.
Similar methods can be used to evaluate the specificity of other Cas9 orthologs and to establish criteria for the selection of specific target sites within the genomes of target species.
Target selection for sgRNA: There are two main considerations in the selection of the 20-nt guide sequence for gene targeting: 1) the target sequence should precede the 5'-NGG PAM for S. pyogenes Cas9, and 2) guide sequences should be chosen to minimize off-target activity. Applicants provided an online Cas9 targeting design tool (available at the website genome-engineering.org/tools; see Examples above and FIG. 23) that takes an input sequence of interest and identifies suitable target sites. To experimentally assess off-target modifications for each sgRNA, Applicants also provide computationally predicted off-target sites for each intended target, ranked according to Applicants" quantitative specificity analysis on the effects of base-pairing mismatch identity, position, and distribution.
The detailed information on computationally predicted off-target sites is as follows: Considerations for Off-target Cleavage Activities: Similar to other nucleases, Cas9 can cleave off-target DNA targets in the genome at reduced frequencies. The extent to which a given guide sequence exhibit off-target activity depends on a combination of factors including enzyme concentration, thermodynamics of the specific guide sequence employed, and the abundance of similar sequences in the target genome. For routine application of Cas9, it is important to consider ways to minimize the degree of off-target cleavage and also to be able to detect the presence of off-target cleavage.
Minimizing off-target activity: For application in cell lines, Applicants recommend following two steps to reduce the degree of off-target genome modification. First, using Applicants' online CRISPR target selection tool, it is possible to computationally assess the likelihood of a given guide sequence to have off-target sites. These analyses are performed through an exhaustive search in the genome for off-target sequences that are similar sequences as the guide sequence. Comprehensive experimental investigation of the effect of mismatching bases between the sgRNA and its target DNA revealed that mismatch tolerance is 1) position dependent--the 8-14 bp on the 3' end of the guide sequence are less tolerant of mismatches than the 5' bases, 2) quantity dependent--in general more than 3 mismatches are not tolerated, 3) guide sequence dependent--some guide sequences are less tolerant of mismatches than others, and 4) concentration dependent--off-target cleavage is highly sensitive to the amount of transfected DNA. The Applicants' target site analysis web tool (available at the website genome-engineering.org/tools) integrates these criteria to provide predictions for likely off-target sites in the target genome. Second, Applicants recommend titrating the amount of Cas9 and sgRNA expression plasmid to minimize off-target activity.
Detection of off-target activities: Using Applicants' CRISPR targeting web tool, it is possible to generate a list of most likely off-target sites as well as primers performing SURVEYOR or sequencing analysis of those sites. For isogenic clones generated using Cas9, Applicants strongly recommend sequencing these candidate off-target sites to check for any undesired mutations. It is worth noting that there may be off target modifications in sites that are not included in the predicted candidate list and full genome sequence should be performed to completely verify the absence of off-target sites. Furthermore, in multiplex assays where several DSBs are induced within the same genome, there may be low rates of translocation events and can be evaluated using a variety of techniques such as deep sequencing (48).
The online tool (FIG. 23) provides the sequences for all oligos and primers necessary for 1) preparing the sgRNA constructs, 2) assaying target modification efficiency, and 3) assessing cleavage at potential off-target sites. It is worth noting that because the U6 RNA polymerase III promoter used to express the sgRNA prefers a guanine (G) nucleotide as the first base of its transcript, an extra G is appended at the 5' of the sgRNA where the 20-nt guide sequence does not begin with G (FIG. 24).
Example 7
Base Pair Mismatching Investigations
Applicants tested whether extension of the tracrRNA tail was able to improve SpCas9 activity. Applicants generated a set of sgRNAs targeting multiple sites within the human EMX1 and PVALB loci with different tracrRNA 3' truncations (FIG. 9a). Using the SURVEYOR nuclease assay, Applicants assessed the ability of each Cas9 sgRNA complex to generate indels in HEK 293FT cells through the induction of DNA double-stranded breaks (DSBs) and subsequent non-homologous end joining (NHEJ) DNA damage repair (Methods and Materials). sgRNAs with +67 or +85 nucleotide (nt) tracrRNA tails mediated DNA cleavage at all target sites tested, with up to 5-fold higher levels of indels than the corresponding crRNA-tracrRNA duplexes (FIG. 9). Furthermore, both sgRNA designs efficiently modified PVALB loci that were previously not targetable using crRNA-tracrRNA duplexes (1) (FIG. 9b and FIG. 9b). For all five tested targets, Applicants observed a consistent increase in modification efficiency with increasing tracrRNA length. Applicants performed Northern blots for the guide RNA truncations and found increased levels expression for the longer tracrRNA sequences, suggesting that improved target cleavage was due to higher sgRNA expression or stability (FIG. 9c). Taken together, these data indicate that the tracrRNA tail is important for optimal SpCas9 expression and activity in vivo.
Applicants have previously shown that a catalytic mutant of SpCas9 (D10A nickase) can mediate gene editing by homology-directed repair (HR) without detectable indel formation. Given its higher cleavage efficiency, Applicants tested whether sgRNA(+85), in complex with the Cas9 nickase, can likewise facilitate HR without incurring on-target NHEJ. Using single-stranded oligonucleotides (ssODNs) as repair templates, Applicants observed that both the wild-type and the D10A SpCas9 mediate HR in HEK 293FT cells, while only the former is able to do so in human embryonic stem cells (hESCs; FIG. 9d).
To explore whether the genome targeting ability of sgRNA(+85) is influenced by epigenetic factors that constrain the alternative transcription activator-like effector nuclease (TALENs) and potentially also zinc finger nuclease (ZFNs) technologies, Applicants further tested the ability of SpCas9 to cleave methylated DNA. Using either unmethylated or M. SssI-methylated pUC19 as DNA targets (FIG. 14a,b) in a cell-free cleavage assay, Applicants showed that SpCas9 efficiently cleaves pUC19 regardless of CpG methylation status in either the 20-bp target sequence or the PAM. To test whether this is also true in vivo, Applicants designed sgRNAs to target a highly methylated region of the human SERPINB5 locus (FIG. 9e,f). All three sgRNAs tested were able to mediate indel mutations in endogenously methylated targets (FIG. 9g).
Applicants systematically investigated the effect of base-pairing mismatches between guide RNA sequences and target DNA on target modification efficiency. Applicants chose four target sites within the human EMX1 gene and, for each, generated a set of 57 different guide RNAs containing all possible single nucleotide substitutions in positions 1-19 directly 5' of the requisite NGG PAM (FIG. 25a). The 5' guanine at position 20 is preserved, given that the U6 promoter requires guanine as the first base of its transcript. These `off-target` guide RNAs were then assessed for cleavage activity at the on-target genomic locus.
Consistent with previous findings, SpCas9 tolerates single base mismatches in the PAM-distal region to a greater extent than in the PAM-proximal region. In contrast with a model that implies a prototypical 10-12 bp PAM-proximal seed sequence that determines target specificity, Applicants found that most bases within the target site are specifically recognized, although mismatches are tolerated at different positions in a sequence-context dependent manner. Single-base specificity generally ranges from 8 to 12 bp immediately upstream of the PAM, indicating a sequence-dependent specificity boundary that varies in length (FIG. 25b).
To further investigate the contributions of base identity and position within the guide RNA to SpCas9 specificity, Applicants generated additional sets of mismatched guide RNAs for eleven more target sites within the EMX1 locus (FIG. 28) totaling over 400 sgRNAs. These guide RNAs were designed to cover all 12 possible RNA:DNA mismatches for each position in the guide sequence with at least 2.times. coverage for positions 1-10. Applicants' aggregate single mismatch data reveals multiple exceptions to the seed sequence model of SpCas9 specificity (FIG. 25c). In general, mismatches within the 8-12 PAM-proximal bases were less tolerated by SpCas9, whereas those in the PAM-distal regions had little effect on SpCas9 cleavage. Within the PAM-proximal region, the degree of tolerance varied with the identity of a particular mismatch, with rC:dC base-pairing exhibiting the highest level of disruption to SpCas9 cleavage (FIG. 25c).
In addition to the target specificity, Applicants also investigated the NGG PAM requirement of SpCas9. To vary the second and third positions of PAM, Applicants selected 32 target sites within the EMX1 locus encompassing all 16 possible alternate PAMs with 2.times. coverage (Table 4). Using SURVEYOR assay, Applicants showed that SpCas9 also cleaves targets with NAG PAMs, albeit 5-fold less efficiently than target sites with NGG PAMs (FIG. 25d). The tolerance for an NAG PAM is in agreement with previous bacterial studies (12) and expands the S. pyogenes Cas9 target space to every 4-bp on average within the human genome, not accounting for constraining factors such as guide RNA secondary structure or certain epigenetic modifications (FIG. 25e).
Applicants next explored the effect of multiple base mismatches on SpCas9 target activity. For four targets within the EMX1 gene, Applicants designed sets of guide RNAs that contained varying combinations of mismatches to investigate the effect of mismatch number, position, and spacing on SpCas9 target cleavage activity (FIG. 26a, b).
In general, Applicants observed that the total number of mismatched base-pairs is a key determinant for SpCas9 cleavage efficiency. Two mismatches, particularly those occurring in a PAM-proximal region, significantly reduced SpCas9 activity whether these mismatches are concatenated or interspaced (FIG. 26a, b); this effect is further magnified for three concatenated mismatches (FIG. 20a). Furthermore, three or more interspaced (FIG. 26c) and five concatenated (FIG. 26a) mismatches eliminated detectable SpCas9 cleavage in the vast majority of loci.
The position of mismatches within the guide sequence also affected the activity of SpCas9: PAM-proximal mismatches are less tolerated than PAM-distal counterparts (FIG. 26a), recapitulating Applicants' observations from the single base-pair mismatch data (FIG. 25c). This effect is particularly salient in guide sequences bearing a small number of total mismatches, whether those are concatenated (FIG. 26a) or interspaced (FIG. 26b). Additionally, guide sequences with mismatches spaced four or more bases apart also mediated SpCas9 cleavage in some cases (FIG. 26c). Thus, together with the identity of mismatched base-pairing, Applicants observed that many off-target cleavage effects can be explained by a combination of mismatch number and position.
Given these mismatched guide RNA results, Applicants expected that for any particular sgRNA, SpCas9 may cleave genomic loci that contain small numbers of mismatched bases. For the four EMX1 targets described above, Applicants computationally identified 117 candidate off-target sites in the human genome that are followed by a 5'-NRG PAM and meet any of the additional following criteria: 1. up to 5 mismatches, 2. short insertions or deletions, or 3. mismatches only in the PAM-distal region. Additionally, Applicants assessed off-target loci of high sequence similarity without the PAM requirement. The majority of off-target sites tested for each sgRNA (30/31, 23/23, 48/51, and 12/12 sites for EMX1 targets 1, 2, 3, and 6, respectively) exhibited modification efficiencies at least 100-fold lower than that of corresponding on-targets (FIG. 27a, b). Of the four off-target sites identified, three contained only mismatches in the PAM-distal region, consistent with the Applicants' multiple mismatch sgRNA observations (FIG. 26). Notably, these three loci were followed by 5'-NAG PAMs, demonstrating that off-target analyses of SpCas9 must include 5'-NAG as well as 5'-NGG candidate loci.
Enzymatic specificity and activity strength are often highly dependent on reaction conditions, which at high reaction concentration might amplify off-target activity (26, 27). One potential strategy for minimizing non-specific cleavage is to limit the enzyme concentration, namely the level of SpCas9-sgRNA complex. Cleavage specificity, measured as a ratio of on- to off-target cleavage, increased dramatically as Applicants decreased the equimolar amounts of SpCas9 and sgRNA transfected into 293FT cells (FIG. 27c, d) from 7.1.times.10-10 to 1.8.times.10-11 nmoL/cell (400 ng to 10 ng of Cas9-sgRNA plasmid). qRT-PCR assay confirmed that the level of hSpCas9 mRNA and sgRNA decreased proportionally to the amount of transfected DNA. Whereas specificity increased gradually by nearly 4-fold as Applicants decreased the transfected DNA amount from 7.1.times.10-10 to 9.0.times.10-11 nmol/cell (400 ng to 50 ng plasmid), Applicants observed a notable additional 7-fold increase in specificity upon decreasing transfected DNA from 9.0.times.10-11 to 1.8.times.10-11 nmol/cell (50 ng to 10 ng plasmid; FIG. 27c). These findings suggest that Applicants may minimize the level of off-target activity by titrating the amount of SpCas9 and sgRNA DNA delivered. However, increasing specificity by reducing the amount of transfected DNA also leads to a reduction in on-target cleavage. These measurements enable quantitative integration of specificity and efficiency criteria into dosage choice to optimize SpCas9 activity for different applications. Applicants further explore modifications in SpCas9 and sgRNA design that may improve the intrinsic specificity without sacrificing cleavage efficiency. FIG. 29 shows data for EMX1 target 2 and target 6. For the tested sites in FIGS. 27 and 29 (in this case, sites with 3 mismatches or less), there were no off-target sites identified (defined as off-target site cleavage within 100-fold of the on-target site cleavage).
The ability to program SpCas9 to target specific sites in the genome by simply designing a short sgRNA holds enormous potential for a variety of applications. Applicants' results demonstrate that the specificity of SpCas9-mediated DNA cleavage is sequence- and locus-dependent and governed by the quantity, position, and identity of mismatching bases. Importantly, while the PAM-proximal 8-12 bp of the guide sequence generally defines specificity, the PAM-distal sequences also contribute to the overall specificity of SpCas9-mediated DNA cleavage. Although there may be off-target cleavage for a given guide sequence, they can be predicted and likely minimized by following general design guidelines.
To maximize SpCas9 specificity for editing a particular gene, one should identify potential `off-target` genomic sequences by considering the following four constraints: First and foremost, they should not be followed by a PAM with either 5'-NGG or 5'-NAG sequences. Second, their global sequence similarity to the target sequence should be minimized, and guide sequences with genomic off-target loci that have fewer than 3 mismatches should be avoided. Third, at least 2 mismatches should lie within the PAM-proximal region of the off-target site. Fourth, a maximal number of mismatches should be consecutive or spaced less than four bases apart. Finally, the amount of SpCas9 and sgRNA may be titrated to optimize on- to off-target cleavage ratio.
Using these criteria, Applicants formulated a simple scoring scheme to integrate the contributions of mismatch location, density, and identity for quantifying their contribution to SpCas9 cutting. Applicants applied the aggregate cleavage efficiencies of single-mismatch guide RNAs to test this scoring scheme separately on genome-wide targets. Applicants found that these factors, taken together, accounted for more than 50% of the variance in cutting-frequency rank among the genome-wide targets studied (FIG. 30).
Implementing the guidelines delineated above, Applicants designed a computational tool to facilitate the selection and validation of sgRNAs as well as to predict off-target loci for specificity analyses; this tool may be accessed at the website genome-engineering.org/tools. These results and tools further extend the SpCas9 system as a powerful and versatile alternative to ZFNs and TALENs for genome editing applications. Further work examining the thermodynamics and in vivo stability of sgRNA-DNA duplexes will likely yield additional predictive power for off-target activity, while exploration of SpCas9 mutants and orthologs may yield novel variants with improved specificity. Accession codes All raw reads can be accessed at NCBI BioProject, accession number SRP023129.
Methods and Materials:
Cell culture and transfection--Human embryonic kidney (HEK) cell line 293FT (Life Technologies) was maintained in Dulbecco's modified Eagle's Medium (DMEM) supplemented with 10% fetal bovine serum (HyClone), 2 mM GlutaMAX (Life Technologies), 100 U/mL penicillin, and 100 .mu.g/mL streptomycin at 37.degree. C. with 5% CO2 incubation.
293FT cells were seeded either onto 6-well plates, 24-well plates, or 96-well plates (Corning) 24 hours prior to transfection. Cells were transfected using Lipofectamine 2000 (Life Technologies) at 80-90% confluence following the manufacturer's recommended protocol. For each well of a 6-well plate, a total of 1 ug of Cas9+sgRNA plasmid was used. For each well of a 24-well plate, a total of 500 ng Cas9+sgRNA plasmid was used unless otherwise indicated. For each well of a 96-well plate, 65 ng of Cas9 plasmid was used at a 1:1 molar ratio to the U6-sgRNA PCR product.
Human embryonic stem cell line HUES9 (Harvard Stem Cell Institute core) was maintained in feeder-free conditions on GelTrex (Life Technologies) in mTesR medium (Stemcell Technologies) supplemented with 100 ug/ml Normocin (InvivoGen). HUES9 cells were transfected with Amaxa P3 Primary Cell 4-D Nucleofector Kit (Lonza) following the manufacturer's protocol.
SURVEYOR Nuclease Assay for Genome Modification
293FT cells were transfected with plasmid DNA as described above. Cells were incubated at 37.degree. C. for 72 hours post-transfection prior to genomic DNA extraction. Genomic DNA was extracted using the QuickExtract DNA Extraction Solution (Epicentre) following the manufacturer's protocol. Briefly, pelleted cells were resuspended in QuickExtract solution and incubated at 65.degree. C. for 15 minutes and 98.degree. C. for 10 minutes.
The genomic region flanking the CRISPR target site for each gene was PCR amplified (primers listed in Table 2), and products were purified using QiaQuick Spin Column (Qiagen) following the manufacturer's protocol. 400 ng total of the purified PCR products were mixed with 2 .mu.l 10.times.Taq DNA Polymerase PCR buffer (Enzymatics) and ultrapure water to a final volume of 20 .mu.l, and subjected to a re-annealing process to enable heteroduplex formation: 95.degree. C. for 10 min, 95.degree. C. to 85.degree. C. ramping at -2.degree. C./s, 85.degree. C. to 25.degree. C. at -0.25.degree. C./s, and 25.degree. C. hold for 1 minute. After re-annealing, products were treated with SURVEYOR nuclease and SURVEYOR enhancer S (Transgenomics) following the manufacturer's recommended protocol, and analyzed on 4-20% Novex TBE poly-acrylamide gels (Life Technologies). Gels were stained with SYBR Gold DNA stain (Life Technologies) for 30 minutes and imaged with a Gel Doc gel imaging system (Bio-rad). Quantification was based on relative band intensities.
Northern blot analysis of tracrRNA expression in human cells: Northern blots were performed as previously described 1. Briefly, RNAs were heated to 95.degree. C. for 5 min before loading on 8% denaturing polyacrylamide gels (SequaGel, National Diagnostics). Afterwards, RNA was transferred to a pre-hybridized Hybond N+ membrane (GE Healthcare) and crosslinked with Stratagene UV Crosslinker (Stratagene). Probes were labeled with [gamma-32P] ATP (Perkin Elmer) with T4 polynucleotide kinase (New England Biolabs). After washing, membrane was exposed to phosphor screen for one hour and scanned with phosphorimager (Typhoon).
Bisulfite sequencing to assess DNA methylation status: HEK 293FT cells were transfected with Cas9 as described above. Genomic DNA was isolated with the DNeasy Blood & Tissue Kit (Qiagen) and bisulfite converted with EZ DNA Methylation-Lightning Kit (Zymo Research). Bisulfite PCR was conducted using KAPA2G Robust HotStart DNA Polymerase (KAPA Biosystems) with primers designed using the Bisulfite Primer Seeker (Zymo Research, Table 6). Resulting PCR amplicons were gel-purified, digested with EcoRI and HindIII, and ligated into a pUC19 backbone prior to transformation. Individual clones were then Sanger sequenced to assess DNA methylation status.
In vitro transcription and cleavage assay: HEK 293FT cells were transfected with Cas9 as described above. Whole cell lysates were then prepared with a lysis buffer (20 mM HEPES, 100 mM KCl, 5 mM MgCl2, 1 mM DTT, 5% glycerol, 0.1% Triton X-100) supplemented with Protease Inhibitor Cocktail (Roche). T7-driven sgRNA was in vitro transcribed using custom oligos (Sequences) and HiScribe T7 In Vitro Transcription Kit (NEB), following the manufacturer's recommended protocol. To prepare methylated target sites, pUC19 plasmid was methylated by M.SssI and then linearized by NheI. The in vitro cleavage assay was performed as follows: for a 20 uL cleavage reaction, 10 uL of cell lysate with incubated with 2 uL cleavage buffer (100 mM HEPES, 500 mM KCl, 25 mM MgCl2, 5 mM DTT, 25% glycerol), the in vitro transcribed RNA, and 300 ng pUC19 plasmid DNA.
Deep sequencing to assess targeting specificity: HEK 293FT cells plated in 96-well plates were transfected with Cas9 plasmid DNA and single guide RNA (sgRNA) PCR cassette 72 hours prior to genomic DNA extraction (FIG. 14). The genomic region flanking the CRISPR target site for each gene was amplified by a fusion PCR method to attach the Illumina P5 adapters as well as unique sample-specific barcodes to the target amplicons. PCR products were purified using EconoSpin 96-well Filter Plates (Epoch Life Sciences) following the manufacturer's recommended protocol.
Barcoded and purified DNA samples were quantified by Quant-iT PicoGreen dsDNA Assay Kit or Qubit 2.0 Fluorometer (Life Technologies) and pooled in an equimolar ratio. Sequencing libraries were then deep sequenced with the Illumina MiSeq Personal Sequencer (Life Technologies).
Sequencing data analysis and indel detection: MiSeq reads were filtered by requiring an average Phred quality (Q score) of at least 23, as well as perfect sequence matches to barcodes and amplicon forward primers. Reads from on- and off-target loci were analyzed by first performing Smith-Waterman alignments against amplicon sequences that included 50 nucleotides upstream and downstream of the target site (a total of 120 bp). Alignments, meanwhile, were analyzed for indels from 5 nucleotides upstream to 5 nucleotides downstream of the target site (a total of 30 bp). Analyzed target regions were discarded if part of their alignment fell outside the MiSeq read itself, or if matched base-pairs comprised less than 85% of their total length.
Negative controls for each sample provided a gauge for the inclusion or exclusion of indels as putative cutting events. For each sample, an indel was counted only if its quality score exceeded .mu.-.sigma., where .mu. was the mean quality-score of the negative control corresponding to that sample and .sigma. was the standard deviation of same. This yielded whole target-region indel rates for both negative controls and their corresponding samples. Using the negative control's per-target-region-per-read error rate, q, the sample's observed indel count n, and its read-count R, a maximum-likelihood estimate for the fraction of reads having target-regions with true-indels, p, was derived by applying a binomial error model, as follows.
Letting the (unknown) number of reads in a sample having target regions incorrectly counted as having at least 1 indel be E, Applicants can write (without making any assumptions about the number of true indels)
.function..function..times..function..function. ##EQU00005## since R(1-p) is the number of reads having target-regions with no true indels. Meanwhile, because the number of reads observed to have indels is n, n=E+Rp, in other words the number of reads having target-regions with errors but no true indels plus the number of reads whose target-regions correctly have indels. Applicants can then re-write the above
.function..function..function..times..function. ##EQU00006##
Taking all values of the frequency of target-regions with true-indels p to be equally probable a priori, Prob(n|p).varies.Prob(p|n). The maximum-likelihood estimate (MLE) for the frequency of target regions with true-indels was therefore set as the value of p that maximized Prob(n|p). This was evaluated numerically.
In order to place error bounds on the true-indel read frequencies in the sequencing libraries themselves, Wilson score intervals (2) were calculated for each sample, given the MLE-estimate for true-indel target-regions, Rp, and the number of reads R. Explicitly, the lower bound l and upper bound u were calculated as
.times..function. ##EQU00007## .times..function. ##EQU00007.2## where z, the standard score for the confidence required in normal distribution of variance 1, was set to 1.96, meaning a confidence of 95%.
qRT-PCR analysis of relative Cas9 and sgRNA expression: 293FT cells plated in 24-well plates were transfected as described above. 72 hours post-transfection, total RNA was harvested with miRNeasy Micro Kit (Qiagen). Reverse-strand synthesis for sgRNAs was performed with qScript Flex cDNA kit (VWR) and custom first-strand synthesis primers (Table 6). qPCR analysis was performed with Fast SYBR Green Master Mix (Life Technologies) and custom primers (Table 2), using GAPDH as an endogenous control. Relative quantification was calculated by the .DELTA..DELTA.CT method.
TABLE-US-00007 TABLE 1 Target site sequences. Tested target sites for S. pyogenes type II CRISPR system with the requisite PAM. Cells were transfected with Cas9 and either crRNA-tracrRNA or chimeric sgRNA for each target. Target site genomic Target site sequence ID target (5' to 3') PAM strand 1 EMX1 GTCACCTCCAATGACTAGGG TGG + 2 EMX1 GACATCGATGTCCTCCCCAT TGG - 3 EMX1 GAGTCCGAGCAGAAGAAGAA GGG + 6 EMX1 GCGCCACCGGTTGATGTGAT GGG - 10 EMX1 GGGGCACAGATGAGAAACTC AGG - 11 EMX1 GTACAAACGGCAGAAGCTGG AGG + 12 EMX1 GGCAGAAGCTGGAGGAGGAA GGG + 13 EMX1 GGAGCCCTTCTTCTTCTGCT CGG - 14 EMX1 GGGCAACCACAAACCCACGA GGG + 15 EMX1 GCTCCCATCACATCAACCGG TGG + 16 EMX1 GTGGCGCATTGCCACGAAGC AGG + 17 EMX1 GGCAGAGTGCTGCTTGCTGC TGG + 18 EMX1 GCCCCTGCGTGGGCCCAAGC TGG + 19 EMX1 GAGTGGCCAGAGTCCAGCTT GGG - 20 EMX1 GGCCTCCCCAAAGCCTGGCC AGG - 4 PVALB GGGGCCGAGATTGGGTGTTC AGG + 5 PVALB GTGGCGAGAGGGGCCGAGAT TGG + 1 SERPINB5 GAGTGCCGCCGAGGCGGGGC GGG + 2 SERPINB5 GGAGTGCCGCCGAGGCGGGG CGG + 3 SERPINB5 GGAGAGGAGTGCCGCCGAGG CGG +
TABLE-US-00008 TABLE 2 Primer sequences SURVEYOR assay genomic primer sequence primer name target (5' to 3') Sp-EMX1-F1 EMX1 AAAACCACCCTTCTCTCTGGC Sp-EMX1-R1 EMX1 GGAGATTGGAGACACGGAGAG Sp-EMX1-F2 EMX1 CCATCCCCTTCTGTGAATGT Sp-EMX1-R2 EMX1 GGAGATTGGAGACACGGAGA Sp-PVALB-F PVALB CTGGAAAGCCAATGCCTGAC Sp-PVALB-R PVALB GGCAGCAAACTCCTTGTCCT primer sequence primer name (5' to 3') qRT-PCR for Cas9 and sgRNA expression sgRNA reverse- AAGCACCGACTCGGTGCCAC strand synthesis EMX1.1 sgRNA TCACCTCCAATGACTAGGGG qPCR F EMX1.1 sgRNA CAAGTTGATAACGGACTAGCCT qPCR R EMX1.3 sgRNA AGTCCGAGCAGAAGAAGAAGTTT qPCR F EMX1.3 sgRNA TTTCAAGTTGATAACGGACTAGCCT qPCR R Cas9 qPCR F AAACAGCAGATTCGCCTGGA Cas9 qPCR R TCATCCGCTCGATGAAGCTC GAPDH qPCR F TCCAAAATCAAGTGGGGCGA GAPDH qPCR R TGATGACCCTTTTGGCTCCC Bisulfite PCR and sequencing Bisulfite PCR F GAGGAATTCTTTTTTTGTTYGAATATGTTG (SERPINB5 locus) GAGGTTTTTTGGAAG Bisulfite PCR R GAGAAGCTTAAATAAAAAACRACAATACTC (SERPINB5 locus) AACCCAACAACC pUC19 sequencing CAGGAAACAGCTATGAC
TABLE-US-00009 TABLE 3 Sequences for primers to test sgRNA architecture. Primers hybridize to the reverse strand of the U6 promoter unless otherwise indicated. The U6 priming site is in bold, the guide sequence is indicated by the stretch of "N"s, the direct repeat sequence is in italics, and the tracrRNA sequence is underlined. The secondary structure of each sgRNA architecture is shown in FIG. 71. primer sequence primer name (5' to 3') U6-Forward GCCTCTAGAGGTACCTGAGGGCCTAT TTCCCATGATTCC I: sgRNA (DR + 12, ACCTCTAGAAAAAAAGCACCGACTCG tracrRNA + 85) GTGCCACTTTTTCAAGTTGATAACGG ACTAGCCTTATTTTAACTTGCTATTT CTAGCTCTAAAACNNNNNNNNNNNNN NNNNNNNGGTGTTTCGTCCTTTCCAC AAG II: sgRNA (DR + 12, ACCTCTAGAAAAAAAGCACCGACTCG tracrRNA + 85) mut2 GTGCCACTTTTTCAAGTTGATAACGG ACTAGCCTTATATTAACTTGCTATTT CTAGCTCTAATACNNNNNNNNNNNNN NNNNNNNGGTGTTTCGTCCTTTCCAC AAG III: sgRNA (DR + ACCTCTAGAAAAAAAGCACCGACTCG 22, tracrRNA + 85) GTGCCACTTTTTCAAGTTGATAACGG ACTAGCCTTATTTTAACTTGCTATGC TGTTTTGTTTCCAAAACAGCATAGCT CTAAAACNNNNNNNNNNNNNNNNNNN NGGTGTTTCGTCCTTTCCACAAG IV: sgRNA (DR + 22, ACCTCTAGAAAAAAAGCACCGACTCG tracrRNA + 85) mut4 GTGCCACTTTTTCAAGTTGATAACGG ACTAGCCTTATATTAACTTGCTATGC TGTATTGTTTCCAATACAGCATAGCT CTAATACNNNNNNNNNNNNNNNNNNN NGGTGTTTCGTCCTTTCCACAAG
TABLE-US-00010 TABLE 4 Target sites with alternate PAMs for testing PAM specificity of Cas9. All target sites for PAM specificity testing are found within the human EMX1 locus. Target site sequence (5' to 3') PAM AGGCCCCAGTGGCTGCTCT NAA ACATCAACCGGTGGCGCAT NAT AAGGTGTGGTTCCAGAACC NAC CCATCACATCAACCGGTGG NAG AAACGGCAGAAGCTGGAGG NTA GGCAGAAGCTGGAGGAGGA NTT GGTGTGGTTCCAGAACCGG NTC AACCGGAGGACAAAGTACA NTG TTCCAGAACCGGAGGACAA NCA GTGTGGTTCCAGAACCGGA NCT TCCAGAACCGGAGGACAAA NCC CAGAAGCTGGAGGAGGAAG NCG CATCAACCGGTGGCGCATT NGA GCAGAAGCTGGAGGAGGAA NGT CCTCCCTCCCTGGCCGAGG NGC TCATCTGTGCCCCTCCCTC NAA GGGAGGACATCGATGTCAC NAT CAAACGGCAGAAGCTGGAG NAC GGGTGGGCAACCACAAACC NAG GGTGGGCAACCACAAACCC NTA GGCTCCCATCACATCAACC NTT GAAGGGCCTGAGTCCGAGC NTC CAACCGGTGGCGCATTGCC NTG AGGAGGAAGGGCCTGAGTC NCA AGCTGGAGGAGGAAGGGCC NCT GCATTGCCACGAAGCAGGC NCC ATTGCCACGAAGCAGGCCA NCG AGAACCGGAGGACAAAGTA NGA TCAACCGGTGGCGCATTGC NGT GAAGCTGGAGGAGGAAGGG NGC
Sequences
All sequences are in the 5' to 3' direction. For U6 transcription, the string of underlined Ts serve as the transcriptional terminator.
TABLE-US-00011 > U6-short tracrRNA (Streptococcus pyogenes SF370) gagggcctatttcccatgattccttcatatttgcatatacgatacaagga gttagagagataattggaattaatttgactgtaaacacaaagatattagt acaaaatacgtgacgtagaaagtaataatttcttgggtagtttgcagttt taaaattatgttttaaaatggactatcatatgcttaccgtaacttgaaag tatttcgatttcaggctttatatatcagtggaaaggacgaaacaccGGAA CCATTCAAAACAGCATACCAACTTAAAATAAGGCTAGTCCGTTATCAACT TGAAAAAGTGGCACCGAGTCGGTGCTTTTTTT (tracrRNA sequence is in bold) > U6-DR-guide sequence-DR (Streptococcus pyogenes SF370) gagggcctatttcccatgattccttcatatttgcatatacgatacaagga gttagagagataattggaattaatttgactgtaaacacaaagatattagt acaaaatacgtgacgtagaaagtaataatttcttgggtagtttgcagttt taaaattatgttttaaaatggactatcatatgcttaccgtaacttgaaag tatttcgatttcttggctttatatatcagtggaaaggacgaaacaccggg ttttagagctatgctgttttgaatggtcccaaaacNNNNNNNNNNNNNNN NNNNNNNNNNNNNNNgttttagagctatgctgttttgaatggtcccaaaa cTTTTTTT (direct repeat sequence is in italics and the guide sequence is indicated by the stretch of "N"s) > sgRNA containing + 48 tracrRNA (Streptococcus pyogenes SF370) gagggcctatttcccatgattccttcatatttgcatatacgatacaaggc tgttagagagataattggaattaatttgactgtaaacacaaagatattag tacaaaatacgtgacgtagaaagtaataatttcttgggtagtttgcagtt ttaaaattatgttttaaaatggactatcatatgcttaccgtaacttgaaa gtatttcgatttcttggctttatatatcttgtggaaaggacgaaacaccN NNNNNNNNNNNNNNNNNNNgttttagagctagaaatagcaagttaaaata aggctagtccgTTTTTTT (guide sequence is highlighted in blue and the tracrRNA fragment is in bold) > sgRNA containing + 54 tracrRNA (Streptococcus pyogenes SF370) gagggcctatacccatgattccttcatatttgcatatacgatacaaggct gttagagagataattggaattaatttgactgtaaacacaaagatattagt acaaaatacgtgacgtagaaagtaataatttcttgggtagtttgcagttt taaaattatgttttaaaatggactatcatatgcttaccgtaacttgaaag tatttcgatttcttggctttatatatcttgtggaaaggacgaaacaccNN NNNNNNNNNNNNNNNNNNgttttagagctagaaatagcaagttaaaataa ggctagtccgttatcaTTTTTTTT (guide sequence is indicated by the stretch of "N"s and the tracrRNA fragment is in bold) > sgRNA containing + 67 tracrRNA (Streptococcus pyogenes SF370) gagggcctatttcccatgattccttcatatttgcatatacgatacaaggc tgttagagagataattggaattaatttgactgtaaacacaaagatattag tacaaaatacgtgacgtagaaagtaataatttcttgggtagtttgcagat taaaattatgattaaaatggactatcatatgcttaccgtaacttgaaagt atttcgatttcttggctttatatatcttgtggaaaggacgaaacaccNNN NNNNNNNNNNNNNNNNNgttttagagctagaaatagcaagttaaaataag gctagtccgttatcaactttgaaaaagtgTTTTTTT (guide sequence is indicated by the stretch of "N"s and the tracrRNA fragment is in bold)) > sgRNA containing + 85 tracrRNA (Streptococcus pyogenes SF370) gagggcctatttcccatgattccttcatatttgcatatacgatacaaggc tgttagagagataattggaattaatttgactgtaaacacaaagatattag tacaaaatacgtgacgtagaaagtaataatttcttgggtagtttgcagtt ttaaaattatgattaaaatggactatcatatgcttaccgtaacttgaaag tatttcgatttcttggctttatatatcttgtggaaaggacgaaacaccNN NNNNNNNNNNNNNNNNNNgttttagagctagaaatagcaagttaaaataa ggctagtccgttatcaacttgaaaaagtggcaccgagtcggtgcTTTTTT T (guide sequence is indicated by the stretch of "N"s and the tracrRNA fragment is in bold) > CBh-NLS-SpCas9-NLS CGTTACATAACTTACGGTAAATGGCCCGCCTGGCTGACCGCCCAACGACC CCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAACGCCAATA GGGACTTTCCATTGACGTCAATGGGTGGAGTATTTACGGTAAACTGCCCA CTTGGCAGTACATCAAGTGTATCATATGCCAAGTACGCCCCCTATTGACG TCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTA TGGGACTTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTAC CATGGTCGAGGTGAGCCCCACGTTCTGCTTCACTCTCCCCATCTCCCCCC CCTCCCCACCCCCAATTTTGTATTTATTTATTTTTTAATTATTTTGTGCA GCGATGGGGGCGGGGGGGGGGGGGGGGCGCGCGCCAGGCGGGGCGGGGCG GGGCGAGGGGCGGGGCGGGGCGAGGCGGAGAGGTGCGGCGGCAGCCAATC AGAGCGGCGCGCTCCGAAAGTTTCCTTTTATGGCGAGGCGGCGGCGGCGG CGGCCCTATAAAAAGCGAAGCGCGCGGCGGGCGGGAGTCGCTGCGACGCT GCCTTCGCCCCGTGCCCCGCTCCGCCGCCGCCTCGCGCCGCCCGCCCCGG CTCTGACTGACCGCGTTACTCCCACAGGTGAGCGGGCGGGACGGCCCTTC TCCTCCGGGCTGTAATTAGCTGAGCAAGAGGTAAGGGTTTAAGGGATGGT TGGTTGGTGGGGTATTAATGTTTAATTACCTGGAGCACCTGCCTGAAATC ACTTTTTTTCAGGTTGGaccggtgccaccATGGACTATAAGGACCACGAC GGAGACTACAAGGATCATGATATTGATTACAAAGACGATGACGATAAGAT GGCCCCAAAGAAGAAGCGGAAGGTCGGTATCCACGGAGTCCCAGCAGCCG ACAAGAAGTACAGCATCGGCCTGGACATCGGCACCAACTCTGTGGGCTGG GCCGTGATCACCGACGAGTACAAGGTGCCCAGCAAGAAATTCAAGGTGCT GGGCAACACCGACCGGCACAGCATCAAGAAGAACCTGATCGGAGCCCTGC TGTTCGACAGCGGCGAAACAGCCGAGGCCACCCGGCTGAAGAGAACCGCC AGAAGAAGATACACCAGACGGAAGAACCGGATCTGCTATCTGCAAGAGAT CTTCAGCAACGAGATGGCCAAGGTGGACGACAGCTTCTTCCACAGACTGG AAGAGTCCTTCCTGGTGGAAGAGGATAAGAAGCACGAGCGGCACCCCATC TTCGGCAACATCGTGGACGAGGTGGCCTACCACGAGAAGTACCCCACCAT CTACCACCTGAGAAAGAAACTGGTGGACAGCACCGACAAGGCCGACCTGC GGCTGATCTATCTGGCCCTGGCCCACATGATCAAGTTCCGGGGCCACTTC CTGATCGAGGGCGACCTGAACCCCGACAACAGCGACGTGGACAAGCTGTT CATCCAGCTGGTGCAGACCTACAACCAGCTGTTCGAGGAAAACCCCATCA ACGCCAGCGGCGTGGACGCCAAGGCCATCCTGTCTGCCAGACTGAGCAAG AGCAGACGGCTGGAAAATCTGATCGCCCAGCTGCCCGGCGAGAAGAAGAA TGGCCTGTTCGGCAACCTGATTGCCCTGAGCCTGGGCCTGACCCCCAACT TCAAGAGCAACTTCGACCTGGCCGAGGATGCCAAACTGCAGCTGAGCAAG GACACCTACGACGACGACCTGGACAACCTGCTGGCCCAGATCGGCGACCA GTACGCCGACCTCTTTCTGGCCGCCAAGAACCTGTCCGACGCCATCCTGC TGAGCGACATCCTGAGAGTGAACACCGAGATCACCAAGGCCCCCCTGAGC GCCTCTATGATCAAGAGATACGACGAGCACCACCAGGACCTGACCCTGCT GAAAGCTCTCGTGCGGCAGCAGCTGCCTGAGAAGTACAAAGAGATTTTCT TCGACCAGAGCAAGAACGGCTACGCCGGCTACATTGACGGCGGAGCCAGC CAGGAAGAGTTCTACAAGTTCATCAAGCCCATCCTGGAAAAGATGGACGG CACCGAGGAACTGCTCGTGAAGCTGAACAGAGAGGACCTGCTGCGGAAGC AGCGGACCTTCGACAACGGCAGCATCCCCCACCAGATCCACCTGGGAGAG CTGCACGCCATTCTGCGGGGGCAGGAAGATTTTTACCCATTCCTGAAGGA CAACCGGGAAAAGATCGAGAAGATCCTGACCTTCCGCATCCCCTACTACG TGGGCCCTCTGGCCAGGGGAAACAGCAGATTCGCCTGGATGACCAGAAAG AGCGAGGAAACCATCACCCCCTGGAACTTCGAGGAAGTGGTGGACAAGGG CGCTTCCGCCCAGAGCTTCATCGAGCGGATGACCAACTTCGATAAGAACC TGCCCAACGAGAAGGTGCTGCCCAAGCACAGCCTGCTGTACGAGTACTTC ACCGTGTATAACGAGCTGACCAAAGTGAAATAGGTGACCGAGGGAATGAG AAAGCCCGCCTTCCTGAGCGGCGAGCAGAAAAAGGCCATCGTGGACCTGC TGTTCAAGACCAACCGGAAAGTGACCGTGAAGCAGCTGAAAGAGGACTAC TTCAAGAAAATCGAGTGCTTCGACTCCGTGGAAATCTCCGGCGTGGAAGA TCGGTTCAACGCCTCCCTGGGCACATACCACGATCTGCTGAAAATTATCA AGGACAAGGACTTCCTGGACAATGAGGAAAACGAGGACATTCTGGAAGAT ATCGTGCTGACCCTGACACTGTTTGAGGACAGAGAGATGATCGAGGAACG GCTGAAAACCTATGCCCACCTGTTCGACGACAAAGTGATGAAGCAGCTGA AGCGGCGGAGATACACCGGCTGGGGCAGGCTGAGCCGGAAGCTGATCAAC GGCATCCGGGACAAGCAGTCCGGCAAGACAATCCTGGATTTCCTGAAGTC CGACGGCTTCGCCAACAGAAACTTCATGCAGCTGATCCACGACGACAGCC TGACCTTTAAAGAGGACATCCAGAAAGCCCAGGTGTCCGGCCAGCGCGAT AGCCTGCACGAGCACATTGCCAATCTGGCCGGCAGCCCCGCCATTAAGAA GGGCATCCTGCAGACAGTGAAGGTGGTGGACGAGCTCGTGAAAGTGATGG GCCGGCACAAGCCCGAGAACATCGTGATCGAAATGGCCAGAGAGAACCAG ACCACCCAGAAGGGACAGAAGAACAGCCGCGAGAGAATGAAGCGGATCGA AGAGGGCATCAAAGAGCTGGGCAGCCAGATCCTGAAAGAACACCCCGTGG AAAACACCCAGCTGCAGAACGAGAAGCTGTACCTGTACTACCTGCAGAAT GGGCGGGATATGTACGTGGACCAGGAACTGGACATCAACCGGCTGTCCGA
CTACGATGTGGACCATATCGTGCCTCAGAGCTTTCTGAAGGACGACTCCA TCGACAACAAGGTGCTGACCAGAAGCGACAAGAACCGGGGCAAGAGCGAC AACGTGCCCTCCGAACAGGTCGTGAAGAAGATGAAGAACTACTGGCGGCA GCTGCTGAACGCCAAGCTGATTACCCAGAGAAAGTTCGACAATCTGACCA AGGCCGAGAGAGGCGGCCTGAGCGAACTGGATAAGGCCGGCTTCATCAAG AGACAGCTGGTGGAAACCCGGCAGATCACAAAGCACGTGGCACAGATCCT GGACTCCCGGATGAACACTAAGTACGACGAGAATGACAAGCTGATCCGGG AAGTGAAAGTGATCACCCTGAAGTCCAAGCTGGTGTCCGATTTCCGGAAG GATTTCCAGTTTTACAAAGTGCGCGAGATCAACAACTACCACCACGCCCA CGACGCCTACCTGAACGCCGTCGTGGGAACCGCCCTGATCAAAAAGTACC CTAAGCTGGAAAGCGAGTTCGTGTACGGCGACTACAAGGTGTACGACGTG CGGAAGATGATCGCCAAGAGCGAGCAGGAAATCGGCAAGGCTACCGCCAA GTACTTCTTCTACAGCAACATCATGAACTTTTTCAAGACCGAGATTACCC TGGCCAACGGCGAGATCCGGAAGCGGCCTCTGATCGAGACAAACGGCGAA ACCGGGGAGATCGTGTGGGATAAGGGCCGGGATTTTGCCACCGTGCGGAA AGTGCTGAGCATGCCCCAAGTGAATATCGTGAAAAAGACCGAGGTGCAGA CAGGCGGCTTCAGCAAAGAGTCTATCCTGCCCAAGAGGAACAGCGATAAG CTGATCGCCAGAAAGAAGGACTGGGACCCTAAGAAGTACGGCGGCTTCGA CAGCCCCACCGTGGCCTATTCTGTGCTGGTGGTGGCCAAAGTGGAAAAGG GCAAGTCCAAGAAACTGAAGAGTGTGAAAGAGCTGCTGGGGATCACCATC ATGGAAAGAAGCAGCTTCGAGAAGAATCCCATCGACTTTCTGGAAGCCAA GGGCTACAAAGAAGTGAAAAAGGACCTGATCATCAAGCTGCCTAAGTACT CCCTGTTCCAGCTGGAAAACGGCCGGAAGAGAATGCTGGCCTCTGCCGGC GAACTGCAGAAGGGAAACGAACTGGCCCTGCCCTCCAAATATGTGAACTT CCTGTACCTGGCCAGCCACTATGAGAAGCTGAAGGGCTCCCCCGAGGATA ATGAGCAGAAACAGCTGTTTGTGGAACAGCACAAGCACTACCTGGACGAG ATCATCGAGCAGATCAGCGAGTTCTCCAAGAGAGTGATCCTGGCCGACGC TAATCTCGACAAAGTGCTGTCCGCCTACAACAAGCACCGGGATAAGCCCA TCAGAGAGCAGGCCGAGAATATCATCCACCTGTTTACCCTGACCAATCTG GGAGCCCCTGCCGCCTTCAAGTACTTTGACACCACCATCGACCGGAAGAG GTACACCAGCACCAAAGAGGTGCTGGACGCCACCCTGATCCACCAGAGCA TCACCGGCCTGTACGAGACACGGATCGACCTGTCTCAGCTGGGAGGCGAC TTTCTTTTTCTTAGCTTGACCAGCTTTCTTAGTAGCAGCAGGACGCTTTA A (NLS-hSpCas9-NLS is in bold) > Sequencing amplicon for EMX1 guides 1.1, 1.14, 1.17 CCAATGGGGAGGACATCGATGTCACCTCCAATGACTAGGGTGGGCAACCA CAAACCCACGAGGGCAGAGTGCTGCTTGCTGCTGGCCAGGCCCCTGCGTG GGCCCAAGGTGGACTCTGGCCAC > Sequencing amplicon for EMX1 guides 1.2, 1.16 CGAGCAGAAGAAGAAGGGCTCCCATCACATCAACCGGTGGCGCATTGCCA CGAAGCAGGCCAATGGGGAGGACATCGATGTCACCTCCAATGACTAGGGT GGGCAACCACAAACCCACGAG > Sequencing amplicon for EMX1 guides 1.3, 1.13, 1.15 GGAGGACAAAGTACAAACGGCAGAAGCTGGAGGAGGAAGGGCCTGAGTCC GAGCAGAAGAAGAAGGGCTCCCATCACATCAACCGGTGGCGCATTGCCAC GAAGCAGGCCAATGGGGAGGACATCGAT > Sequencing amplicon for EMX1 guides 1.6 AGAAGCTGGAGGAGGAAGGGCCTGAGTCCGAGCAGAAGAAGAAGGGCTCC CATCACATCAACCGGTGGCGCATTGCCACGAAGCAGGCCAATGGGGAGGA CATCGATGTCACCTCCAATGACTAGGGTGG > Sequencing amplicon for EMX1 guides 1.10 CCTCAGTCTTCCCATCAGGCTCTCAGCTCAGCCTGAGTGTTGAGGCCCCA GTGGCTGCTGTGGGGGCCTCCTGAGTTTCTCATCTGTGCCCCTCCCTCCC TGGCCCAGGTGAAGGTGTGGTTCCA > Sequencing amplicon for EMX1 guides 1.11, 1.12 TCATCTGTGCCCCTCCCTCCCTGGCCCAGGTGAAGGTGTGGTTCCAGAAC CGGAGGACAAAGTACAAACGGCAGAAGCTGGAGGAGGAAGGGCCTGAGTC CGAGCAGAAGAAGAAGGGCTCCCATCACA > Sequencing amplicon for EMX1 guides 1.18, 1.19 CTCCAATGACTAGGGTGGGCAACCACAAACCCACGAGGGCAGAGTGCTGC TTGCTGCTGGCCAGGCCCCTGCGTGGGCCCAAGCTGGACTCTGGCCACTC CCTGGCCAGGCTTTGGGGAGGCCTGGAGT > Sequencing amplicon for EMX1 guides 1.20 CTGCTTGCTGCTGGCCAGGCCCCTGCGTGGGCCCAAGCTGGACTCTGGCC ACTCCCTGGCCAGGCTTTGGGGAGGCCTGGAGTCATGGCCCCACAGGGCT TGAAGCCCGGGGCCGCCATTGACAGAG > T7 promoter F primer for annealing with target strand GAAATTAATACGACTCACTATAGGG > oligo containing pUC19 target site 1 for methylation (T7 reverse) AAAAAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAACGGACTAGCC TTATTTTAACTTTGCTATTTCTAGCTCTAAAACAACGACGAGCGTGACAC CACCCTATAGTGAGTCGTATTAATTTC > oligo containing pUC19 target site 2 for methylation (T7 reverse) AAAAAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAACGGACTAGCC TTATTTTAACTTGCTATTTCTAGCTCTAAAACGCAACAATTAATAGACTG GACCTATAGTGAGTCGTATTAATTTC
Example 7
Base Pair Mismatching Investigations
Applicants have tested several mismatch guide sequences have been tested to probe cas9 protein off-target effect (Hsu et al., DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol. 2013 September; 31(9):827-32). However the number of mismatch guide sequences used may have been far less than the complete set of all possible mismatch guide sequences. In order to predict off target cutting efficiency of non-tested guide-target pairs, a thermodynamic model is built to fit existing experimental data and is used to predict new guide target pair.
Applicants tested cutting frequency data for tested guide target pair collected from Hsu P. D et al., 2013. Applicants collected the nearest neighbor thermodynamic parameters for nucleic acid pairing from ViennaRNA package distribution. Applicants obtained first bi-base pair thermal dynamical parameters per position for each guide target pair. These parameters were referred to as features. Data, comprising of cutting frequency and features for each guide target pair, was divided into 5 groups randomly in order to perform 5 fold cross validation. In each training session, 4 groups of data were used as training data. A modified radial basis kernel that consisted of radial basis kernel with an additive term accounting for different PAM was used in nonlinear regression. In each testing session, the remaining 1 group was used to test the model. Coarse grid search for fitting parameters was performed. The best fitting parameters were used to generate new parameters in fine grid search.
The Applicants' algorithm flows as follows: 1. Choose data with duplicate. 2. Calculate confidence interval for cutting frequency based on binomial distribution. Select data with p value passing a threshold. 3. Calculate thermal dynamic parameters for each guide-target pair to obtain its feature. 4. Transform non-linearly cutting frequency data in order to balance high and low cutting frequency contributions in model fitting. 5. Divide data into 5 groups. 6. Perform coarse grid search for fitting parameter using 5 fold cross validation. 7. Best parameters are selected, and fine grid search is performed.
Result: Training data and testing data are shown as blue dots and red dots respectively in the scatter plot. Spearman correlation coefficient is 0.88, and Pearson correlation coefficient is 0.91 (FIG. 36). It demonstrates good model prediction of the data. In order to check model over fitting, data was randomized again, trained and tested using the same parameters. The result is 0.82 for both correlation coefficients (FIG. 37).
REFERENCES
1. Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819-823 (2013) 2. Mali, P. et al. RNA-Guided Human Genome Engineering via Cas9. Science 339, 823-826 (2013). 3. Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013). 4. Cho, S. W., Kim, S., Kim, J. M. & Kim, J. S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31, 230-232 (2013). 5. Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602-607 (2011). 6. Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816-821 (2012). 7. Wang, H. et al. One-Step Generation of Mice Carrying Mutations in Multiple Genes by CRISPR/Cas-Mediated Genome Engineering. Cell 153, 910-918 (2013). 8. Guschin, D. Y. et al. A rapid and general assay for monitoring endogenous gene modification. Methods Mol Biol 649, 247-256 (2010). 9. Bogenhagen, D. F. & Brown, D. D. Nucleotide sequences in Xenopus 5S DNA required for transcription termination. Cell 24, 261-270 (1981). 10. Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol 31, 227-229 (2013). 11. Bultmann, S. et al. Targeted transcriptional activation of silent oct4 pluripotency gene by combining designer TALEs and inhibition of epigenetic modifiers. Nucleic Acids Res 40, 5368-5377 (2012). 12. Valton, J. et al. Overcoming transcription activator-like effector (TALE) DNA binding domain sensitivity to cytosine methylation. J Biol Chem 287, 38427-38432 (2012). 13. Christian, M. et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186, 757-761 (2010). 14. Miller, J. C. et al. A TALE nuclease architecture for efficient genome editing. Nat Biotechnol 29, 143-148 (2011). 15. Mussolino, C. et al. A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic acids research 39, 9283-9293 (2011). 16. Hsu, P. D. & Zhang, F. Dissecting neural function using targeted genome engineering technologies. ACS chemical neuroscience 3, 603-610 (2012). 17. Sanjana, N. E. et al. A transcription activator-like effector toolbox for genome engineering. Nature protocols 7, 171-192 (2012). 18. Porteus, M. H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003). 19. Miller, J. C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat Biotechnol 25, 778-785 (2007). 20. Sander, J. D. et al. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat Methods 8, 67-69 (2011). 21. Wood, A. J. et al. Targeted genome editing across species using ZFNs and TALENs.
Science 333, 307 (2011). 22. Bobis-Wozowicz, S., Osiak, A., Rahman, S. H. & Cathomen, T. Targeted genome editing in pluripotent stem cells using zinc-finger nucleases. Methods 53, 339-346 (2011). 23. Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L. A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol 31, 233-239 (2013). 24. Qi, L. S. et al. Repurposing CRISPR as an RNA-Guided Platform for Sequence-Specific Control of Gene Expression. Cell 152, 1173-1183 (2013). 25. Michaelis, L. M., Maud "Die kinetik der invertinwirkung.". Biochem. z (1913). 26. Mahfouz, M. M. et al. De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks. Proc Natl Acad Sci USA 108, 2623-2628 (2011). 27. Wilson, E. B. Probable inference, the law of succession, and statistical inference. J Am Stat Assoc 22, 209-212 (1927). 28. Ding, Q. et al. A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell 12, 238-251 (2013). 29. Soldner, F. et al. Generation of isogenic pluripotent stem cells differing exclusively at two early onset Parkinson point mutations. Cell 146, 318-331 (2011). 30. Carlson, D. F. et al. Efficient TALEN-mediated gene knockout in livestock. Proc Natl Acad Sci USA 109, 17382-17387 (2012). 31. Geurts, A. M. et al. Knockout Rats via Embryo Microinjection of Zinc-Finger Nucleases. Science 325, 433-433 (2009). 32. Takasu, Y. et al. Targeted mutagenesis in the silkworm Bombyx mori using zinc finger nuclease mRNA injection. Insect Biochem Molec 40, 759-765 (2010). 33. Watanabe, T. et al. Non-transgenic genome modifications in a hemimetabolous insect using zinc-finger and TAL effector nucleases. Nat Commun 3 (2012). 34. Reyon, D. et al. FLASH assembly of TALENs for high-throughput genome editing. Nat Biotechnol 30, 460-465 (2012). 35. Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type 111 effectors. Science 326, 1509-1512 (2009). 36. Moscou, M. J. & Bogdanove, A. J. A simple cipher governs DNA recognition by TAL effectors. Science 326, 1501 (2009). 37. Deveau, H., Garneau, J. E. & Moineau, S. CRISPR/Cas system and its role in phage-bacteria interactions. Annu Rev Microbiol 64, 475-493 (2010). 38. Horvath, P. & Barrangou, R. CRISPR/Cas, the immune system of bacteria and archaea. Science 327, 167-170 (2010). 39. Makarova, K. S. et al. Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 9, 467-477 (2011). 40. Bhaya, D., Davison, M. & Barrangou, R. CRISPR-Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annu Rev Genet 45, 273-297 (2011). 41. Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67-71 (2010). 42. Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci USA 109, E2579-2586 (2012). 43. Urnov, F. D., Rebar, E. J., Holmes, M. C., Zhang, H. S. & Gregory, P. D. Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11, 636-646 (2010). 44. Perez, E. E. et al. Establishment of HIV-1 resistance in CD4(+) T cells by genome editing using zinc-finger nucleases. Nat Biotechnol 26, 808-816 (2008). 45. Chen, F. Q. et al. High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases. Nat Methods 8, 753-U796 (2011). 46. Bedell, V. M. et al. In vivo genome editing using a high-efficiency TALEN system. Nature 491, 114-U133 (2012). 47. Saleh-Gohari, N. & Helleday, T. Conservative homologous recombination preferentially repairs DNA double-strand breaks in the S phase of the cell cycle in human cells. Nucleic Acids Res 32, 3683-3688 (2004). 48. Sapranauskas, R. et al. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res 39, 9275-9282 (2011). 49. Shen, B. et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res 23, 720-723 (2013). 50. Tuschl, T. Expanding small RNA interference. Nat Biotechnol 20, 446-448 (2002). 51. Smithies, O., Gregg, R. G., Boggs, S. S., Koralewski, M. A. & Kucherlapati, R. S. Insertion of DNA sequences into the human chromosomal beta-globin locus by homologous recombination. Nature 317, 230-234 (1985). 52. Thomas, K. R., Folger, K. R. & Capecchi, M. R. High frequency targeting of genes to specific sites in the mammalian genome. Cell 44, 419-428 (1986). 53. Hasty, P., Rivera-Perez, J. & Bradley, A. The length of homology required for gene targeting in embryonic stem cells. Mol Cell Biol 11, 5586-5591 (1991). 54. Wu, S., Ying, G. X., Wu, Q. & Capecchi, M. R. A protocol for constructing gene targeting vectors: generating knockout mice for the cadherin family and beyond. Nat Protoc 3, 1056-1076 (2008). 55. Oliveira, T. Y. et al. Translocation capture sequencing: a method for high throughput mapping of chromosomal rearrangements. J Immunol Methods 375, 176-181 (2012). 56. Tremblay et al., Transcription Activator-Like Effector Proteins Induce the Expression of the Frataxin Gene; Human Gene Therapy. August 2012, 23(8): 883-890. 57. Shalek et al. Nanowire-mediated delivery enables functional interrogation of primary immune cells: application to the analysis of chronic lymphocytic leukemia. Nano Letters, 2012, Dec. 12; 12(12):6498-504. 58. Pardridge et al. Preparation of Trojan horse liposomes (THLs) for gene transfer across the blood-brain barrier; Cold Spring Harb Protoc; 2010; April; 2010 (4) 59. Plosker G L et al. Fluvastatin: a review of its pharmacology and use in the management of hypercholesterolaemia; Drugs 1996, 51(3):433-459). 60. Trapani et al. Potential role of nonstatin cholesterol lowering agents; IUBMB Life, Volume 63, Issue 11, pages 964-971, November 2011 61. Birch A M et al. DGAT1 inhibitors as anti-obesity and anti-diabetic agents; Current Opinion in Drug Discovery & Development, 2010, 13(4):489-496 62. Fuchs et al. Killing of leukemic cells with a BCR/ABL fusion gene by RNA interference (RNAi), Oncogene 2002, 21(37):5716-5724. 63. McManaman J L et al. Perilipin-2 Null Mice are Protected Against Diet-Induced Obesity, Adipose Inflammation and Fatty Liver Disease; The Journal of Lipid Research, jlr.M035063. First Published on Feb. 12, 2013. 64. Tang J et al. Inhibition of SREBP by a Small Molecule, Betulin, Improves Hyperlipidemia and Insulin Resistance and Reduces Atherosclerotic Plaques: Cell Metabolism, Volume 13, Issue 1, 44-56, 5 Jan. 2011. 65. Dumitrache et al. Trex2 enables spontaneous sister chromatid exchanges without facilitating DNA double-strand break repair; Genetics. 2011 August; 188(4): 787-797
While preferred embodiments of the present invention have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the invention. It should be understood that various alternatives to the embodiments of the invention described herein may be employed in practicing the invention.
SEQUENCE LISTINGS
1
1076112RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1guuuuagagc ua 1227PRTSimian virus 40 2Pro Lys Lys Lys Arg Lys Val1 5316PRTUnknownsource/note="Description of Unknown Nucleoplasmin bipartite NLS sequence" 3Lys Arg Pro Ala Ala Thr Lys Lys Ala Gly Gln Ala Lys Lys Lys Lys1 5 10 1549PRTUnknownsource/note="Description of Unknown C-myc NLS sequence" 4Pro Ala Ala Lys Arg Val Lys Leu Asp1 5511PRTUnknownsource/note="Description of Unknown C-myc NLS sequence" 5Arg Gln Arg Arg Asn Glu Leu Lys Arg Ser Pro1 5 10638PRTHomo sapiens 6Asn Gln Ser Ser Asn Phe Gly Pro Met Lys Gly Gly Asn Phe Gly Gly1 5 10 15Arg Ser Ser Gly Pro Tyr Gly Gly Gly Gly Gln Tyr Phe Ala Lys Pro 20 25 30Arg Asn Gln Gly Gly Tyr 35742PRTUnknownsource/note="Description of Unknown IBB domain from importin-alpha sequence" 7Arg Met Arg Ile Glx Phe Lys Asn Lys Gly Lys Asp Thr Ala Glu Leu1 5 10 15Arg Arg Arg Arg Val Glu Val Ser Val Glu Leu Arg Lys Ala Lys Lys 20 25 30Asp Glu Gln Ile Leu Lys Arg Arg Asn Val 35 4088PRTUnknownsource/note="Description of Unknown Myoma T protein sequence" 8Val Ser Arg Lys Arg Pro Arg Pro1 598PRTUnknownsource/note="Description of Unknown Myoma T protein sequence" 9Pro Pro Lys Lys Ala Arg Glu Asp1 5108PRTHomo sapiens 10Pro Gln Pro Lys Lys Lys Pro Leu1 51112PRTMus musculus 11Ser Ala Leu Ile Lys Lys Lys Lys Lys Met Ala Pro1 5 10125PRTInfluenza virus 12Asp Arg Leu Arg Arg1 5137PRTInfluenza virus 13Pro Lys Gln Lys Lys Arg Lys1 51410PRTHepatitus delta virus 14Arg Lys Leu Lys Lys Lys Ile Lys Lys Leu1 5 101510PRTMus musculus 15Arg Glu Lys Lys Lys Phe Leu Lys Arg Arg1 5 101620PRTHomo sapiens 16Lys Arg Lys Gly Asp Glu Val Asp Gly Val Asp Glu Val Ala Lys Lys1 5 10 15Lys Ser Lys Lys 201717PRTHomo sapiens 17Arg Lys Cys Leu Gln Ala Gly Met Asn Leu Glu Ala Arg Lys Thr Lys1 5 10 15Lys1827DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(1)..(20)a, c, t or gmodified_base(21)..(22)a, c, t, g, unknown or other 18nnnnnnnnnn nnnnnnnnnn nnagaaw 271919DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(1)..(12)a, c, t or gmodified_base(13)..(14)a, c, t, g, unknown or other 19nnnnnnnnnn nnnnagaaw 192027DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(1)..(20)a, c, t or gmodified_base(21)..(22)a, c, t, g, unknown or other 20nnnnnnnnnn nnnnnnnnnn nnagaaw 272118DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(1)..(11)a, c, t or gmodified_base(12)..(13)a, c, t, g, unknown or other 21nnnnnnnnnn nnnagaaw 1822137DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(1)..(20)a, c, t, g, unknown or other 22nnnnnnnnnn nnnnnnnnnn gtttttgtac tctcaagatt tagaaataaa tcttgcagaa 60gctacaaaga taaggcttca tgccgaaatc aacaccctgt cattttatgg cagggtgttt 120tcgttattta atttttt 13723123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(1)..(20)a, c, t, g, unknown or other 23nnnnnnnnnn nnnnnnnnnn gtttttgtac tctcagaaat gcagaagcta caaagataag 60gcttcatgcc gaaatcaaca ccctgtcatt ttatggcagg gtgttttcgt tatttaattt 120ttt 12324110DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(1)..(20)a, c, t, g, unknown or other 24nnnnnnnnnn nnnnnnnnnn gtttttgtac tctcagaaat gcagaagcta caaagataag 60gcttcatgcc gaaatcaaca ccctgtcatt ttatggcagg gtgttttttt 11025102DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(1)..(20)a, c, t, g, unknown or other 25nnnnnnnnnn nnnnnnnnnn gttttagagc tagaaatagc aagttaaaat aaggctagtc 60cgttatcaac ttgaaaaagt ggcaccgagt cggtgctttt tt 1022688DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(1)..(20)a, c, t, g, unknown or other 26nnnnnnnnnn nnnnnnnnnn gttttagagc tagaaatagc aagttaaaat aaggctagtc 60cgttatcaac ttgaaaaagt gttttttt 882776DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(1)..(20)a, c, t, g, unknown or other 27nnnnnnnnnn nnnnnnnnnn gttttagagc tagaaatagc aagttaaaat aaggctagtc 60cgttatcatt tttttt 762822DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 28aggccccagt ggctgctctn aa 222922DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 29acatcaaccg gtggcgcatn at 223022DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 30aaggtgtggt tccagaaccn ac 223122DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 31ccatcacatc aaccggtggn ag 223222DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 32aaacggcaga agctggaggn ta 223322DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 33ggcagaagct ggaggaggan tt 223422DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 34ggtgtggttc cagaaccggn tc 223522DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 35aaccggagga caaagtacan tg 223622DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 36ttccagaacc ggaggacaan ca 223722DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 37gtgtggttcc agaaccggan ct 223822DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 38tccagaaccg gaggacaaan cc 223922DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 39cagaagctgg aggaggaagn cg 224022DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 40catcaaccgg tggcgcattn ga 224122DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 41gcagaagctg gaggaggaan gt 224222DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 42cctccctccc tggcccaggn gc 224322DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 43tcatctgtgc ccctccctcn aa 224422DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 44gggaggacat cgatgtcacn at 224522DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 45caaacggcag aagctggagn ac 224622DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 46gggtgggcaa ccacaaaccn ag 224722DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 47ggtgggcaac cacaaacccn ta 224822DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 48ggctcccatc acatcaaccn tt 224922DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 49gaagggcctg agtccgagcn tc 225022DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 50caaccggtgg cgcattgccn tg 225122DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 51aggaggaagg gcctgagtcn ca 225222DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 52agctggagga ggaagggccn ct 225322DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 53gcattgccac gaagcaggcn cc 225422DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 54attgccacga agcaggccan cg 225522DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 55agaaccggag gacaaagtan ga 225622DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 56tcaaccggtg gcgcattgcn gt 225722DNAHomo sapiensmodified_base(20)..(20)a, c, t, g, unknown or other 57gaagctggag gaggaagggn gc 225823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 58gccaaattgg acgaccctcg cgg 235923DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 59cgaggagacc cccgtttcgg tgg 236023DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 60cccgccgccg ccgtggctcg agg 236123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 61tgagctctac gagatccaca agg 236223DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 62ctcaaaattc ataccggttg tgg 236323DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 63cgttaaacaa caaccggact tgg 236423DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 64ttcaccccgc ggcgctgaat ggg 236523DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 65accactacca gtccgtccac agg 236623DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 66cactgcttaa gcctcgctcg agg 236723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 67tcaccagcaa tattcgctcg agg 236823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 68caccagcaat attccgctcg agg 236923DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 69tagcaacaga catacgctcg agg 237023DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 70gggcagtagt aatacgctcg agg 237123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 71ccaattccca tacattattg tac 237233DNAHomo sapiens 72ggacatcgat gtcacctcca atgactaggg tgg 337333DNAHomo sapiens 73cattggaggt gacatcgatg tcctccccat tgg 337433DNAHomo sapiens 74ggaagggcct gagtccgagc agaagaagaa ggg 337533DNAHomo sapiens 75ggtggcgaga ggggccgaga ttgggtgttc agg 337633DNAHomo sapiens 76atgcaggagg gtggcgagag gggccgagat tgg 337723DNAHomo sapiens 77gtcacctcca atgactaggg tgg 237823DNAHomo sapiens 78gacatcgatg tcctccccat tgg 237923DNAHomo sapiens 79gagtccgagc agaagaagaa ggg 238023DNAHomo sapiens 80gcgccaccgg ttgatgtgat ggg 238123DNAHomo sapiens 81ggggcacaga tgagaaactc agg 238223DNAHomo sapiens 82gtacaaacgg cagaagctgg agg 238323DNAHomo sapiens 83ggcagaagct ggaggaggaa ggg 238423DNAHomo sapiens 84ggagcccttc ttcttctgct cgg 238523DNAHomo sapiens 85gggcaaccac aaacccacga ggg 238623DNAHomo sapiens 86gctcccatca catcaaccgg tgg 238723DNAHomo sapiens 87gtggcgcatt gccacgaagc agg 238823DNAHomo sapiens 88ggcagagtgc tgcttgctgc tgg 238923DNAHomo sapiens 89gcccctgcgt gggcccaagc tgg 239023DNAHomo sapiens 90gagtggccag agtccagctt ggg 239123DNAHomo sapiens 91ggcctcccca aagcctggcc agg 239223DNAHomo sapiens 92ggggccgaga ttgggtgttc agg 239323DNAHomo sapiens 93gtggcgagag gggccgagat tgg 239423DNAHomo sapiens 94gagtgccgcc gaggcggggc ggg 239523DNAHomo sapiens 95ggagtgccgc cgaggcgggg cgg 239623DNAHomo sapiens 96ggagaggagt gccgccgagg cgg 239721DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 97aaaaccaccc ttctctctgg c 219821DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 98ggagattgga gacacggaga g 219920DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 99ccatcccctt ctgtgaatgt 2010020DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 100ggagattgga gacacggaga 2010120DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 101ctggaaagcc aatgcctgac 2010220DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 102ggcagcaaac tccttgtcct 2010320DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 103aagcaccgac tcggtgccac 2010420DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 104tcacctccaa tgactagggg 2010522DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 105caagttgata acggactagc ct 2210623DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 106agtccgagca gaagaagaag ttt 2310725DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 107tttcaagttg ataacggact agcct 2510820DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 108aaacagcaga ttcgcctgga 2010920DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 109tcatccgctc gatgaagctc 2011020DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 110tccaaaatca agtggggcga 2011120DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 111tgatgaccct tttggctccc 2011245DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 112gaggaattct ttttttgtty gaatatgttg gaggtttttt ggaag
4511342DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 113gagaagctta aataaaaaac racaatactc aacccaacaa cc 4211417DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 114caggaaacag ctatgac 1711539DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 115gcctctagag gtacctgagg gcctatttcc catgattcc 39116133DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer"modified_base(92)..(111)a, c, t, g, unknown or other 116acctctagaa aaaaagcacc gactcggtgc cactttttca agttgataac ggactagcct 60tattttaact tgctatttct agctctaaaa cnnnnnnnnn nnnnnnnnnn nggtgtttcg 120tcctttccac aag 133117133DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer"modified_base(92)..(111)a, c, t, g, unknown or other 117acctctagaa aaaaagcacc gactcggtgc cactttttca agttgataac ggactagcct 60tatattaact tgctatttct agctctaata cnnnnnnnnn nnnnnnnnnn nggtgtttcg 120tcctttccac aag 133118153DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer"modified_base(112)..(131)a, c, t, g, unknown or other 118acctctagaa aaaaagcacc gactcggtgc cactttttca agttgataac ggactagcct 60tattttaact tgctatgctg ttttgtttcc aaaacagcat agctctaaaa cnnnnnnnnn 120nnnnnnnnnn nggtgtttcg tcctttccac aag 153119153DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer"modified_base(112)..(131)a, c, t, g, unknown or other 119acctctagaa aaaaagcacc gactcggtgc cactttttca agttgataac ggactagcct 60tatattaact tgctatgctg tattgtttcc aatacagcat agctctaata cnnnnnnnnn 120nnnnnnnnnn nggtgtttcg tcctttccac aag 153120335DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 120gagggcctat ttcccatgat tccttcatat ttgcatatac gatacaaggc tgttagagag 60ataattggaa ttaatttgac tgtaaacaca aagatattag tacaaaatac gtgacgtaga 120aagtaataat ttcttgggta gtttgcagtt ttaaaattat gttttaaaat ggactatcat 180atgcttaccg taacttgaaa gtatttcgat ttcttggctt tatatatctt gtggaaagga 240cgaaacaccg gaaccattca aaacagcata gcaagttaaa ataaggctag tccgttatca 300acttgaaaaa gtggcaccga gtcggtgctt ttttt 335121360DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(288)..(317)a, c, t, g, unknown or other 121gagggcctat ttcccatgat tccttcatat ttgcatatac gatacaaggc tgttagagag 60ataattggaa ttaatttgac tgtaaacaca aagatattag tacaaaatac gtgacgtaga 120aagtaataat ttcttgggta gtttgcagtt ttaaaattat gttttaaaat ggactatcat 180atgcttaccg taacttgaaa gtatttcgat ttcttggctt tatatatctt gtggaaagga 240cgaaacaccg ggttttagag ctatgctgtt ttgaatggtc ccaaaacnnn nnnnnnnnnn 300nnnnnnnnnn nnnnnnngtt ttagagctat gctgttttga atggtcccaa aacttttttt 360122318DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(250)..(269)a, c, t, g, unknown or other 122gagggcctat ttcccatgat tccttcatat ttgcatatac gatacaaggc tgttagagag 60ataattggaa ttaatttgac tgtaaacaca aagatattag tacaaaatac gtgacgtaga 120aagtaataat ttcttgggta gtttgcagtt ttaaaattat gttttaaaat ggactatcat 180atgcttaccg taacttgaaa gtatttcgat ttcttggctt tatatatctt gtggaaagga 240cgaaacaccn nnnnnnnnnn nnnnnnnnng ttttagagct agaaatagca agttaaaata 300aggctagtcc gttttttt 318123325DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(250)..(269)a, c, t, g, unknown or other 123gagggcctat ttcccatgat tccttcatat ttgcatatac gatacaaggc tgttagagag 60ataattggaa ttaatttgac tgtaaacaca aagatattag tacaaaatac gtgacgtaga 120aagtaataat ttcttgggta gtttgcagtt ttaaaattat gttttaaaat ggactatcat 180atgcttaccg taacttgaaa gtatttcgat ttcttggctt tatatatctt gtggaaagga 240cgaaacaccn nnnnnnnnnn nnnnnnnnng ttttagagct agaaatagca agttaaaata 300aggctagtcc gttatcattt ttttt 325124337DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(250)..(269)a, c, t, g, unknown or other 124gagggcctat ttcccatgat tccttcatat ttgcatatac gatacaaggc tgttagagag 60ataattggaa ttaatttgac tgtaaacaca aagatattag tacaaaatac gtgacgtaga 120aagtaataat ttcttgggta gtttgcagtt ttaaaattat gttttaaaat ggactatcat 180atgcttaccg taacttgaaa gtatttcgat ttcttggctt tatatatctt gtggaaagga 240cgaaacaccn nnnnnnnnnn nnnnnnnnng ttttagagct agaaatagca agttaaaata 300aggctagtcc gttatcaact tgaaaaagtg ttttttt 337125352DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide"modified_base(250)..(269)a, c, t, g, unknown or other 125gagggcctat ttcccatgat tccttcatat ttgcatatac gatacaaggc tgttagagag 60ataattggaa ttaatttgac tgtaaacaca aagatattag tacaaaatac gtgacgtaga 120aagtaataat ttcttgggta gtttgcagtt ttaaaattat gttttaaaat ggactatcat 180atgcttaccg taacttgaaa gtatttcgat ttcttggctt tatatatctt gtggaaagga 240cgaaacaccn nnnnnnnnnn nnnnnnnnng ttttagagct agaaatagca agttaaaata 300aggctagtcc gttatcaact tgaaaaagtg gcaccgagtc ggtgcttttt tt 3521265101DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 126cgttacataa cttacggtaa atggcccgcc tggctgaccg cccaacgacc cccgcccatt 60gacgtcaata atgacgtatg ttcccatagt aacgccaata gggactttcc attgacgtca 120atgggtggag tatttacggt aaactgccca cttggcagta catcaagtgt atcatatgcc 180aagtacgccc cctattgacg tcaatgacgg taaatggccc gcctggcatt atgcccagta 240catgacctta tgggactttc ctacttggca gtacatctac gtattagtca tcgctattac 300catggtcgag gtgagcccca cgttctgctt cactctcccc atctcccccc cctccccacc 360cccaattttg tatttattta ttttttaatt attttgtgca gcgatggggg cggggggggg 420gggggggcgc gcgccaggcg gggcggggcg gggcgagggg cggggcgggg cgaggcggag 480aggtgcggcg gcagccaatc agagcggcgc gctccgaaag tttcctttta tggcgaggcg 540gcggcggcgg cggccctata aaaagcgaag cgcgcggcgg gcgggagtcg ctgcgacgct 600gccttcgccc cgtgccccgc tccgccgccg cctcgcgccg cccgccccgg ctctgactga 660ccgcgttact cccacaggtg agcgggcggg acggcccttc tcctccgggc tgtaattagc 720tgagcaagag gtaagggttt aagggatggt tggttggtgg ggtattaatg tttaattacc 780tggagcacct gcctgaaatc actttttttc aggttggacc ggtgccacca tggactataa 840ggaccacgac ggagactaca aggatcatga tattgattac aaagacgatg acgataagat 900ggccccaaag aagaagcgga aggtcggtat ccacggagtc ccagcagccg acaagaagta 960cagcatcggc ctggacatcg gcaccaactc tgtgggctgg gccgtgatca ccgacgagta 1020caaggtgccc agcaagaaat tcaaggtgct gggcaacacc gaccggcaca gcatcaagaa 1080gaacctgatc ggagccctgc tgttcgacag cggcgaaaca gccgaggcca cccggctgaa 1140gagaaccgcc agaagaagat acaccagacg gaagaaccgg atctgctatc tgcaagagat 1200cttcagcaac gagatggcca aggtggacga cagcttcttc cacagactgg aagagtcctt 1260cctggtggaa gaggataaga agcacgagcg gcaccccatc ttcggcaaca tcgtggacga 1320ggtggcctac cacgagaagt accccaccat ctaccacctg agaaagaaac tggtggacag 1380caccgacaag gccgacctgc ggctgatcta tctggccctg gcccacatga tcaagttccg 1440gggccacttc ctgatcgagg gcgacctgaa ccccgacaac agcgacgtgg acaagctgtt 1500catccagctg gtgcagacct acaaccagct gttcgaggaa aaccccatca acgccagcgg 1560cgtggacgcc aaggccatcc tgtctgccag actgagcaag agcagacggc tggaaaatct 1620gatcgcccag ctgcccggcg agaagaagaa tggcctgttc ggcaacctga ttgccctgag 1680cctgggcctg acccccaact tcaagagcaa cttcgacctg gccgaggatg ccaaactgca 1740gctgagcaag gacacctacg acgacgacct ggacaacctg ctggcccaga tcggcgacca 1800gtacgccgac ctgtttctgg ccgccaagaa cctgtccgac gccatcctgc tgagcgacat 1860cctgagagtg aacaccgaga tcaccaaggc ccccctgagc gcctctatga tcaagagata 1920cgacgagcac caccaggacc tgaccctgct gaaagctctc gtgcggcagc agctgcctga 1980gaagtacaaa gagattttct tcgaccagag caagaacggc tacgccggct acattgacgg 2040cggagccagc caggaagagt tctacaagtt catcaagccc atcctggaaa agatggacgg 2100caccgaggaa ctgctcgtga agctgaacag agaggacctg ctgcggaagc agcggacctt 2160cgacaacggc agcatccccc accagatcca cctgggagag ctgcacgcca ttctgcggcg 2220gcaggaagat ttttacccat tcctgaagga caaccgggaa aagatcgaga agatcctgac 2280cttccgcatc ccctactacg tgggccctct ggccagggga aacagcagat tcgcctggat 2340gaccagaaag agcgaggaaa ccatcacccc ctggaacttc gaggaagtgg tggacaaggg 2400cgcttccgcc cagagcttca tcgagcggat gaccaacttc gataagaacc tgcccaacga 2460gaaggtgctg cccaagcaca gcctgctgta cgagtacttc accgtgtata acgagctgac 2520caaagtgaaa tacgtgaccg agggaatgag aaagcccgcc ttcctgagcg gcgagcagaa 2580aaaggccatc gtggacctgc tgttcaagac caaccggaaa gtgaccgtga agcagctgaa 2640agaggactac ttcaagaaaa tcgagtgctt cgactccgtg gaaatctccg gcgtggaaga 2700tcggttcaac gcctccctgg gcacatacca cgatctgctg aaaattatca aggacaagga 2760cttcctggac aatgaggaaa acgaggacat tctggaagat atcgtgctga ccctgacact 2820gtttgaggac agagagatga tcgaggaacg gctgaaaacc tatgcccacc tgttcgacga 2880caaagtgatg aagcagctga agcggcggag atacaccggc tggggcaggc tgagccggaa 2940gctgatcaac ggcatccggg acaagcagtc cggcaagaca atcctggatt tcctgaagtc 3000cgacggcttc gccaacagaa acttcatgca gctgatccac gacgacagcc tgacctttaa 3060agaggacatc cagaaagccc aggtgtccgg ccagggcgat agcctgcacg agcacattgc 3120caatctggcc ggcagccccg ccattaagaa gggcatcctg cagacagtga aggtggtgga 3180cgagctcgtg aaagtgatgg gccggcacaa gcccgagaac atcgtgatcg aaatggccag 3240agagaaccag accacccaga agggacagaa gaacagccgc gagagaatga agcggatcga 3300agagggcatc aaagagctgg gcagccagat cctgaaagaa caccccgtgg aaaacaccca 3360gctgcagaac gagaagctgt acctgtacta cctgcagaat gggcgggata tgtacgtgga 3420ccaggaactg gacatcaacc ggctgtccga ctacgatgtg gaccatatcg tgcctcagag 3480ctttctgaag gacgactcca tcgacaacaa ggtgctgacc agaagcgaca agaaccgggg 3540caagagcgac aacgtgccct ccgaagaggt cgtgaagaag atgaagaact actggcggca 3600gctgctgaac gccaagctga ttacccagag aaagttcgac aatctgacca aggccgagag 3660aggcggcctg agcgaactgg ataaggccgg cttcatcaag agacagctgg tggaaacccg 3720gcagatcaca aagcacgtgg cacagatcct ggactcccgg atgaacacta agtacgacga 3780gaatgacaag ctgatccggg aagtgaaagt gatcaccctg aagtccaagc tggtgtccga 3840tttccggaag gatttccagt tttacaaagt gcgcgagatc aacaactacc accacgccca 3900cgacgcctac ctgaacgccg tcgtgggaac cgccctgatc aaaaagtacc ctaagctgga 3960aagcgagttc gtgtacggcg actacaaggt gtacgacgtg cggaagatga tcgccaagag 4020cgagcaggaa atcggcaagg ctaccgccaa gtacttcttc tacagcaaca tcatgaactt 4080tttcaagacc gagattaccc tggccaacgg cgagatccgg aagcggcctc tgatcgagac 4140aaacggcgaa accggggaga tcgtgtggga taagggccgg gattttgcca ccgtgcggaa 4200agtgctgagc atgccccaag tgaatatcgt gaaaaagacc gaggtgcaga caggcggctt 4260cagcaaagag tctatcctgc ccaagaggaa cagcgataag ctgatcgcca gaaagaagga 4320ctgggaccct aagaagtacg gcggcttcga cagccccacc gtggcctatt ctgtgctggt 4380ggtggccaaa gtggaaaagg gcaagtccaa gaaactgaag agtgtgaaag agctgctggg 4440gatcaccatc atggaaagaa gcagcttcga gaagaatccc atcgactttc tggaagccaa 4500gggctacaaa gaagtgaaaa aggacctgat catcaagctg cctaagtact ccctgttcga 4560gctggaaaac ggccggaaga gaatgctggc ctctgccggc gaactgcaga agggaaacga 4620actggccctg ccctccaaat atgtgaactt cctgtacctg gccagccact atgagaagct 4680gaagggctcc cccgaggata atgagcagaa acagctgttt gtggaacagc acaagcacta 4740cctggacgag atcatcgagc agatcagcga gttctccaag agagtgatcc tggccgacgc 4800taatctggac aaagtgctgt ccgcctacaa caagcaccgg gataagccca tcagagagca 4860ggccgagaat atcatccacc tgtttaccct gaccaatctg ggagcccctg ccgccttcaa 4920gtactttgac accaccatcg accggaagag gtacaccagc accaaagagg tgctggacgc 4980caccctgatc caccagagca tcaccggcct gtacgagaca cggatcgacc tgtctcagct 5040gggaggcgac tttctttttc ttagcttgac cagctttctt agtagcagca ggacgcttta 5100a 5101127123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 127ccaatgggga ggacatcgat gtcacctcca atgactaggg tgggcaacca caaacccacg 60agggcagagt gctgcttgct gctggccagg cccctgcgtg ggcccaagct ggactctggc 120cac 123128121DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 128cgagcagaag aagaagggct cccatcacat caaccggtgg cgcattgcca cgaagcaggc 60caatggggag gacatcgatg tcacctccaa tgactagggt gggcaaccac aaacccacga 120g 121129128DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 129ggaggacaaa gtacaaacgg cagaagctgg aggaggaagg gcctgagtcc gagcagaaga 60agaagggctc ccatcacatc aaccggtggc gcattgccac gaagcaggcc aatggggagg 120acatcgat 128130130DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 130agaagctgga ggaggaaggg cctgagtccg agcagaagaa gaagggctcc catcacatca 60accggtggcg cattgccacg aagcaggcca atggggagga catcgatgtc acctccaatg 120actagggtgg 130131125DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 131cctcagtctt cccatcaggc tctcagctca gcctgagtgt tgaggcccca gtggctgctc 60tgggggcctc ctgagtttct catctgtgcc cctccctccc tggcccaggt gaaggtgtgg 120ttcca 125132129DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 132tcatctgtgc ccctccctcc ctggcccagg tgaaggtgtg gttccagaac cggaggacaa 60agtacaaacg gcagaagctg gaggaggaag ggcctgagtc cgagcagaag aagaagggct 120cccatcaca 129133129DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 133ctccaatgac tagggtgggc aaccacaaac ccacgagggc agagtgctgc ttgctgctgg 60ccaggcccct gcgtgggccc aagctggact ctggccactc cctggccagg ctttggggag 120gcctggagt 129134127DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 134ctgcttgctg ctggccaggc ccctgcgtgg gcccaagctg gactctggcc actccctggc 60caggctttgg ggaggcctgg agtcatggcc ccacagggct tgaagcccgg ggccgccatt 120gacagag 12713525DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic primer" 135gaaattaata cgactcacta taggg 25136126DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 136aaaaaagcac cgactcggtg ccactttttc aagttgataa cggactagcc ttattttaac 60ttgctatttc tagctctaaa acaacgacga gcgtgacacc accctatagt gagtcgtatt 120aatttc 126137126DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 137aaaaaagcac cgactcggtg ccactttttc aagttgataa cggactagcc ttattttaac 60ttgctatttc tagctctaaa acgcaacaat taatagactg gacctatagt gagtcgtatt 120aatttc 126138102DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 138gttttagagc tatgctgttt tgaatggtcc caaaacggaa gggcctgagt ccgagcagaa 60gaagaagttt tagagctatg ctgttttgaa tggtcccaaa ac 102139100DNAHomo sapiens 139cggaggacaa agtacaaacg gcagaagctg gaggaggaag ggcctgagtc cgagcagaag 60aagaagggct cccatcacat caaccggtgg cgcattgcca 10014050DNAHomo sapiens 140agctggagga ggaagggcct gagtccgagc agaagaagaa gggctcccac 5014130RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 141gaguccgagc agaagaagaa guuuuagagc 3014249DNAHomo sapiens 142agctggagga ggaagggcct gagtccgagc agaagagaag ggctcccat 4914353DNAHomo sapiens 143ctggaggagg aagggcctga gtccgagcag aagaagaagg gctcccatca cat 5314452DNAHomo sapiens 144ctggaggagg aagggcctga gtccgagcag aagagaaggg ctcccatcac at 5214554DNAHomo sapiens 145ctggaggagg aagggcctga gtccgagcag aagaaagaag ggctcccatc acat 5414650DNAHomo sapiens 146ctggaggagg aagggcctga gtccgagcag aagaagggct cccatcacat 5014747DNAHomo sapiens 147ctggaggagg aagggcctga gcccgagcag aagggctccc atcacat 4714823DNAHomo sapiens 148gtcacctcca atgactaggg tgg 2314923DNAHomo sapiens 149gcgccaccgg ttgatgtgat ggg 2315099RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(1)..(20)a, c, u, g, unknown or other 150nnnnnnnnnn nnnnnnnnnn guuuuagagc uagaaauagc aaguuaaaau aaggcuaguc 60cguuaucaac uugaaaaagu ggcaccgagu cggugcuuu 9915164DNAHomo sapiens 151caagaggctt gagtaggaga ggagtgccgc cgaggcgggg cggggcgggg cgtggagctg 60ggct 6415220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 152gucaccucca augacuaggg 2015320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 153gucaccucca augacuaaga 2015420RNAArtificial Sequencesource/note="Description of Artificial
Sequence Synthetic oligonucleotide" 154gucaccucca augauuagga 2015520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 155gucaccucca gugacuagga 2015620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 156gucacuucca augacuagga 2015720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 157aucaccucca augacuagga 2015820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 158gucaccucca augaccaagg 2015920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 159gucaccucca auaacuaagg 2016020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 160gucaccucua augacuaagg 2016120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 161gucgccucca augacuaagg 2016220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 162gucaccucca auggccaggg 2016320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 163gucaccucca gugaccaggg 2016420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 164gucaccccca augaccaggg 2016520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 165gccaccucca augaccaggg 2016620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 166gucaccucca augaccaaga 2016720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 167gucaccucca auggcuggga 2016820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 168gucaccucca acgaccagga 2016920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 169gucaccuccg augauuagga 2017020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 170gucaccucca auggccaagg 2017120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 171gucaccucca acgauuaagg 2017220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 172gucaccuccg auggcuaagg 2017320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 173gucaccuuca auaacuaagg 2017420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 174gucaccucca auggccaaga 2017520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 175gucaccucca guggcuggga 2017620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 176gucaccuuca acgaccagga 2017720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 177gucaucuccg augauuagga 2017820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 178gccaccuuca auggcuagga 2017920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 179gucaccucca augacuagaa 2018020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 180gucaccucca augacugagg 2018120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 181gucaccucca augaucaggg 2018220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 182gucaccucca auagcuaggg 2018320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 183gucaccucca gcgacuaggg 2018420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 184gucaccucug augacuaggg 2018520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 185gucacccuca augacuaggg 2018620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 186gucauuucca augacuaggg 2018720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 187guugccucca augacuaggg 2018820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 188accaccucca augacuaggg 2018920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 189gucaccucca augacugaag 2019020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 190gucaccucca auggucaggg 2019120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 191gucaccucca gcaacuaggg 2019220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 192gucaccuuug augacuaggg 2019320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 193gucauuccca augacuaggg 2019420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 194gcugccucca augacuaggg 2019520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 195gucaccucca augaccgaaa 2019620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 196gucaccucca auagucgggg 2019720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 197gucaccuccg gcagcuaggg 2019820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 198gucacccuug gugacuaggg 2019920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 199gucguucuca augacuaggg 2020020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 200acugucucca augacuaggg 2020120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 201gcgccaccgg uugaugugau 2020220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 202gcgccaccgg uugauguaac 2020320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 203gcgccaccgg uugacgugac 2020420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 204gcgccaccgg cugaugugac 2020520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 205gcgccgccgg uugaugugac 2020620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 206acgccaccgg uugaugugac 2020720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 207gcgccaccgg uugauauaau 2020820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 208gcgccaccgg uuaauguaau 2020920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 209gcgccaccag uugauguaau 2021020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 210gcgucaccgg uugauguaau 2021120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 211gcgccaccgg uugguaugau 2021220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 212gcgccaccgg cugauaugau 2021320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 213gcgccaucgg uugauaugau 2021420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 214gugccaccgg uugauaugau 2021520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 215gcgccaccgg uugauauaac 2021620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 216gcgccaccgg uuggugcgac 2021720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 217gcgccaccgg ucgauaugac 2021820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 218gcgccaccga uugacgugac 2021920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 219gcgccaccgg uugguauaau 2022020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 220gcgccaccgg ucgacguaau 2022120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 221gcgccaccga uugguguaau 2022220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 222gcgccacugg uuaauguaau 2022320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 223gcgccaccgg uugguauaac 2022420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 224gcgccaccgg cuggugcgac 2022520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 225gcgccacugg ucgauaugac 2022620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 226gcgcuaccga uugacgugac 2022720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 227gugccacugg uuggugugac 2022820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 228gcgccaccgg uugauguggc 2022920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 229gcgccaccgg uugaugcaau 2023020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 230gcgccaccgg uugacaugau 2023120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 231gcgccaccgg uuagugugau 2023220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 232gcgccaccgg ccgaugugau 2023320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 233gcgccaccaa uugaugugau 2023420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 234gcgccauugg uugaugugau 2023520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 235gcgcugccgg uugaugugau 2023620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 236gcaucaccgg uugaugugau 2023720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 237augccaccgg uugaugugau 2023820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 238gcgccaccgg uugaugcagu 2023920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 239gcgccaccgg uuggcaugau 2024020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 240gcgccaccgg ccaaugugau 2024120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 241gcgccacuaa uugaugugau 2024220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 242gcgcugucgg uugaugugau 2024320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 243guaucaccgg uugaugugau 2024420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 244gcgccaccgg uugauacagc 2024520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 245gcgccaccgg uuagcacgau 2024620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 246gcgccaccga ccagugugau 2024720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 247gcgccauuaa cugaugugau 2024820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic
oligonucleotide" 248gcguuguugg uugaugugau 2024920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 249auauuaccgg uugaugugau 2025020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 250gucaccucca augacuaggg 2025120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 251gucaccugga augacuaggg 2025220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 252gucacgacca augacuaggg 2025320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 253gucugcucca augacuaggg 2025420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 254gagaccucca augacuaggg 2025520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 255gucaccagga augacuaggg 2025620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 256gucuggucca augacuaggg 2025720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 257gaguccucca augacuaggg 2025820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 258gucacgagga augacuaggg 2025920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 259gucuggacca augacuaggg 2026020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 260gagugcucca augacuaggg 2026120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 261gucugcugga augacuaggg 2026220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 262gagacgacca augacuaggg 2026320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 263gagaccugga augacuaggg 2026440DNAHomo sapiens 264ggacatcgat gtcacctcca atgactaggg tgggcaacca 4026520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 265gucaccucca augacuaggg 2026620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(20)..(20)a, c, u, g, unknown or other 266gucaccucca augacuaggn 2026720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(19)..(19)a, c, u, g, unknown or other 267gucaccucca augacuagng 2026820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(18)..(18)a, c, u, g, unknown or other 268gucaccucca augacuangg 2026920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(17)..(17)a, c, u, g, unknown or other 269gucaccucca augacunggg 2027020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(16)..(16)a, c, u, g, unknown or other 270gucaccucca augacnaggg 2027120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(15)..(15)a, c, u, g, unknown or other 271gucaccucca auganuaggg 2027220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(14)..(14)a, c, u, g, unknown or other 272gucaccucca augncuaggg 2027320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(13)..(13)a, c, u, g, unknown or other 273gucaccucca aunacuaggg 2027420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(12)..(12)a, c, u, g, unknown or other 274gucaccucca angacuaggg 2027520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(11)..(11)a, c, u, g, unknown or other 275gucaccucca nugacuaggg 2027620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(10)..(10)a, c, u, g, unknown or other 276gucaccuccn augacuaggg 2027720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(9)..(9)a, c, u, g, unknown or other 277gucaccucna augacuaggg 2027820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(8)..(8)a, c, u, g, unknown or other 278gucaccunca augacuaggg 2027920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(7)..(7)a, c, u, g, unknown or other 279gucaccncca augacuaggg 2028020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(6)..(6)a, c, u, g, unknown or other 280gucacnucca augacuaggg 2028120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(5)..(5)a, c, u, g, unknown or other 281gucancucca augacuaggg 2028220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(4)..(4)a, c, u, g, unknown or other 282gucnccucca augacuaggg 2028320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(3)..(3)a, c, u, g, unknown or other 283gunaccucca augacuaggg 2028420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(2)..(2)a, c, u, g, unknown or other 284gncaccucca augacuaggg 2028520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 285gucaccucca augacuaggg 2028620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 286gacaucgaug uccuccccau 2028720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 287gcgccaccgg uugaugugau 2028820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 288gucaccucca augacuaggg 2028920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 289gucaccucca augacuagaa 2029020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 290gucaccucca augacugagg 2029120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 291gucaccucca augaucaggg 2029220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 292gucaccucca auagcuaggg 2029320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 293gucaccucca gcgacuaggg 2029420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 294gucaccucug augacuaggg 2029520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 295gucacccuca augacuaggg 2029620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 296gucauuucca augacuaggg 2029720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 297guugccucca augacuaggg 2029820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 298accaccucca augacuaggg 2029920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 299gucaccucca augacugaag 2030020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 300gucaccucca auggucaggg 2030120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 301gucaccucca gcaacuaggg 2030220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 302gucaccuuug augacuaggg 2030320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 303gucauuccca augacuaggg 2030420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 304gcugccucca augacuaggg 2030520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 305gucaccucca augaccgaaa 2030620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 306gucaccucca auagucgggg 2030720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 307gucaccuccg gcagcuaggg 2030820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 308gucacccuug gugacuaggg 2030920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 309gucguucuca augacuaggg 2031020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 310acugucucca augacuaggg 2031120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 311gcgccaccgg uugaugugau 2031220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 312gcgccaccgg uugauguggc 2031320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 313gcgccaccgg uugaugcaau 2031420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 314gcgccaccgg uugacaugau 2031520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 315gcgccaccgg uuagugugau 2031620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 316gcgccaccgg ccgaugugau 2031720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 317gcgccaccaa uugaugugau 2031820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 318gcgccauugg uugaugugau 2031920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 319gcgcugccgg uugaugugau 2032020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 320gcaucaccgg uugaugugau 2032120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 321augccaccgg uugaugugau 2032220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 322gcgccaccgg uugaugcagu 2032320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 323gcgccaccgg uuggcaugau 2032420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 324gcgccaccgg ccaaugugau 2032520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 325gcgccacuaa uugaugugau 2032620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 326gcgcugucgg uugaugugau 2032720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 327guaucaccgg uugaugugau 2032820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 328gcgccaccgg uugauacagc 2032920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 329gcgccaccgg uuagcacgau 2033020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 330gcgccaccga ggagugugau 2033120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 331gcgccauuaa gugaugugau 2033220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 332gcguuguugg uugaugugau 2033320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 333auauuaccgg uugaugugau 2033420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 334gucaccucca augacuaggg 2033520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 335gucaccucca augacuaaga 2033620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 336gucaccucca augauuagga 2033720RNAArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 337gucaccucca gugacuagga 2033820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 338gucacuucca augacuagga 2033920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 339aucaccucca augacuagga 2034020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 340gucaccucca augacuaggg 2034120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 341gucaccucca augaccaaga 2034220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 342gucaccucca auggcuggga 2034320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 343gucaccucca acgaccagga 2034420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 344gucaccuccg augauuagga 2034520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 345gucaccucca auggccaagg 2034620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 346gucaccucca acgauuaagg 2034720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 347gucaccuccg auggcuaagg 2034820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 348gucaccuuca auaacuaagg 2034920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 349gucaccucca auggccaaga 2035020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 350gucaccucca guggcuggga 2035120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 351gucaccuuca acgaccagga 2035220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 352gucaucuccg augauuagga 2035320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 353gccaccuuca auggcuagga 2035420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 354gcgccaccgg uugaugugau 2035520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 355gcgccaccgg uugauguaac 2035620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 356gcgccaccgg uugacgugac 2035720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 357gcgccaccgg cugaugugac 2035820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 358gcgccgccgg uugaugugac 2035920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 359acgccaccgg uugaugugac 2036020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 360gcgccaccgg uugaugugau 2036120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 361gcgccaccgg uugauauaac 2036220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 362gcgccaccgg uuggugcgac 2036320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 363gcgccaccgg ucgauaugac 2036420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 364gcgccaccga uugacgugac 2036520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 365gcgccaccgg uugguauaau 2036620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 366gcgccaccgg ucgacguaau 2036720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 367gcgccaccga uugguguaau 2036820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 368gcgccacugg uuaauguaau 2036920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 369gcgccaccgg uugguauaac 2037020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 370gcgccaccgg cuggugcgac 2037120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 371gcgccacugg ucgauaugac 2037220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 372gcgcuaccga uugacgugac 2037320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 373gugccacugg uuggugugac 2037423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 374gaguccgagc agaagaagaa ggg 2337523DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 375gauuccuacc agaagaagaa tgg 2337623RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 376aaguccgagg agaggaagaa agg 2337723RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 377aaguccgaga agaagcagaa aag 2337823RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 378gaguuccaga agaagaagaa gag 2337923RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 379gagucugaac ggaagaagaa aag 2338023RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 380gaguuugagu agaagaagaa gag 2338123RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 381gagucccaga agaagaaaaa aag 2338223DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 382gacuccgagc agcagaagga tgg 2338323RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 383gagucagagc agaacuagaa ggg 2338423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 384gaggccgagc agaagaaaga cgg 2338523RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 385gagucagacc aggagaagaa gag 2338623DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 386gaguccaaga agaauaagaa tag 2338723RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 387gaguaagaga agaagaagaa ggg 2338823RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 388gaguaggagg agaagaagaa agg 2338923DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 389aagucugagc acaagaagaa tgg 2339023RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 390gagcccgagc agaaggagga ggg 2339123RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 391gaguuagagc agaagaagaa agg 2339223RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 392gagucagaac agaagaacaa cag 2339323DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 393acgucugagc agaagaagaa tgg 2339423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 394gaguacuaga agaagaagaa aag 2339523RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 395aagucugagc agaagaagca cag 2339623RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 396gagucccagc agaggaagca gag 2339723RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 397gagucugggc aggagaagaa gag 2339823RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 398gaguccuagc aggagaagaa gag 2339923RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 399gaggaggagc agaagaagaa aag 2340023RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 400gaguccgaga aaaugaagaa gag 2340123RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 401gaggcccagc agaggaagaa gag 2340223RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 402gaauccaagc agaagaagag aag 2340323DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 403gacuccuagc aaaagaagaa tgg 2340423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 404gagucccagc aaaagaagaa aag 2340523RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 405gagucuaagc agaagaagaa gag 2340623RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 406aagucagagg agaagaagaa ggg 2340723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 407gaguccaagc agaagaagga tag 2340823RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 408gagacugaga agaagaagaa agg 2340923RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 409gagucccagg agaagaagag agg 2341023RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 410gagucccagg agaagaaaaa cag 2341123RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 411gagagcaagc agaagaagaa aag 2341223RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 412gaguccaagc auaagaaaaa cag 2341323RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 413gaguccaagc aguagaggaa ggg 2341423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 414gaguccggga aggagaagaa agg 2341520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 415gcgccaccgg uugaugugau 2041623RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 416gggccauggg uugaugugau gag 2341723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 417gtcacctcca atgactaggg tgg 2341823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 418gcuacctcca gtgactaggg agg 2341923DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 419gtgacctcca atgcctagag ggg 2342023DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 420gtcacctcca ctucctaggg cag 2342123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 421utcacctcca aaaactaggg aag 2342223DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 422gtcaactcca atggctuggg agg 2342323DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 423gtcacctuaa atgactuggg aag 2342423DNAArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 424gtcacutcca aggactagag aag 2342523DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 425gtcacctcca ggguctaggg cag 2342623DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 426gtcacctcca ctguataggg agg 2342723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 427gtcaactcca atgautagga cag 2342823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 428gtggtgtcac gctcgtcgtt tgg 2342923DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 429tccagtctat taattgttgc cgg 2343012DNAHomo sapiens 430tagcgggtaa gc 1243112DNAHomo sapiens 431tcggtcacat gt 1243212DNAHomo sapiens 432actccccgta gg 1243312DNAHomo sapiens 433actgcgtgtt aa 1243412DNAHomo sapiens 434acgtcgcctg at 1243512DNAHomo sapiens 435taggtcgacc ag 1243612DNAHomo sapiens 436ggcgttaatg at 1243712DNAHomo sapiens 437tgtcgcatgt ta 1243812DNAHomo sapiens 438atggaaacgc at 1243912DNAHomo sapiens 439gccgaattcc tc 1244012DNAHomo sapiens 440gcatggtacg ga 1244112DNAHomo sapiens 441cggtactctt ac 1244212DNAHomo sapiens 442gcctgtgccg ta 1244312DNAHomo sapiens 443tacggtaagt cg 1244412DNAHomo sapiens 444cacgaaatta cc 1244512DNAHomo sapiens 445aaccaagata cg 1244612DNAHomo sapiens 446gagtcgatac gc 1244712DNAHomo sapiens 447gtctcacgat cg 1244812DNAHomo sapiens 448tcgtcgggtg ca 1244912DNAHomo sapiens 449actccgtagt ga 1245012DNAHomo sapiens 450caggacgtcc gt 1245112DNAHomo sapiens 451tcgtatccct ac 1245212DNAHomo sapiens 452tttcaaggcc gg 1245312DNAHomo sapiens 453cgccggtgga at 1245412DNAHomo sapiens 454gaacccgtcc ta 1245512DNAHomo sapiens 455gattcatcag cg 1245612DNAHomo sapiens 456acaccggtct tc 1245712DNAHomo sapiens 457atcgtgccct aa 1245812DNAHomo sapiens 458gcgtcaatgt tc 1245912DNAHomo sapiens 459ctccgtatct cg 1246012DNAHomo sapiens 460ccgattcctt cg 1246112DNAHomo sapiens 461tgcgcctcca gt 1246212DNAHomo sapiens 462taacgtcgga gc 1246312DNAHomo sapiens 463aaggtcgccc at 1246412DNAHomo sapiens 464ctcggggact at 1246512DNAHomo sapiens 465ttcgagcgat tt 1246612DNAHomo sapiens 466tgagtcgtcg ag 1246712DNAHomo sapiens 467tttacgcaga gg 1246812DNAHomo sapiens 468aggaagtatc gc 1246912DNAHomo sapiens 469actcgatacc at 1247012DNAHomo sapiens 470cgctacatag ca 1247112DNAHomo sapiens 471ttcataaccg gc 1247212DNAHomo sapiens 472ccaaacggtt aa 1247312DNAHomo sapiens 473cgattccttc gt 1247412DNAHomo sapiens 474cgtcatgaat aa 1247512DNAHomo sapiens 475agtggcgatg ac 1247612DNAHomo sapiens 476cccctacggc ac 1247712DNAHomo sapiens 477gccaacccgc ac 1247812DNAHomo sapiens 478tgggacaccg gt 1247912DNAHomo sapiens 479ttgactgcgg cg 1248012DNAHomo sapiens 480actatgcgta gg 1248112DNAHomo sapiens 481tcacccaaag cg 1248212DNAHomo sapiens 482gcaggacgtc cg 1248312DNAHomo sapiens 483acaccgaaaa cg 1248412DNAHomo sapiens 484cggtgtattg ag 1248512DNAHomo sapiens 485cacgaggtat gc 1248612DNAHomo sapiens 486taaagcgacc cg 1248712DNAHomo sapiens 487cttagtcggc ca 1248812DNAHomo sapiens 488cgaaaacgtg gc 1248912DNAHomo sapiens 489cgtgccctga ac 1249012DNAHomo sapiens 490tttaccatcg aa 1249112DNAHomo sapiens 491cgtagccatg tt 1249212DNAHomo sapiens 492cccaaacggt ta 1249312DNAHomo sapiens 493gcgttatcag aa 1249412DNAHomo sapiens 494tcgatggtaa ac 1249512DNAHomo sapiens 495cgactttttg ca 1249612DNAHomo sapiens 496tcgacgactc ac 1249712DNAHomo sapiens 497acgcgtcaga ta 1249812DNAHomo sapiens 498cgtacggcac ag 1249912DNAHomo sapiens 499ctatgccgtg ca 1250012DNAHomo sapiens 500cgcgtcagat at 1250112DNAHomo sapiens 501aagatcggta gc 1250212DNAHomo sapiens 502cttcgcaagg ag 1250312DNAHomo sapiens 503gtcgtggact ac 1250412DNAHomo sapiens 504ggtcgtcatc aa 1250512DNAHomo sapiens 505gttaacagcg tg 1250612DNAHomo sapiens 506tagctaaccg tt 1250712DNAHomo sapiens 507agtaaaggcg ct 1250812DNAHomo sapiens 508ggtaatttcg tg 1250920DNAMus musculus 509gactgattca gctagaccgg 2051020DNAMus musculus 510ggcacaaacg cgttcaatac 2051120DNAMus musculus 511aacgcgttca ataccggtct 2051220DNARattus norvegicus 512gctaggagga tctcgggcgg 2051320DNARattus norvegicus 513gttccagccc gcgactaagc 2051420DNARattus norvegicus 514gtgagggggt tgcaacgagg 2051520DNARattus norvegicus 515accgcaactc ttggcgcgtc 2051620DNARattus norvegicus 516accagatgcg gttgcagtcg 2051720DNARattus norvegicus 517tacttcggcg cgtattttta 2051820DNARattus norvegicus 518aaatgtgaga ttccggacta 2051920DNARattus norvegicus 519tgtgaccgcg cttgcgaact 2052020DNARattus norvegicus 520tctcgggcac cgcgagccgg 2052120DNARattus norvegicus 521cgctcctggc tggtccgcaa 2052220DNARattus norvegicus 522tgagggtagt tgagcgccgt 2052320DNARattus norvegicus 523tgagattggg accgccacgc 2052420DNARattus norvegicus 524ttgatataga ctttacgtgc 2052520DNARattus norvegicus 525agctaactac ccccccgggt 2052620DNARattus norvegicus 526cacctccagc atcgggcgag 2052720DNARattus norvegicus 527aggcggcgtc gtcgggatgt 2052820DNARattus norvegicus 528ccgtttggcc cccagttcgg 2052920DNARattus norvegicus 529ttccagcccg cgactaagcg 2053020DNARattus norvegicus 530agagtgaggg ggttgcaacg 2053120DNARattus norvegicus 531ccgcaactct tggcgcgtcg 2053220DNARattus norvegicus 532tgatcccgct cgcccgatgc 2053320DNARattus norvegicus 533ctggaagagg cggcgtcgtc 2053420DNARattus norvegicus 534aaccccgctt agtcgcgggc 2053520DNARattus norvegicus 535attgggaccg ccacgccggt 2053620DNARattus norvegicus 536tccagaaccg cgtctcgacg 2053720DNARattus norvegicus 537cgcgcacaga acgcgtacgc 2053820DNARattus norvegicus 538gtgaaacccc gcttagtcgc 2053920DNARattus norvegicus 539ccccgacgcg ccaagagttg 2054020DNARattus norvegicus 540accgcgtctc gacgcggaga 2054120DNARattus norvegicus 541ccgcgcacag aacgcgtacg 2054220DNARattus norvegicus 542ggtgaaaccc cgcttagtcg 2054320DNARattus norvegicus 543actcttggcg cgtcggggca 2054420DNARattus norvegicus 544tccttctccg cgtcgagacg 2054520DNARattus norvegicus 545acggtccgga cctccgaact 2054620DNARattus norvegicus 546ggcgccagcc gctcgctcta 2054720DNARattus norvegicus 547gacttccgag ttcgcaagcg 2054820DNARattus norvegicus 548tgcgaactcg gaagtccaat 2054920DNARattus norvegicus 549tctggaagag gcggcgtcgt 2055020DNARattus norvegicus 550tcttagcccc gagacccggt 2055120DNARattus norvegicus 551ccctccggcg ccgtgttttc 2055220DNARattus norvegicus 552ccagaaaaca cggcgccgga 2055317DNADanio rerio 553atcaatatct cgcccag 1755417DNADanio rerio 554attaaacgtg ttccact 1755517DNADanio rerio 555acgtttaatc gctttct 1755617DNADanio rerio 556caagatgcgt acggtca 1755717DNADanio rerio 557tctgtgtact cgaatat 1755817DNADanio rerio 558tcctgacttc tgggcac 1755917DNADanio rerio 559cccggtgccc agaagtc 1756017DNADanio rerio 560ccggagatcc attcatg 1756117DNADanio rerio 561tgatgaccca gaatcaa 1756217DNADanio rerio 562tgaatgaccg gtcctcc 1756317DNADanio rerio 563tccaaagggg cttcgga 1756417DNADanio rerio 564cttcagttcg ctgtgca 1756517DNADanio rerio 565gatacacgcc ggcataa 1756617DNADanio rerio 566cacacgagga tacacgc 1756717DNADanio rerio 567tcgctgtcct cttcact 1756817DNADanio rerio 568ttaaagtgat tcccagt 1756917DNADanio rerio 569cagctgatcc tccaact 1757017DNADanio rerio 570atatttagag taagaat 1757117DNADrosophila melanogaster 571aatttttatc attcagt 1757220DNADrosophila melanogaster 572gattcttcca tgatcggcca 2057320DNADrosophila melanogaster 573cgattgaaaa tccacataaa 2057420DNADrosophila melanogaster 574gctgatttcg gacaaaagtc 2057520DNADrosophila melanogaster 575aaacggatga cgtgcaacgt 2057620DNADrosophila melanogaster 576cgttttgtgt gcgtgcgcac 2057720DNADrosophila melanogaster 577ggaataatct tcgtctatcc 2057820DNADrosophila melanogaster 578ttcgctcacg aacaccaaca 2057920DNADrosophila melanogaster 579acaaagtaca tagggtgttt 2058020DNADrosophila melanogaster 580ccatcccccg taaagagggt 2058120DNADrosophila melanogaster 581tatttttagt tggtaaatta 2058217DNADrosophila melanogaster 582attcactctg ggttatg 1758320DNADrosophila melanogaster 583ctgctttcct tggccgatca 2058420DNADrosophila melanogaster 584gattatttgc tccatttatg 2058520DNADrosophila melanogaster 585ctctgtacag aatattttct 2058620DNADrosophila melanogaster 586gcagcggcag tgagagacgt 2058720DNADrosophila melanogaster 587tgtgcgcacg cacacaaaac 2058820DNADrosophila melanogaster 588aattcgctaa ttgggcagcc 2058920DNADrosophila melanogaster 589caaggaaatt gacagatgta 2059020DNADrosophila melanogaster 590gctctagcaa ccacgtcaga 2059120DNADrosophila melanogaster 591acaatacgaa tgcgagaagg 2059220DNADrosophila melanogaster 592aaataaaaat tagtgtccgc 2059317DNADrosophila melanogaster 593agagtgaata aattaca 1759420DNADrosophila melanogaster 594ttattcaaga cttacaccat
2059520DNADrosophila melanogaster 595attatgacgt tttttggtac 2059620DNADrosophila melanogaster 596ttcgggcggt ctaaagcagt 2059720DNADrosophila melanogaster 597tgttactttt tcgaacgttg 2059820DNADrosophila melanogaster 598gttttgtgtg cgtgcgcaca 2059920DNADrosophila melanogaster 599tgcccaatta gcgaattagg 2060020DNADrosophila melanogaster 600cgccccaaaa gaaaatacgt 2060120DNADrosophila melanogaster 601gctgcggctt tcgtgaaatg 2060220DNADrosophila melanogaster 602aaaaggagga gaacatatct 2060317DNADrosophila melanogaster 603atgtaattta ttcactc 1760420DNADrosophila melanogaster 604agaactaagc acgtattttt 2060520DNADrosophila melanogaster 605acaaacaatc caaataaaag 2060620DNADrosophila melanogaster 606caacaaaaaa gtgagctctg 2060720DNADrosophila melanogaster 607cgttgttcgt ttttgtagtt 2060820DNADrosophila melanogaster 608ctctttctgg taaatgctgt 2060920DNADrosophila melanogaster 609ttaggtggga ttttcgtgta 2061020DNADrosophila melanogaster 610ctacaaaact acacgaatta 2061120DNADrosophila melanogaster 611gctctcgctc tgcctcgctg 2061220DNADrosophila melanogaster 612aaaggaggag aacatatctt 2061317DNADrosophila melanogaster 613aatcaaatcg gtacatt 1761420DNADrosophila melanogaster 614atttcagctc tcaacgcaaa 2061520DNADrosophila melanogaster 615attctgaaac cacttttatt 2061620DNADrosophila melanogaster 616gagctctgcg gtagtgacgc 2061720DNADrosophila melanogaster 617ctacaaaaac gaacaacgca 2061820DNADrosophila melanogaster 618tactaaacac atactaaaac 2061920DNADrosophila melanogaster 619gtcgatttgt agaactggtg 2062020DNADrosophila melanogaster 620tggtactttt ctctcgtaca 2062120DNADrosophila melanogaster 621ttccatgctt ctttagtgca 2062220DNADrosophila melanogaster 622attaaaagcg tatttttagt 2062317DNADrosophila melanogaster 623ggttcgacac gcagaaa 1762420DNADrosophila melanogaster 624tggagcaaat aatccattca 2062520DNADrosophila melanogaster 625gctctgtaca gaatattttc 2062620DNADrosophila melanogaster 626atgttacttt ttcgaacgtt 2062720DNADrosophila melanogaster 627atcccgttag tgtgtgagat 2062820DNADrosophila melanogaster 628taggtgggat tttcgtgtag 2062920DNADrosophila melanogaster 629tctttatgta cacgagtata 2063020DNADrosophila melanogaster 630ttccatgcac taaagaagca 2063120DNADrosophila melanogaster 631ggtcgcgtcg ataccatttt 2063220DNADrosophila melanogaster 632ctttcttcct cttcttcggg 2063320DNADrosophila melanogaster 633catgttactt tttcgaacgt 2063420DNADrosophila melanogaster 634tccccaatct cacacactaa 2063520DNADrosophila melanogaster 635tgtaggggtc cctatagtag 2063620DNADrosophila melanogaster 636cacaaagtac atagggtgtt 2063720DNADrosophila melanogaster 637tcccccatcc cccgtaaaga 2063820DNADrosophila melanogaster 638ctttagaccg cccgaagaag 2063920DNADrosophila melanogaster 639tacaaaaacg aacaacgcaa 2064020DNADrosophila melanogaster 640gttgaatcgc ctgtctttaa 2064120DNADrosophila melanogaster 641ccgcgctgct catccaaact 2064220DNADrosophila melanogaster 642cttcagcgca caaagtacat 2064320DNADrosophila melanogaster 643gcaacaatac gaatgcgaga 2064420DNADrosophila melanogaster 644gccggcagag gcagtgaccg 2064520DNADrosophila melanogaster 645cgcaagggag caatgggtac 2064620DNADrosophila melanogaster 646ttgaatcgcc tgtctttaat 2064720DNADrosophila melanogaster 647agaaactttc acggcaccgt 2064820DNADrosophila melanogaster 648gcaagcgaaa tggcgcttcg 2064920DNADrosophila melanogaster 649agaggcagtg accgaggcag 2065020DNADrosophila melanogaster 650ggaaaaaacc cattaaagac 2065120DNADrosophila melanogaster 651atgcacggaa taagctccaa 2065220DNADrosophila melanogaster 652tgtaccacca accctcttta 2065320DNADrosophila melanogaster 653gaaatcttca ttccctccaa 2065420DNADrosophila melanogaster 654aaggaaattg acagatgtat 2065520DNADrosophila melanogaster 655gtaccaccaa ccctctttac 2065620DNADrosophila melanogaster 656cgcaaaaagc ctctactata 2065720DNADrosophila melanogaster 657taccaccaac cctctttacg 2065820DNADrosophila melanogaster 658tcgcaaaaag cctctactat 2065920DNADrosophila melanogaster 659accaccaacc ctctttacgg 2066020DNADrosophila melanogaster 660ccaaccctct ttacggggga 2066120DNADrosophila melanogaster 661caaccctctt tacgggggat 2066220DNADrosophila melanogaster 662atcccccatc ccccgtaaag 2066317DNACaenorhabditis elegans 663aagagagtca gattgga 1766417DNACaenorhabditis elegans 664tctgactctc ttgtgtt 1766517DNACaenorhabditis elegans 665acacaagaga gtcagat 1766617DNACaenorhabditis elegans 666attttcaggt aaaagtt 1766717DNACaenorhabditis elegans 667gaagacgctc gaattct 1766817DNACaenorhabditis elegans 668tatgtgacgt cttatct 1766917DNACaenorhabditis elegans 669tgttcctctt gatcatc 1767017DNACaenorhabditis elegans 670aagaggaaca taatcta 1767117DNACaenorhabditis elegans 671tctaggccac gtcttgc 1767217DNACaenorhabditis elegans 672agtattctat attcagt 1767317DNACaenorhabditis elegans 673tgcatggttt gaatctt 1767417DNACaenorhabditis elegans 674gcaactgatg tgtttct 1767517DNACaenorhabditis elegans 675gatgtgtttc taggctc 1767617DNACaenorhabditis elegans 676aagttctaga tcacgag 1767717DNACaenorhabditis elegans 677tcccaaatca tccttga 1767817DNACaenorhabditis elegans 678tgggttttaa accatca 1767917DNACaenorhabditis elegans 679gacgggttgg aggcaga 1768017DNACaenorhabditis elegans 680cctccaaccc gtccaat 1768117DNACaenorhabditis elegans 681aaaccgattg gacgggt 1768217DNACaenorhabditis elegans 682ttggaaaccg attggac 1768317DNACaenorhabditis elegans 683tgtctattgt acaagct 1768417DNACaenorhabditis elegans 684gtctattgta caagctt 1768517DNACaenorhabditis elegans 685tctattgtac aagcttg 1768617DNACaenorhabditis elegans 686tagcactcga cccgaaa 1768717DNACaenorhabditis elegans 687cactcgaccc gaaacgg 1768817DNACaenorhabditis elegans 688actcgacccg aaacggt 1768916DNACaenorhabditis elegans 689acccgaaacg gtggga 1669017DNACaenorhabditis elegans 690cccgaaacgg tgggaag 1769117DNACaenorhabditis elegans 691ccgaaacggt gggaagg 1769217DNACaenorhabditis elegans 692aacggtggga agggggg 1769317DNACaenorhabditis elegans 693ccccttccca ccgtttc 1769417DNACaenorhabditis elegans 694cccccttccc accgttt 1769517DNACaenorhabditis elegans 695ggggggagga catccgc 1769617DNACaenorhabditis elegans 696taggtgactg actccgg 1769717DNACaenorhabditis elegans 697ttgtaggtga ctgactc 1769817DNACaenorhabditis elegans 698ggattagctg ctccaat 1769917DNACaenorhabditis elegans 699attagctgct ccaattg 1770017DNACaenorhabditis elegans 700acttagcaca ccccaat 1770117DNACaenorhabditis elegans 701aattgtctat gtttaat 1770217DNACaenorhabditis elegans 702taatgggtgt atcgtat 1770317DNACaenorhabditis elegans 703agcacttttt gagtggt 1770417DNACaenorhabditis elegans 704aggaagtgag agtcact 1770517DNACaenorhabditis elegans 705ggaagtgaga gtcacta 1770617DNACaenorhabditis elegans 706gaatggataa agccacg 1770717DNACaenorhabditis elegans 707ccacgtggca atcatat 1770817DNACaenorhabditis elegans 708gtttatagta ggatttt 1770920DNASus scrofa 709aggatacctc aacgactacg 2071020DNASus scrofa 710gcgctgcccg acgtgtggca 2071120DNASus scrofa 711cttcaagcaa tagggtcccg 2071220DNASus scrofa 712atgacgcata tcaaatcgca 2071320DNASus scrofa 713tattgctcta tagtcggctt 2071420DNASus scrofa 714ataaccgact tagcgaattc 2071520DNASus scrofa 715aggctgctga ccgtattgcc 2071620DNASus scrofa 716ggatacctca acgactacgc 2071720DNASus scrofa 717ctcagccccg tgcaggccta 2071820DNASus scrofa 718taggaaagcg gctcggattg 2071920DNASus scrofa 719ggcatagacg ggcgttaagt 2072020DNASus scrofa 720gcggtcccta gacccaaccg 2072120DNASus scrofa 721agcccgggag tctttcgcta 2072220DNASus scrofa 722ctgctagtgg cggccgttta 2072320DNASus scrofa 723gatacctcaa cgactacgcg 2072420DNASus scrofa 724ttttcccagc aatagggtcg 2072520DNASus scrofa 725cgaatgcccg ggtacgaggg 2072620DNASus scrofa 726ggaaagaaag tctgcgcgtc 2072720DNASus scrofa 727cgggccccgc gtagtcgttg 2072820DNASus scrofa 728ggtctcctcg accctattgc 2072920DNASus scrofa 729tttatgaacg gattgcactc 2073020DNASus scrofa 730gcagaggccg gcgggtgtaa 2073120DNASus scrofa 731ggcgtcgcca tcgtagacgg 2073220DNASus scrofa 732gtctctgcga cgcgatccag 2073320DNASus scrofa 733tcacccacca accgttcatt 2073420DNASus scrofa 734cgtctacgat ggcgacgccg 2073520DNASus scrofa 735agtctctgcg acgcgatcca 2073620DNASus scrofa 736gtaggtgcct aatgaacggt 2073720DNASus scrofa 737tggctacatc aacggcgacg 2073820DNASus scrofa 738gagtctctgc gacgcgatcc 2073920DNASus scrofa 739ggggcccacg gttgggtcta 2074020DNASus scrofa 740acctccgccg tcgccgtcgg 2074120DNASus scrofa 741ttggcagagg atgcgaggcg 2074220DNASus scrofa 742cccgggccct gagcgcgcct 2074320DNASus scrofa 743cggcacgctt gcggcgagtg 2074420DNABos taurus 744cctgccagta tctcaaaccg 2074523DNAHomo sapiens 745cacatcgatg tcctccccat tcc 2374620DNAHomo sapiens 746atggggagga catcgatgtc 2074720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 747cacaucgaug uccuccccau 2074824DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 748caccgacatc gatgtcctcc ccat 2474924DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 749aaacatgggg aggacatcga tgtc 2475026DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 750gagggcctat ttcccatgat tccttc 26751125DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic polynucleotide" 751aaaaaaagca ccgactcggt gccacttttt caagttgata acggactagc cttattttaa 60cttgctattt ctagctctaa aacatgggga ggacatcgat gtcggtgttt cgtcctttcc 120acaag 12575223DNAHomo sapiens 752cacataggtg tcctccccat agg 2375323DNAHomo sapiens 753gaaatcaagg tcctccccat agg 2375423DNAHomo sapiens 754aacttccatc tcctccccat ggg 2375523DNAHomo sapiens 755gccttctctg tcctccccat ggg 2375623DNAHomo sapiens 756gagacccatt tcctccccat tag 2375723DNAHomo sapiens 757aaaaacaatg tcctccccat tag 2375823DNAHomo sapiens 758gccctcgctg ccctccccat cag 2375923DNAHomo sapiens 759cacggctgtg tcctccccat tgg 2376023DNAHomo sapiens 760tgcttcactg tcctccccat agg 2376123DNAHomo sapiens 761gtcaacgttg tcctcctcat tgg 2376223DNAArtificial
Sequencesource/note="Description of Artificial Sequence Synthetic primer" 762gagggcctat ttcccatgat tcc 2376323DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 763gagggcctat ttcccatgat tcc 2376422DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 764cttgtggaaa ggacgaaaca cc 2276545DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(4)..(23)a, c, t, g, unknown or other 765aacnnnnnnn nnnnnnnnnn nnnggtgttt cgtcctttcc acaag 4576625DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(6)..(25)a, c, t, g, unknown or other 766caccgnnnnn nnnnnnnnnn nnnnn 2576725DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"modified_base(5)..(24)a, c, t, g, unknown or other 767aaacnnnnnn nnnnnnnnnn nnnnc 2576854DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 768aacaccgggt cttcgagaag acctgtttta gagctagaaa tagcaagtta aaat 5476920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 769gaguccgagc agaagaagaa 2077020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 770gucaccucca augacuaggg 2077120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 771gucaccucca augacuagaa 2077220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 772gucaccucca augacugagg 2077320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 773gucaccucca augaucaggg 2077420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 774gucaccucca auagcuaggg 2077520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 775gucaccucca gcgacuaggg 2077620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 776gucaccucug augacuaggg 2077720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 777gucacccuca augacuaggg 2077820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 778gucauuucca augacuaggg 2077920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 779guugccucca augacuaggg 2078020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 780gucaccucca augacugaag 2078120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 781gucaccucca auggucaggg 2078220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 782gucaccucca gcaacuaggg 2078320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 783gucaccuucg augacuaggg 2078420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 784gucauuccca augacuaggg 2078520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 785gcugccucca augacuaggg 2078620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 786gucaccucca augaccgaaa 2078720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 787gucaccucca auagucgggg 2078820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 788gucaccuccg gcagcuaggg 2078920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 789gucacccuug gugacuaggg 2079020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 790gucguucuca augacuaggg 2079120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 791gacaucgaug uccuccccau 2079220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 792gacaucgaug uccuccccgc 2079320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 793gacaucgaug uccuccuuau 2079420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 794gacaucgaug uccuuuccau 2079520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 795gacaucgaug ucucccccau 2079620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 796gacaucgaug cucuccccau 2079720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 797gacaucgaca uccuccccau 2079820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 798gacaucagug uccuccccau 2079920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 799gacacugaug uccuccccau 2080020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 800gaugucgaug uccuccccau 2080120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 801gacaucgaug uccuccuugu 2080220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 802gacaucgaug ucccuuccau 2080320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 803gacaucgaug cuuuccccau 2080420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 804gacaucggca uccuccccau 2080520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 805gacacuaaug uccuccccau 2080620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 806ggugucgaug uccuccccau 2080720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 807gacaucgaug uccucuuugc 2080820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 808gacaucgaug ucucuuucau 2080920RNAArtificial Sequencesource/note="Description of Artificial SequenceSynthetic oligonucleotide" 809gacaucgaua cuucccccau 2081020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 810gacaucagca cccuccccau 2081120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 811gacgcuagug uccuccccau 2081220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 812gaguccgagc agaagaagaa 2081320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 813gaguccgagc agaagaaggg 2081420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 814gaguccgagc agaagagaaa 2081520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 815gaguccgagc agaaagagaa 2081620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 816gaguccgagc aggggaagaa 2081720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 817gaguccgagc gaaagaagaa 2081820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 818gaguccgaau agaagaagaa 2081920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 819gaguccaggc agaagaagaa 2082020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 820gagucugagc agaagaagaa 2082120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 821gaacccgagc agaagaagaa 2082220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 822gaguccgagc agaagagaga 2082320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 823gaguccgagc agagagagaa 2082420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 824gaguccgagc gagagaagaa 2082520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 825gaguccggau agaagaagaa 2082620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 826gagucuaagc agaagaagaa 2082720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 827ggacccgagc agaagaagaa 2082820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 828gaguccgagc agaagggagg 2082920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 829gaguccgagc agggagggaa 2083020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 830gaguccgagu gagggaagaa 2083120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 831gaguccagau ggaagaagaa 2083220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 832gagccuaggc agaagaagaa 2083320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 833gcgccaccgg uugaugugau 2083420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 834gcgccaccgg uugauguggc 2083520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 835gcgccaccgg uugaugcaau 2083620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 836gcgccaccgg uugacaugau 2083720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 837gcgccaccgg uuagugugau 2083820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 838gcgccaccgg ccgaugugau 2083920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 839gcgccaccaa uugaugugau 2084020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 840gcgccauugg uugaugugau 2084120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 841gcgcugccgg uugaugugau 2084220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 842gcaucaccgg uugaugugau 2084320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 843gcgccaccgg uugaugcagu 2084420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 844gcgccaccgg uuggcaugau 2084520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 845gcgccaccgg ccaaugugau 2084620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 846gcgccacuaa uugaugugau 2084720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 847gcgcugucgg uugaugugau 2084820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 848guaucaccgg uugaugugau 2084920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 849gcgccaccgg uugauacagc 2085020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 850gcgccaccgg uuagcacgau 2085120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 851gcgccaccga ccagugugau 2085220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 852gcgccauuaa cugaugugau 2085320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 853gcguuguugg uugaugugau 2085420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 854gucaccucca augacuaggg 2085520RNAArtificial Sequencesource/note="Description of Artificial
Sequence Synthetic oligonucleotide" 855gucaccucca augacuaaga 2085620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 856gucaccucca augauuagga 2085720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 857gucaccucca gugacuagga 2085820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 858gucacuucca augacuagga 2085920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 859aucaccucca augacuagga 2086020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 860gucaccucca augaccaagg 2086120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 861gucaccucca auaacuaagg 2086220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 862gucaccucua augacuaagg 2086320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 863gucgccucca augacuaagg 2086420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 864gucaccucca auggccaggg 2086520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 865gucaccucca gugaccaggg 2086620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 866gucaccccca augaccaggg 2086720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 867gccaccucca augaccaggg 2086820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 868gacaucgaug uccuccccau 2086920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 869gacaucgaug uccucccuag 2087020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 870gacaucgaug uccuucccag 2087120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 871gacaucgaug cccuccccag 2087220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 872gacauugaug uccuccccag 2087320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 873aacaucgaug uccuccccag 2087420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 874gacaucgaug uccucucuau 2087520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 875gacaucgaug ucuucccuau 2087620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 876gacaucgacg uccucccuau 2087720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 877gacgucgaug uccucccuau 2087820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 878gacaucgaug uccccuccau 2087920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 879gacaucgaug cccucuccau 2088020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 880gacaucaaug uccucuccau 2088120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 881ggcaucgaug uccucuccau 2088220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 882gaguccgagc agaagaagaa 2088320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 883gaguccgagc agaagaaaag 2088420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 884gaguccgagc agaaaaagag 2088520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 885gaguccgagc ggaagaagag 2088620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 886gagucugagc agaagaagag 2088720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 887aaguccgagc agaagaagag 2088820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 888gaguccgagc agaagcaaaa 2088920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 889gaguccgagc aggagaaaaa 2089020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 890gaguccgaac agaagaaaaa 2089120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 891gagcccgagc agaagaaaaa 2089220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 892gaguccgagc agagggagaa 2089320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 893gaguccgagc ggaaggagaa 2089420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 894gaguccaagc agaaggagaa 2089520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 895ggguccgagc agaaggagaa 2089620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 896gcgccaccgg uugaugugau 2089720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 897gcgccaccgg uugauguaac 2089820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 898gcgccaccgg uugaugugac 2089920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 899gcgccaccgg cugaugugac 2090020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 900gcgccgccgg uugaugugac 2090120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 901acgccaccgg uugaugugac 2090220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 902gcgccaccgg uugauauaau 2090320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 903gcgccaccgg uuaauguaau 2090420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 904gcgccaccag uugauguaau 2090520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 905gcgucaccgg uugauguaau 2090620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 906gcgccaccgg uugguaugau 2090720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 907gcgccaccgg cugauaugau 2090820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 908gcgccaucgg uugauaugau 2090920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 909gugccaccgg uugauaugau 2091020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 910gucaccucca augacuaggg 2091120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 911gucaccucca augaccaaga 2091220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 912gucaccucca auggcuggga 2091320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 913gucaccucca acgaccagga 2091420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 914gucaccuccg augauuagga 2091520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 915gucaccucca auggccaagg 2091620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 916gucaccucca acgauuaagg 2091720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 917gucaccuccg auggcuaagg 2091820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 918gucaccuuca auaacuaagg 2091920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 919gucaccucca auggccaaga 2092020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 920gucaccucca guggcuggga 2092120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 921gucaccuuca acgaccagga 2092220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 922gucaucuccg augauuagga 2092320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 923gccaccuuca auggcuagga 2092420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 924gacaucgaug uccuccccau 2092520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 925gacaucgaug uccucucuag 2092620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 926gacaucgaug ucccccucag 2092720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 927gacaucgaug uucucuccag 2092820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 928gacaucgaua uccuucccag 2092920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 929gacaucgaug uccccucuau 2093020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 930gacaucgaug uucuuccuau 2093120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 931gacaucgaua uccccccuau 2093220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 932gacaucggug ucaucccuau 2093320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 933gacaucgaug uccccucuag 2093420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 934gacaucgaug ccccccucag 2093520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 935gacaucggug uucucuccag 2093620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 936gacaccgaua uccuucccag 2093720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 937ggcaucggug ucccccccag 2093820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 938gaguccgagc agaagaagaa 2093920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 939gaguccgagc agaaggaaag 2094020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 940gaguccgagc agaggaggag 2094120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 941gaguccgagc aaaaggagag 2094220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 942gaguccgagu agaaaaagag 2094320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 943gaguccgagc agagggaaaa 2094420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 944gaguccgagc aaaaaaaaaa 2094520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 945gaguccgagu agaggaaaaa 2094620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 946gaguccgggc aggagaaaaa 2094720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 947gaguccgagc agagggaaag 2094820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 948gaguccgagc ggaggaggag 2094920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic
oligonucleotide" 949gaguccgggc aaaaggagag 2095020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 950gaguucgagu agaaaaagag 2095120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 951ggguccgggc agaggaagag 2095220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 952gcgccaccgg uugaugugau 2095320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 953gcgccaccgg uugauauaac 2095420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 954gcgccaccgg uuggugcgac 2095520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 955gcgccaccgg ucgauaugac 2095620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 956gcgccaccga uugacgugac 2095720RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 957gcgccaccgg uugguauaau 2095820RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 958gcgccaccgg ucgacguaau 2095920RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 959gcgccaccga uugguguaau 2096020RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 960gcgccacugg uuaauguaau 2096120RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 961gcgccaccgg uugguauaac 2096220RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 962gcgccaccgg cuggugcgac 2096320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 963gcgccacugg ucgauaugac 2096420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 964gcgcuaccga uugacgugac 2096520RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 965gugccacugg uuggugugac 2096620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 966gucaccucca augacuaggg 2096723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 967gucaccucca augacuaggg tgg 2396823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 968gucaacucca augatuagga cag 2396923DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 969gucaccucca cutccuaggg cag 2397023DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 970gucactucca aggacuagag aag 2397123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 971gucaccucca gggtcuaggg cag 2397223DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 972gucaacucca augacutggg agg 2397323DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 973gucaccutaa augacutggg aag 2397423DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 974tucaccucca aaaacuaggg aag 2397523DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 975gctaccucca gugacuaggg aag 2397612DNAHomo sapiens 976tgactagggt gg 1297711DNAHomo sapiens 977tgactgggtg g 1197814DNAHomo sapiens 978tgactatagg gtgg 1497912DNAHomo sapiens 979tgactaggga ag 1298011DNAHomo sapiens 980tgactgggaa g 1198114DNAHomo sapiens 981tgactatagg gaag 1498214DNAHomo sapiens 982tgactggagg gaag 1498313DNAHomo sapiens 983tgactacggg aag 1398420RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 984gaguccgagc agaagaagaa 2098523RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 985gaguccgagc agaagaagaa ggg 2398623RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 986gaggccgagc agaagaaaga cgg 2398723RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 987gagucccagc agaggaagca gag 2398823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 988gaguccaagc agaggaagga tag 2398923DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 989gagucagagc agaactagaa ggg 2399023RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 990gagucccagg agaagaagag agg 2399123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 991gaguccaagc agtagaggaa ggg 2399223DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 992gaguccgaga aaatgaagaa gag 2399323RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 993gagucccaga agaagaaaaa aag 2399423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 994gagucccagg agaagaaaaa cag 2399523RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 995gagucagaac agaagaacaa cag 2399623DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 996gaguccaaga agaataagaa tag 2399723RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 997gaauccaagc agaagaagag aag 2399823RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 998gagucagacc aggagaagaa gag 2399923RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 999aaguccgaga agaagcagaa aag 23100023DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1000gaguctgggc aggagaagaa gag 23100123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1001gaguctgaac ggaagaagaa aag 23100223DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1002aaguctgagc agaagaagca cag 23100323RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1003aaguccgagg agaggaagaa agg 23100423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1004gaggcccagc agaggaagaa gag 23100523DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1005gagutccaga agaagaagaa gag 23100623DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1006gaguactaga agaagaagaa aag 23100723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1007gacucctagc aaaagaagaa tgg 23100823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1008gaguttgagt agaagaagaa gag 23100923RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1009gaguaagaga agaagaagaa ggg 23101023RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1010gaguaggagg agaagaagaa agg 23101123DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1011gagucctagc aggagaagaa gag 23101223DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1012gatucctacc agaagaagaa tgg 23101323RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1013gagucccagc aaaagaagaa aag 23101423RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1014gagagcaagc agaagaagaa ggg 23101523RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1015aagucagagg agaagaagaa aag 23101623DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1016gaguctaagc agaagaagaa gag 23101723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1017acguctgagc agaagaagaa tgg 23101812DNAHomo sapiens 1018gaagaagaag gg 12101913DNAHomo sapiens 1019gaagaaggaa ggg 13102013DNAHomo sapiens 1020gaagaaagaa ggg 13102115DNAHomo sapiens 1021gaagaattag aaggg 15102214DNAHomo sapiens 1022gaagaacaga aggg 14102311DNAHomo sapiens 1023gaagagaagg g 11102411DNAHomo sapiens 1024gaagaaaagg g 11102512DNAHomo sapiens 1025gaagaaagac gg 12102611DNAHomo sapiens 1026gaagaagacg g 11102713DNAHomo sapiens 1027gaagaaaaga cgg 13102813DNAHomo sapiens 1028gaagaataga cgg 13102913DNAHomo sapiens 1029gaagaacaga cgg 13103014DNAHomo sapiens 1030gaagaataag acgg 14103112DNAHomo sapiens 1031ggagaagaag ag 12103213DNAHomo sapiens 1032ggagaaagaa gag 13103311DNAHomo sapiens 1033ggagagaaga g 11103411DNAHomo sapiens 1034ggagaaaaga g 11103514DNAHomo sapiens 1035ggagaataga agag 14103612DNAHomo sapiens 1036gaagaagaag ag 12103713DNAHomo sapiens 1037gaagaaagaa gag 13103811DNAHomo sapiens 1038gaagagaaga g 11103911DNAHomo sapiens 1039gaagaaaaga g 111040384DNAHomo sapiens 1040cccagtggct gctctggggg cctcctgagt ttctcatctg tgcccctccc tccctggccc 60aggtgaaggt gtggttccag aaccggagga caaagtacaa acggcagaag ctggaggagg 120aagggcctga gtccgagcag aagaagaagg gctcccatca catcaaccgg tggcgcattg 180ccacgaagca ggccaatggg gaggacatcg atgtcacctc caatgactag ggtgggcaac 240cacaaaccca cgagggggca gagtgctgct tgctgctggc caggcccctg cgtgggccca 300agctggactc tggccactcc ctggccaggc tttggggagg cctggagtca tggccccaca 360gggcttgaag cccggggccg ccat 384104123DNAHomo sapiens 1041gtcacctcca atgactaggg tgg 23104223DNAHomo sapiens 1042gacatcgatg tcctccccat tgg 23104323DNAHomo sapiens 1043gagtccgagc agaagaagaa ggg 23104423DNAHomo sapiens 1044gcgccaccgg ttgatgtgat ggg 23104523DNAHomo sapiens 1045ggggcacaga tgagaaactc agg 23104623DNAHomo sapiens 1046gtacaaacgg cagaagctgg agg 23104723DNAHomo sapiens 1047ggcagaagct ggaggaggaa
ggg 23104823DNAHomo sapiens 1048ggagcccttc ttcttctgct cgg 23104923DNAHomo sapiens 1049gggcaaccac aaacccacga ggg 23105023DNAHomo sapiens 1050gctcccatca catcaaccgg tgg 23105123DNAHomo sapiens 1051gtggcgcatt gccacgaagc agg 23105223DNAHomo sapiens 1052ggcagagtgc tgcttgctgc tgg 23105323DNAHomo sapiens 1053gcccctgcgt gggcccaagc tgg 23105423DNAHomo sapiens 1054gagtggccag agtccagctt ggg 23105523DNAHomo sapiens 1055ggcctcccca aagcctggcc agg 23105620RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1056gacaucgaug uccuccccau 20105723DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1057gacaucgaug uccuccccau tgg 23105823DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1058gacaucgaua gccuccccac tgg 23105923RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1059gacaccgaug ucaucuccau cag 23106023RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1060gacaugaaug accuccccau cag 23106123RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1061gaaaucaagg uccuccccau agg 23106223RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1062cacauaggug uccuccccau agg 23106320RNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1063gcgccaccgg uugaugugau 20106423DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1064gcgccaccgg uugaugugau tgg 23106523DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide"source/note="Description of Combined DNA/RNA Molecule Synthetic oligonucleotide" 1065gggccatggg uugaugugac gag 23106627DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1066gcccgggtgg aactggtagc catgaat 27106726DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1067gttgaagatg aagcccagag cggagt 26106827DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1068gcttccgacg aggtggccat caaggat 27106927DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1069gcaccatctc tccgtggtac cccgggt 27107026DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1070ggtggaactg gtagccatga atgaga 26107127DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1071gccatgaatg agaccgaccc aaagagc 27107227DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1072gcatcctcgt gggcacttcc gacgagg 27107327DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1073gcagagcgga gtgctgttct cccaagt 27107426DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1074ggtcggtctc attcatggct accagt 26107526DNAArtificial Sequencesource/note="Description of Artificial Sequence Synthetic oligonucleotide" 1075gcaataaaag gtgctattgc tatagt 26107620DNASus scrofa 1076gggagtcttt cgctagggtg 20
* * * * *
References
-
ploscompbiol.org
-
dx.doi.org/10.1016/j.cell.2015.02.038
-
sciencemag.org/content/339/6121/819/suppl/DC1
-
independent.co.uk/news/science/scientific-split--the-human-genome-breakthrough-dividing-former-colleagues-9300456.html
-
Nature.com/doi:10.1038/nature09886:1-35
-
mobilednajournal.com/contents5/1/2
-
dx.doi.org/10.4161/rna.24203
-
dx.doi.org/10.1155/2014/270805
- /biology-direct.com/content/6/1/38
-
asbmb.org/asbmbtoday/201408/Features/Doudna
-
catalyst.berkeley.edu/slideshow/the-crispr-revolution
-
uniprot.org/uniprot/QOP897.txt?version=28
-
ozy.com/rising-stars-and-provocateurs/jennifer-doudna-crispr-code-killer/4690
-
newscenter.berkeley.edu/2013/01/07/cheap-and-easy-technique-to-snip-dna-could-revolutionize-gene-therapy
-
biorxiv.org/content/early/2015/07/28/023366
-
groups.google.com/forum/#!profo_nt50txrP9Yb6e_LXccolb9hNf7gKeMLt6rgaVQ4fOsQ/crispr/fkhX7Fu3r-l/rziHxKT76pYJ
-
pnas.org/cgi/doi/10.1073/pnas.1308335110
-
www/addgenen.org/41824
-
ncbi.nlm.nih.gov/protein/403411236?sat=16&satkey=13804560
-
uniprot.org/uniprot/Q73Qw6.txt?version4
-
uniprot.org/uniprot/D0W2Z9.txt?version=4
-
uniprot.org/uniprot/G1UFN3.txt?version=3
-
uniprot.org/uniprot/J3TRJ9.txt?version=2
-
dx.doi.org/10.1101/091611
-
uniprot.org/uniprot/Q6NK13.txt?version=43
-
uniprot.org/uniprot/Q0P897.txt?version=28
-
news.mit.edu/2017/crispr-carrying-nanoparticles-edit-genome-1113
-
ibis/exam/dbfetch.jsp?id=uniprot:J7RUA5
-
groups.google.com/forum/#!topic/crispr/g9Q8U1tNSis
-
ncbi.nlm.nih.gov/nuccore/HE980450
-
jcvi.org/cgi-bin/tigrfams/HmmReportPage.cgi?acc=TIGR04330
-
groups.google/forum/#!top1c/crispr/5BpJj_Y3ylG
-
novocraft.com
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

D00012

D00013

D00014

D00015
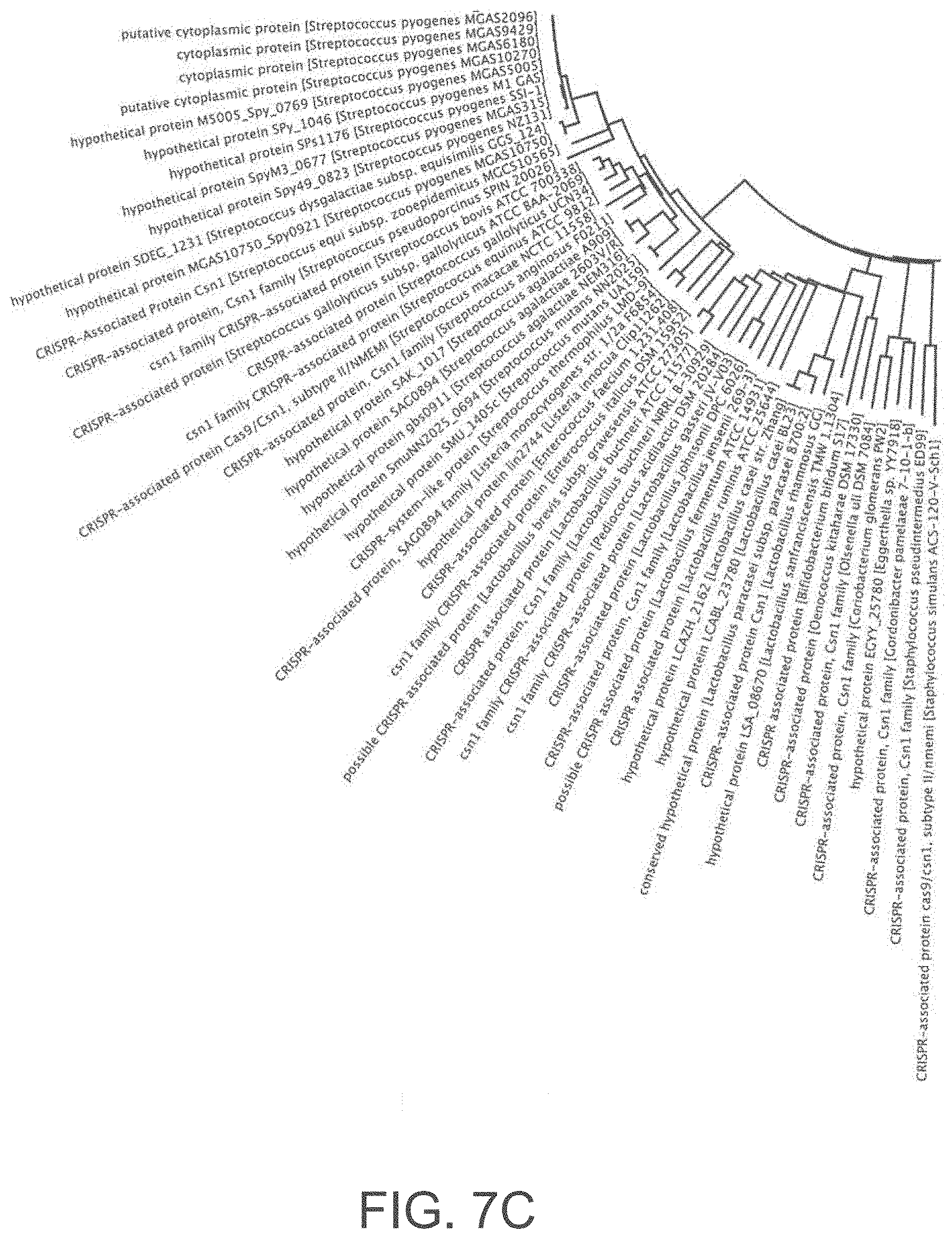
D00016

D00017

D00018

D00019

D00020

D00021

D00022

D00023

D00024

D00025

D00026

D00027

D00028
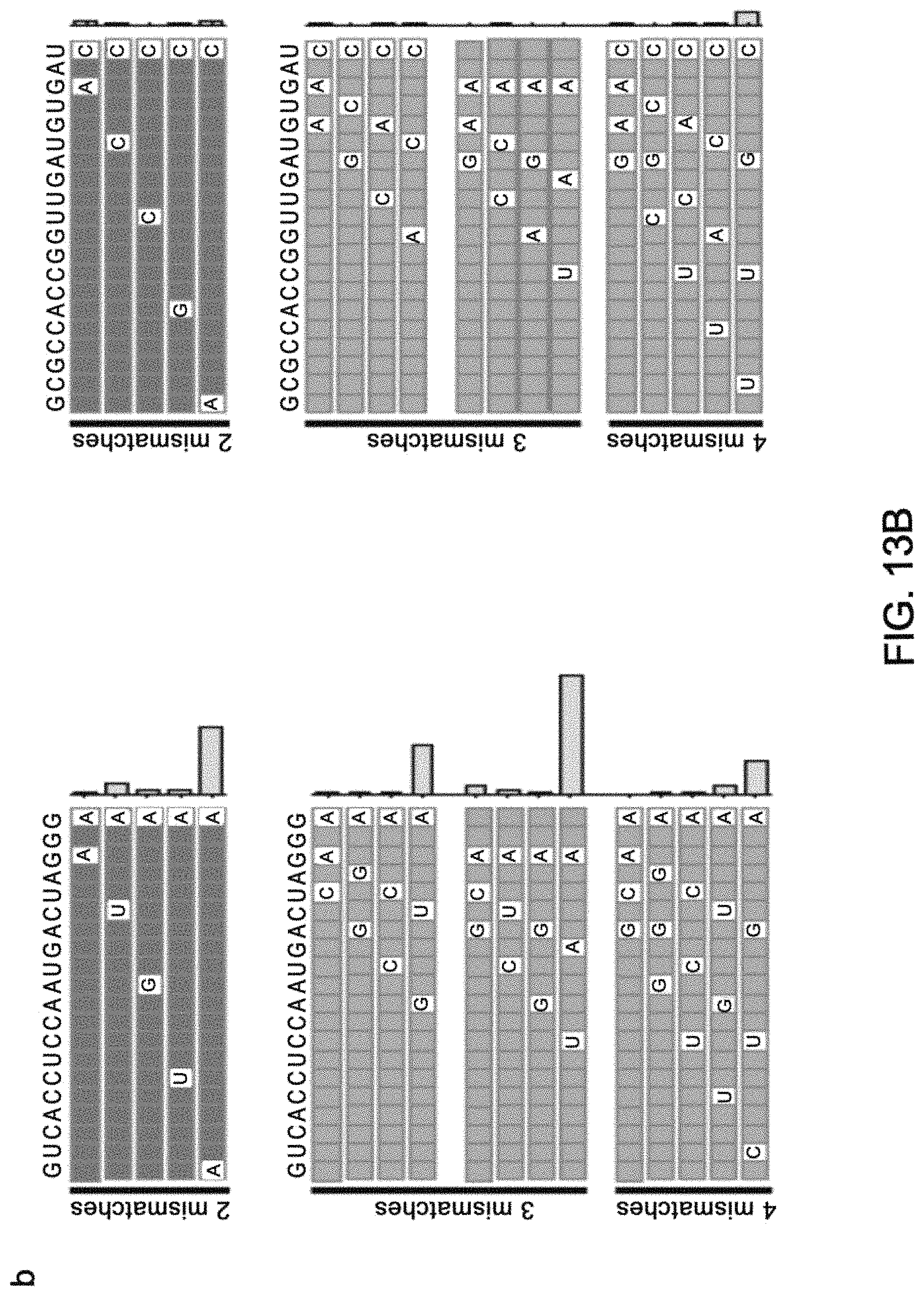

D00029

D00030

D00031

D00032

D00033

D00034

D00035

D00036

D00037

D00038
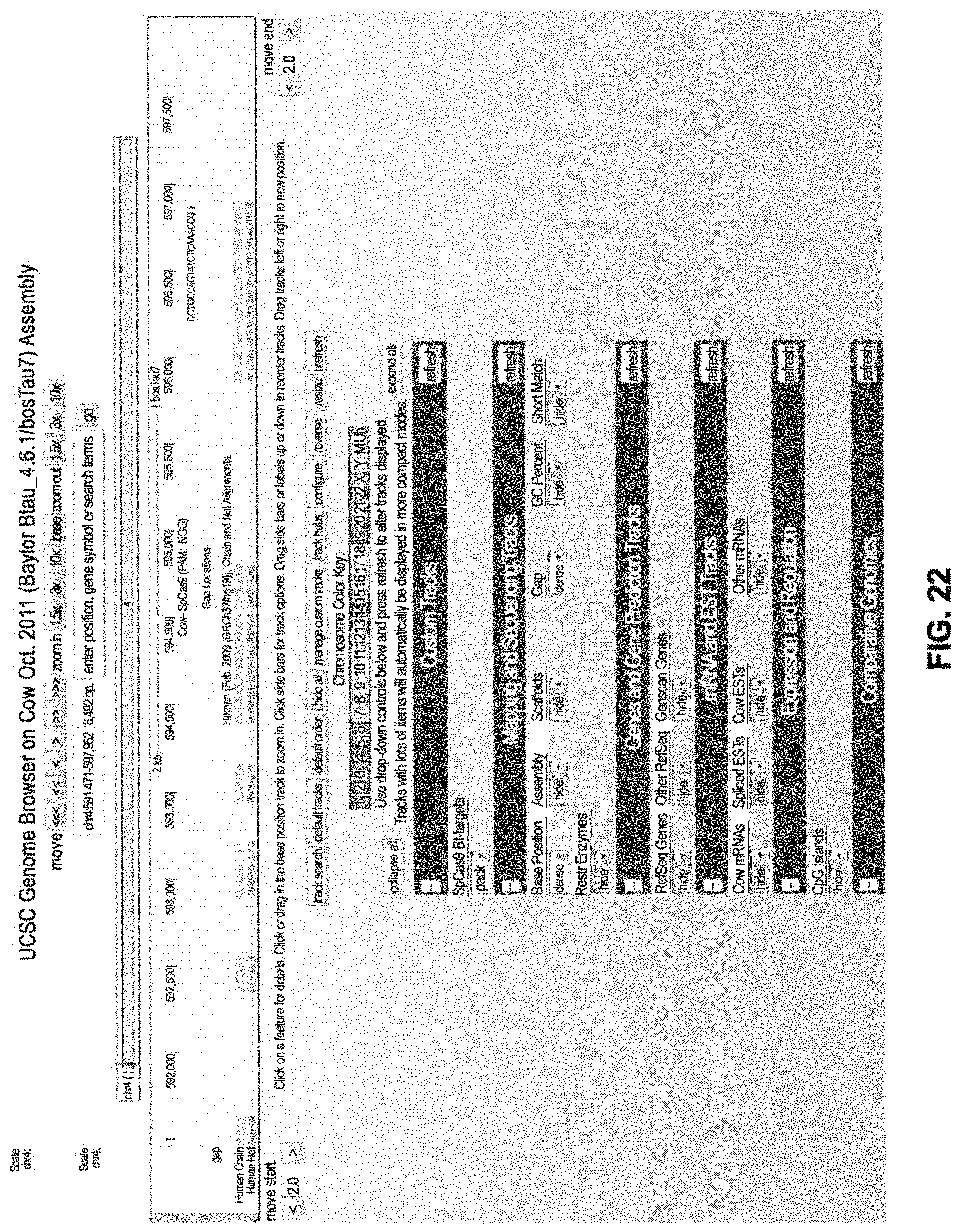
D00039

D00040

D00041

D00042

D00043

D00044

D00045

D00046

D00047

D00048

D00049

D00050

D00051
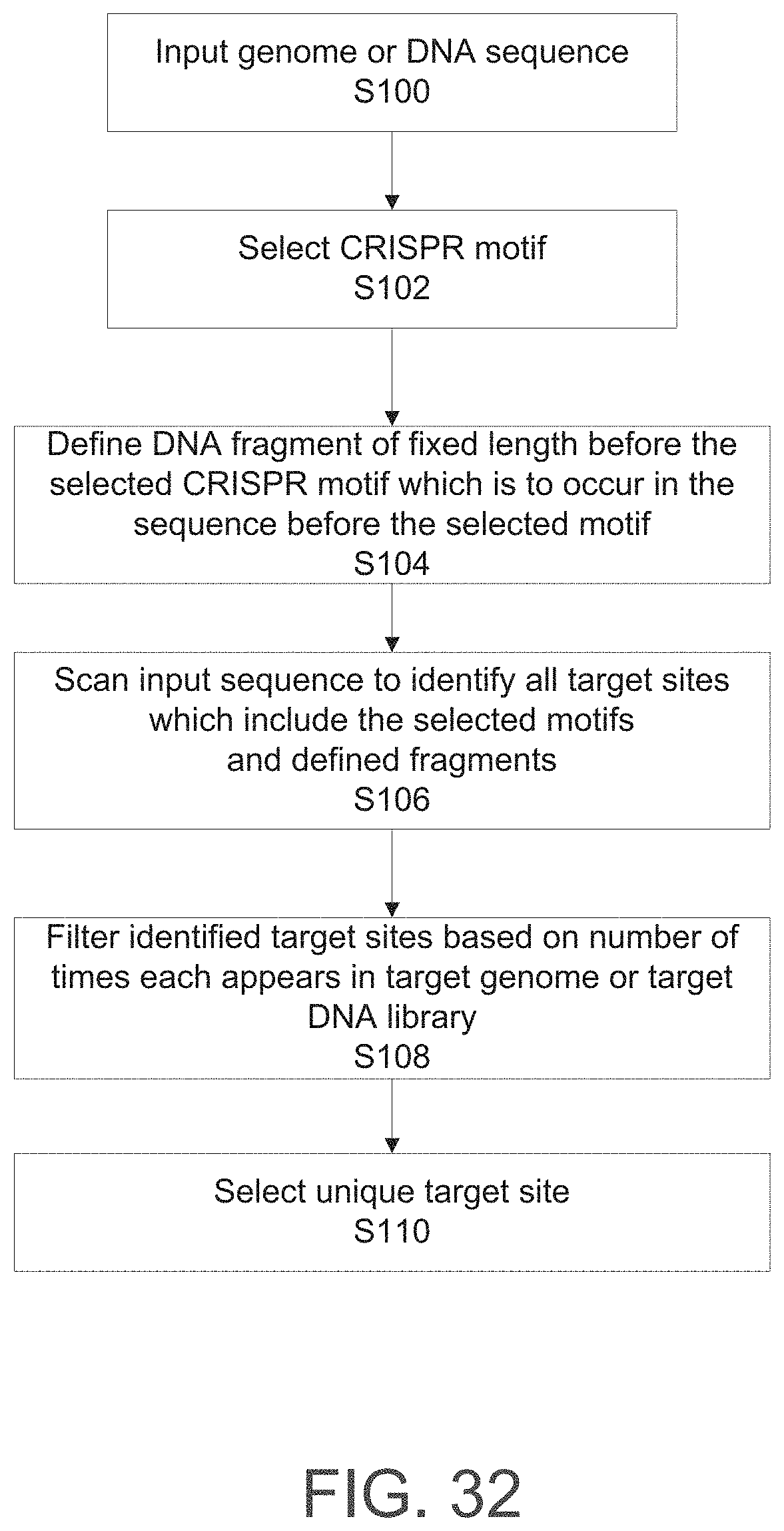
D00052

D00053

D00054

D00055

D00056
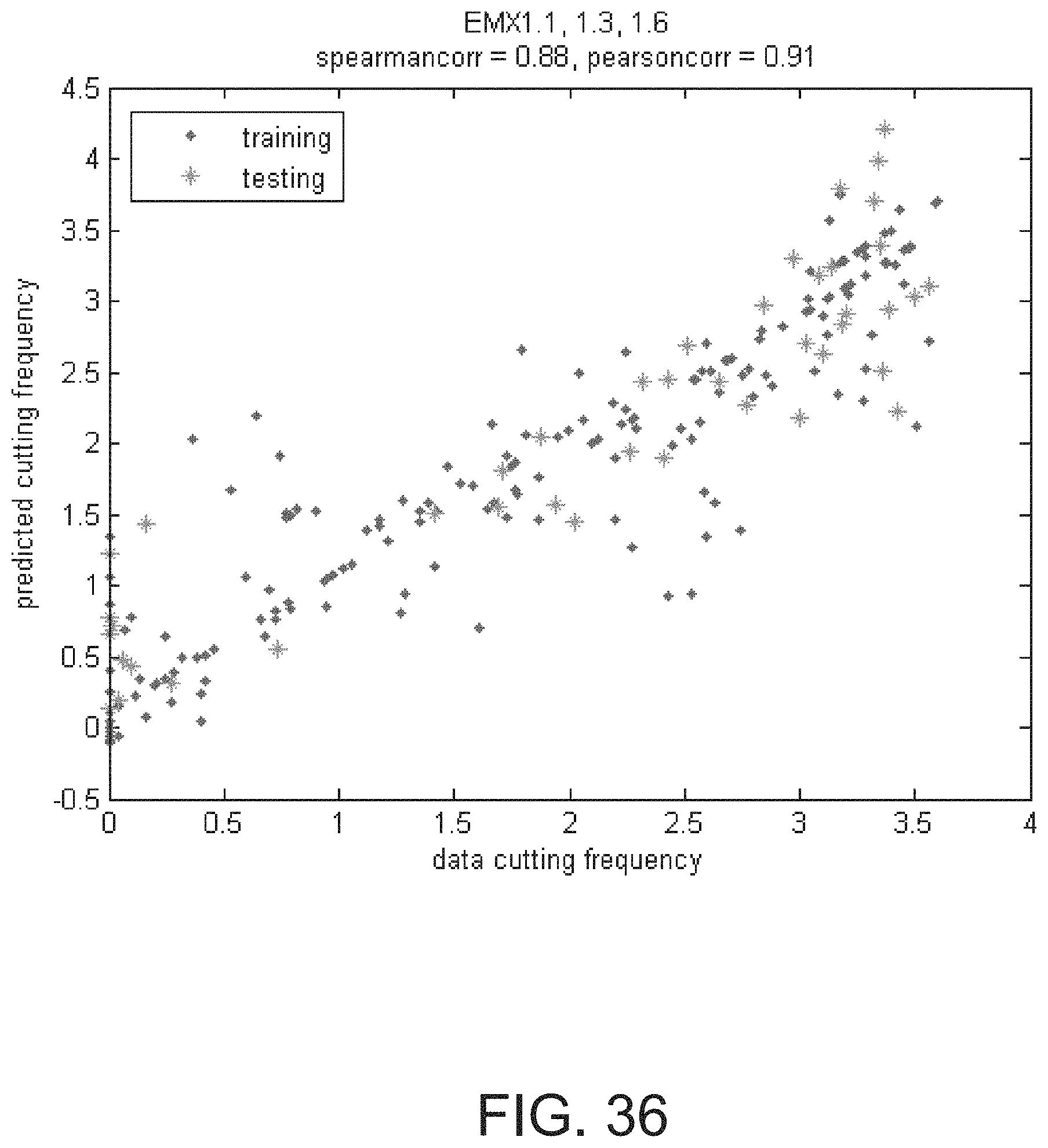
D00057

M00001

M00002

M00003

M00004

M00005

M00006

M00007

M00008

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.