Cyclic RGD cell-binding motif and uses thereof
Hedhammar
U.S. patent number 10,316,069 [Application Number 15/739,057] was granted by the patent office on 2019-06-11 for cyclic rgd cell-binding motif and uses thereof. This patent grant is currently assigned to SPIBER TECHNOLOGIES AB. The grantee listed for this patent is Spiber Technologies AB. Invention is credited to My Hedhammar.







| United States Patent | 10,316,069 |
| Hedhammar | June 11, 2019 |
Cyclic RGD cell-binding motif and uses thereof
Abstract
A recombinant fusion protein is comprising a spider silk fragment and a cyclic RGD cell-binding motif with selectivity for integrins, such as for .alpha.5.beta.1 integrins. The fusion protein is useful as a cell scaffold material and for the cultivation of cells displaying integrins on their cell surface.
| Inventors: | Hedhammar; My (Stockholm, SE) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Applicant: |
|
||||||||||
| Assignee: | SPIBER TECHNOLOGIES AB
(Stockholm, SE) |
||||||||||
| Family ID: | 53513980 | ||||||||||
| Appl. No.: | 15/739,057 | ||||||||||
| Filed: | June 23, 2016 | ||||||||||
| PCT Filed: | June 23, 2016 | ||||||||||
| PCT No.: | PCT/EP2016/064543 | ||||||||||
| 371(c)(1),(2),(4) Date: | December 21, 2017 | ||||||||||
| PCT Pub. No.: | WO2016/207281 | ||||||||||
| PCT Pub. Date: | December 29, 2016 |
Prior Publication Data
| Document Identifier | Publication Date | |
|---|---|---|
| US 20180170977 A1 | Jun 21, 2018 | |
Foreign Application Priority Data
| Jun 26, 2015 [EP] | 15174072 | |||
| Current U.S. Class: | 1/1 |
| Current CPC Class: | C08H 1/00 (20130101); C12N 15/62 (20130101); C08L 89/00 (20130101); C07K 14/43518 (20130101); C12N 5/0068 (20130101); C07K 2319/33 (20130101); C12N 2533/50 (20130101); C07K 2319/70 (20130101); C07K 16/2839 (20130101) |
| Current International Class: | A61K 38/00 (20060101); C12N 5/00 (20060101); C07K 14/435 (20060101); C08H 1/00 (20060101); C12N 15/62 (20060101); C07K 16/28 (20060101) |
| WO 2007/078239 | Jul 2007 | WO | |||
| WO 2013/185027 | Dec 2013 | WO | |||
| WO 2014/027042 | Feb 2014 | WO | |||
| WO 2015/036619 | Mar 2015 | WO | |||
Other References
|
Bini et al., "RGD-Functionalized Bioengineered Spider Dragline Silk Biomaterial", Biomacromolecules, vol. 7, No. 11, 2006, pp. 3139-3145. cited by applicant . European Search Report for EP 15 17 4072, dated Oct. 15, 2015. cited by applicant . International Search Report issued in PCT/EP2016/064543 (PCT/ISA/210), dated Oct. 21, 2016. cited by applicant . Ivanov et al., "Synthesis and Use of a New Bromoacetyl-Derivatized Heterotrifunctional Amino Acid for Conjugation of Cyclic RGD-Containing Peptides Derived from Human Bone Sialoprotein", Bioconjugate Chemistry, vol. 6, No. 3, 1995, pp. 269-277. cited by applicant . Koivunen et al., "Phage Libraries Displaying Cyclic Peptides with Different Ring Sizes: Ligand Specificities of the RGD-Directed Integrins", BioTechnology, vol. 13, Mar. 1995, pp. 265-270. cited by applicant . Shalaly et al., "Silk matrices promote formation of insulin-secreting islet-like clusters", Biomaterials, vol. 90, 2016, pp. 50-61. cited by applicant . Widhe et al., "A fibronectin mimetic motif improves integrin mediated cell biding to recombinant spider silk matrices", Biomaterials, vol. 74, 2016, pp. 256-266. cited by applicant . Written Opinion of the International Searching Authority issued in PCT/EP2016/064543 (PCT/ISA/237), dated Oct. 21, 2016. cited by applicant. |
Primary Examiner: Lieb; Jeanette M
Attorney, Agent or Firm: Birch, Stewart, Kolasch & Birch, LLP
Claims
The invention claimed is:
1. A recombinant fusion protein comprising a spidroin fragment and a cell-binding motif with selectivity for integrins, wherein the cell-binding motif is comprising the amino acid sequence C.sup.1X.sup.1X.sup.2RGDX.sup.3X.sup.4X.sup.5C.sup.2 wherein X.sup.1 is S or T; X.sup.2 is G, A or V; X.sup.3 is S or T; X.sup.4 is G, A, V or P; and X.sup.5 is G, A or V; and C.sup.1 and C.sup.2 are connected via a disulphide bond.
2. A recombinant fusion protein according to claim 1, wherein X.sup.2 is G or A.
3. A recombinant fusion protein according to claim 1, wherein X.sup.2 is G.
4. A recombinant fusion protein according to claim 1, wherein X.sup.3 is S.
5. A recombinant fusion protein according to claim 1, wherein X.sup.4 is G or P.
6. A recombinant fusion protein according to claim 5, wherein X.sup.4 is P.
7. A recombinant fusion protein according to claim 1, wherein X.sup.5 is G or A.
8. A recombinant fusion protein according to claim 7, wherein X.sup.5 is A.
9. A recombinant fusion protein according to claim 1 wherein the cell-binding motif is comprising the amino acid sequence CTGRGDSPAC (SEQ ID NO: 10).
10. A recombinant fusion protein according to claim 1, wherein the cell-binding motif has selectivity for .alpha.5.beta.1 integrins.
11. A recombinant fusion protein according to claim 1, wherein the spidroin fragment is comprising the protein moiety CT, wherein CT is a fragment of from 70 to 120 amino acid residues, having at least 70% identity to any one of SEQ ID NO: 3 and 29-59.
12. A recombinant fusion protein according to claim 11, wherein CT has at least 80% identity to any one of SEQ ID NO: 3 and 29-59.
13. A recombinant fusion protein according to claim 11, wherein CT has at least 80% identity to SEQ ID NO: 3.
14. A recombinant fusion protein according to claim 1, wherein the spidroin fragment is further comprising the protein moiety REP, wherein REP is a repetitive fragment of from 70 to 300 amino acid residues, selected from the group consisting of L(AG).sub.nL, L(AG).sub.nAL, L(GA).sub.nL, and L(GA).sub.nGL, wherein n is an integer from 2 to 10; each individual A segment is an amino acid sequence of from 8 to 18 amino acid residues, wherein from 0 to 3 of the amino acid residues are not Ala, and the remaining amino acid residues are Ala; each individual G segment is an amino acid sequence of from 12 to 30 amino acid residues, wherein at least 40% of the amino acid residues are Gly; and each individual L segment is a linker amino acid sequence of from 0 to 30 amino acid residues.
15. A cell scaffold material comprising a protein polymer which as a repeating unit is containing a recombinant fusion protein comprising a spidroin fragment and a cell-binding motif with selectivity for integrins, wherein the cell-binding motif is comprising the amino acid sequence C.sup.1X.sup.1X.sup.2RGDX.sup.3X.sup.4X.sup.5C.sup.2 wherein X.sup.1 is S or T; X.sup.2 is G, A or V; X.sup.3 is S or T; X.sup.4 is G, A, V or P; and X.sup.5 is G, A or V; and C.sup.1 and C.sup.2 are connected via a disulphide bond.
16. A cell scaffold material according to claim 15, wherein the protein polymer is in a physical form selected from the group consisting of film, coating, foam, fiber and fiber-mesh.
17. A cell scaffold material according to claim 15, wherein the protein polymer is in a physical form of a free-standing matrix.
18. A method for the cultivation of cells, comprising the steps of providing a sample of cells; applying the sample to a cell scaffold material; and maintaining the cell scaffold material having the cells applied thereto under conditions suitable for cell culture; wherein the cell scaffold material is as defined in claim 15.
19. A method according to claim 18, wherein the cells are displaying .alpha.5.beta.1 integrins on their cell surface; and wherein the cell-binding motif of the recombinant fusion protein has selectivity for .alpha.5.beta.1 integrins.
20. A method according to claim 18, wherein the cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines.
Description
TECHNICAL FIELD OF THE INVENTION
The present invention relates to the fields of eukaryotic cell culture and tissue engineering. The invention provides new proteins, a cell scaffold material comprising the proteins, and a method for cultivation of cells wherein polymers of the new proteins are used as a cell scaffold material.
BACKGROUND TO THE INVENTION
The phenotype of a cell is largely influenced by its display of integrins. By expressing several types of integrins on its surface, the cell is able to bind multiple kinds of ligands and thereby interpret parallel signals from the surrounding extracellular matrix (ECM). Cells cultured in vitro often express a different kind of integrin pattern than corresponding cells in vivo. In order to maintain the original phenotype of cells, or to accomplish a specific cellular response (e.g. differentiation, proliferation), it is important to enable integrin binding also during in vitro culture. This is most commonly done by coating cell culture plastics with ECM proteins like laminin, fibronectin, collagen or vitronectin, or mimics thereof. The ECM coatings will provide ligands for various integrins, with activation of different cellular pathways as a result. However, within several cell culture disciplines it is desirable to find ways to accomplish this on a defined matrix without the use of animal derived substrates.
WO 2011/129756 discloses methods and a cell scaffold material based on a miniature spider silk protein for eukaryotic cell culture. The protein may contain various short (3-5 amino acid residues) cell-binding peptides.
WO 2012/055854 discloses polymers consisting of a fusion protein containing a miniature spider silk protein and a large non-spidroin protein fragment of more than 30 amino acid residues which provides affinity to another molecule. The fusion protein may additionally contain various cell-binding peptides.
WO 2015/036619 discloses polymers consisting of a fusion protein containing a miniature spider silk protein and a cell-binding peptide comprising the amino acid residues RGD. The fusion protein is useful for cultivation of human pluripotent stem cells (hPSCs).
Several strategies have been attempted in order to accomplish ligands with high affinity and selectivity for specific integrins. For instance, phage libraries expressing RGD-containing peptides have been used in panning experiments. The outcome of such experiments is however dependent on limitations of the sequence coverage in the phage library. Moreover, epitopes that promote cell adherence might be missed when using a selection method that is based on inhibition of binding to coated integrins by peptides in solution. The interaction between a cell and the surrounding ECM is a crosstalk where initial binding causes intracellular signaling resulting in integrin activation and conformational changes that affects the affinity to the ligand. Thus, a cell-free system with coated integrins might miss the peptides with highest affinity to the activated form of the integrin. Ivanov, B. et al., Bioconjugate Chem. 6: 269-277 (1995) and Koivunen E. et al., Biotechnology 13(3): 265-270 (1995) disclose various RGD-containing peptides.
Several peptidomimetics and non-peptidic small molecules have been designed and synthesized with the purpose to find potent and selective integrin ligands. Rational design of ligands for certain integrins has been hampered by the lack of determined structures.
In most previous studies the goal has been to obtain a potent inhibitor of a specific integrin binding, for example with the purpose to hinder tumor cell invasion or unwanted angiogenesis. In those cases, a functional integrin binding is not required; rather the goal is a soluble molecule that is a potent integrin antagonist. WO 2013/185027 discloses soluble variants of human fibronectin with integrin antagonist activity, i.e. blocking or reducing activities of integrin, such as cell adhesion.
Despite these advances in the field, there is still a need for new cell scaffolds in the field, in particular since various cell types may have preference for different scaffolds and since there is a need for efficient cell scaffolds for wound healing.
SUMMARY OF THE INVENTION
It is an object of the present invention to provide proteins and a cell scaffold that promotes proliferation, differentiation and migration of cells, in particular primary cells.
It is in particular an object of the present invention to provide proteins and a cell scaffold which support proliferation, differentiation and migration of keratinocytes.
It is a further object of the present invention to achieve increased cell adhesion efficacy to a cell scaffold.
It is in particular an object of the present invention to provide proteins and a cell scaffold which provides early attachment of adherent cells.
It is also an object of the present invention to provide proteins and a cell scaffold that are useful for efficient expansion of adherent cells in vitro.
It is also an object of the present invention to provide proteins and a cell scaffold that are useful for transferring cells as a cell sheet, e.g. to a wound area in vivo.
Is it an object of the present invention to provide proteins and a cell scaffold that attract inherent cells for migration into a wound area, e.g. from the wound edges from where dermal keratinocytes are usually recruited during wound healing.
For these and other objects that will be evident from the following disclosure, the present invention provides a cyclic RGD cell-binding motif comprising the amino acid sequence C.sup.1X.sup.1X.sup.2RGDX.sup.3X.sup.4X.sup.5C.sup.2 wherein each of X.sup.1, X.sup.2, X.sup.3, X.sup.4 and X.sup.5 are independently selected from natural amino acid residues other than cysteine; and C.sup.1 and C.sup.2 are connected via a disulphide bond. The cell-binding motif has selectivity for integrins, such as for .alpha.5.beta.1 integrins.
It has surprisingly been found that recombinant proteins containing this cyclic RGD cell-binding motif are useful for the cultivation of cells displaying integrins on their cell surface.
Without limitation thereto, preferred cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines.
Without wishing to be bound to any specific theory, it is contemplated that the cell-binding motif presented herein imitates the .alpha.5.beta.1-specific RGD loop motif of fibronectin by positioning cysteines adjacent to the RGD sequence to allow formation of a disulphide-bridge to constrain the chain into a similar type of turn loop. This cyclic RGD cell-binding motif increases the cell adhesion efficacy to a matrix made of a protein containing the cell-binding motif, such as a recombinantly produced spider silk protein.
The present invention provides according to an aspect a recombinant protein comprising said cell-binding motif with selectivity for integrins, such as for .alpha.5.beta.1 integrins. This recombinant protein is surprisingly useful for the cultivation of cells displaying integrins on their cell surface.
The present invention provides according to a one aspect a recombinant fusion protein comprising a spidroin fragment and said cell-binding motif with selectivity for integrins, such as for .alpha.5.beta.1 integrins. This recombinant fusion protein is surprisingly useful for the cultivation of cells displaying integrins on their cell surface.
In preferred embodiments of the invention, each of X.sup.1, X.sup.2, X.sup.3, X.sup.4 and X.sup.5 are independently selected from the group of amino acid residues consisting of: G, A, V, S, T, D, E, M, P, N and Q.
In other preferred embodiments of the invention, each of X.sup.1 and X.sup.3 are independently selected from the group of amino acid residues consisting of: G, S, T, M, N and Q; and each of X.sup.2, X.sup.4 and X.sup.5 are independently selected from the group of amino acid residues consisting of: G, A, V, S, T, P, N and Q.
In certain preferred embodiments of the invention, X.sup.1 is selected from the group of amino acid residues consisting of: G, S, T, N and Q; X.sup.3 is selected from the group of amino acid residues consisting of: S, T and Q; and each of X.sup.2, X.sup.4 and X.sup.5 are independently selected from the group of amino acid residues consisting of: G, A, V, S, T, P and N.
In some preferred embodiments of the invention, X.sup.1 is S or T; X.sup.2 is G, A or V; preferably G or A; more preferably G; X.sup.3 is S or T; preferably S; X.sup.4 is G, A, V or P; preferably G or P; more preferably P; and X.sup.5 is G, A or V; preferably G or A; more preferably A.
In certain preferred embodiments of the invention, the cell-binding motif is comprising the amino acid sequence CTGRGDSPAC (SEQ ID NO: 10).
Further preferred cyclic RGD cell-binding motifs according to the invention display at least 60%, such as at least 70%, such as at least 80%, such as at least 90% identity to CTGRGDSPAC (SEQ ID NO: 10), with the proviso that position 1 and 10 are always C; position 4 is always R; position 5 is always G; position 6 is always D; and positions 2-3 and 7-9 are never cysteine. It is understood that the non-identical positions among positions 2-3 and 7-9 can be freely selected as set out above.
In some preferred fusion proteins according to the invention, the cell-binding motif is arranged N-terminally of the spidroin fragment.
In certain preferred fusion proteins according to the invention, the spidroin fragment is comprising the protein moieties REP and CT, wherein
REP is a repetitive fragment of from 70 to 300 amino acid residues, selected from the group consisting of L(AG).sub.nL, L(AG).sub.nAL, L(GA).sub.nL, and L(GA).sub.nGL, wherein n is an integer from 2 to 10; each individual A segment is an amino acid sequence of from 8 to 18 amino acid residues, wherein from 0 to 3 of the amino acid residues are not Ala, and the remaining amino acid residues are Ala; each individual G segment is an amino acid sequence of from 12 to 30 amino acid residues, wherein at least 40% of the amino acid residues are Gly; and each individual L segment is a linker amino acid sequence of from 0 to 30 amino acid residues; and
CT is a fragment of from 70 to 120 amino acid residues, having at least 70% identity to SEQ ID NO: 3.
In some preferred fusion proteins according to the invention, the spidroin fragment has at least 70% identity to SEQ ID NO: 2 or to amino acid residues 18-277 of SEQ ID NO: 13.
According to a further aspect, the present invention provides a cell scaffold material comprising a protein polymer which as a repeating unit is containing the recombinant fusion protein according to the invention.
In a preferred embodiment of the cell scaffold material according to the invention, the protein polymer is in a physical form selected from the group consisting of film, coating, foam, fiber and fiber-mesh.
In one preferred embodiment of the cell scaffold material according to the invention, the protein polymer is in a physical form of a free-standing matrix.
According to a related aspect, the present invention provides a method for the cultivation of cells, comprising the steps of providing a sample of cells; applying the sample to a cell scaffold material; and maintaining the cell scaffold material having the cells applied thereto under conditions suitable for cell culture; wherein the cell scaffold material comprises a protein polymer, which is containing the recombinant protein, such as the recombinant fusion protein, according to the invention as a repeating structural unit.
It has surprisingly been found that recombinant proteins containing this cyclic RGD cell-binding motif are useful for the cultivation of cells displaying integrins on their cell surface. Without limitation thereto, preferred cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines.
According to a further aspect, the present invention provides use of the recombinant fusion protein according to the invention, the cell scaffold material according to the invention, or the recombinant protein according to the invention for the cultivation of cells displaying integrins on their cell surface.
It has surprisingly been found that recombinant proteins, such as recombinant fusion proteins, containing this cyclic RGD cell-binding motif are useful for the cultivation of cells displaying integrins on their cell surface. The immobilized (i.e. not in solution) cell-binding motif promotes integrin activation and cell binding.
Without limitation thereto, preferred cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines.
In preferred embodiments of the method or the use according to the invention, the cells are displaying .alpha.5.beta.1 integrins on their cell surface; and the cell-binding motif of the recombinant fusion protein has selectivity for .alpha.5.beta.1 integrins.
BRIEF DESCRIPTION OF THE DRAWINGS
FIG. 1 illustrates silk constructs with cell binding motifs derived from fibronectin.
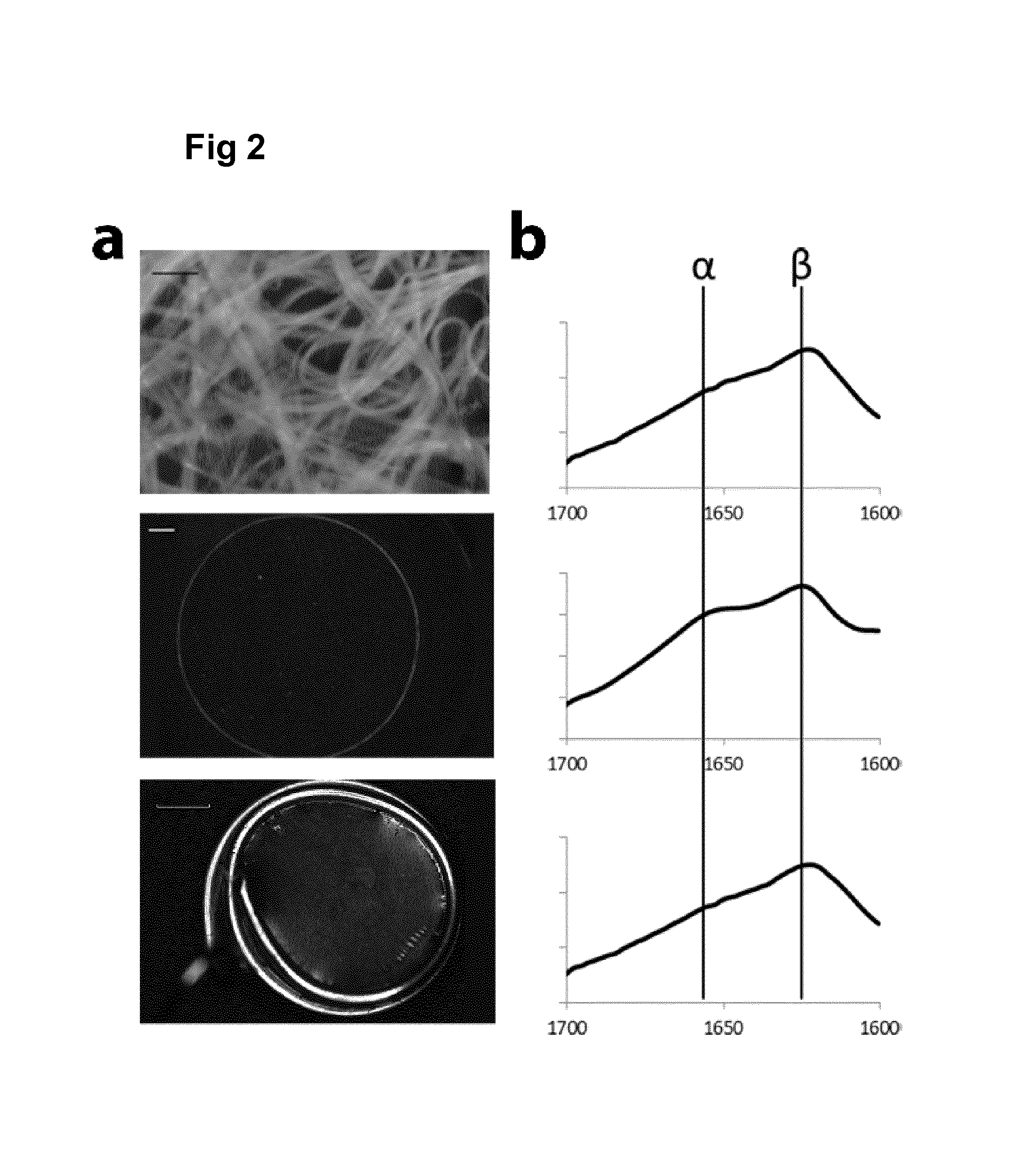
FIG. 2 shows micrographs and FTIR spectra of FN.sub.CC silk (SEQ ID NO: 13) matrices.
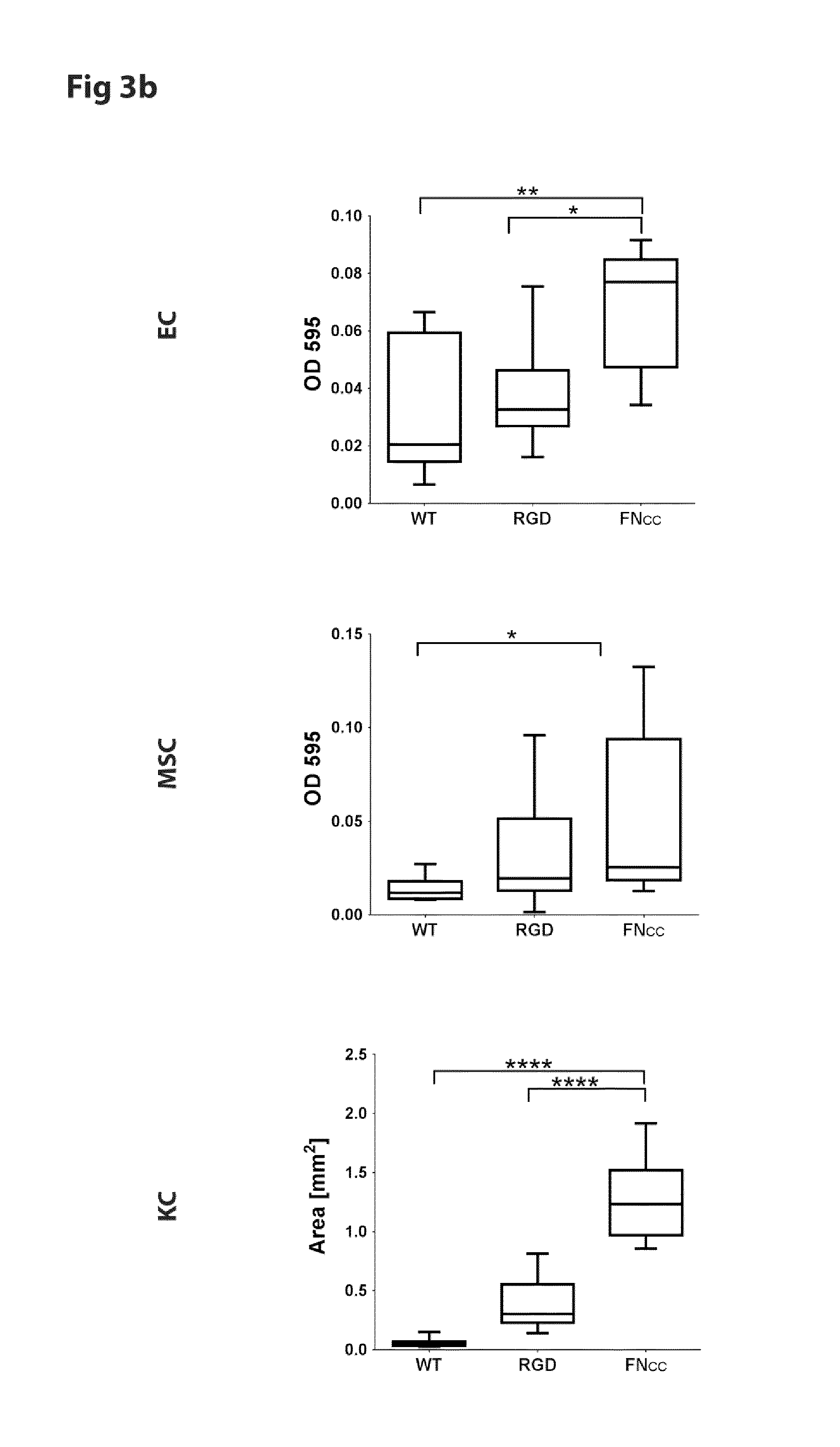
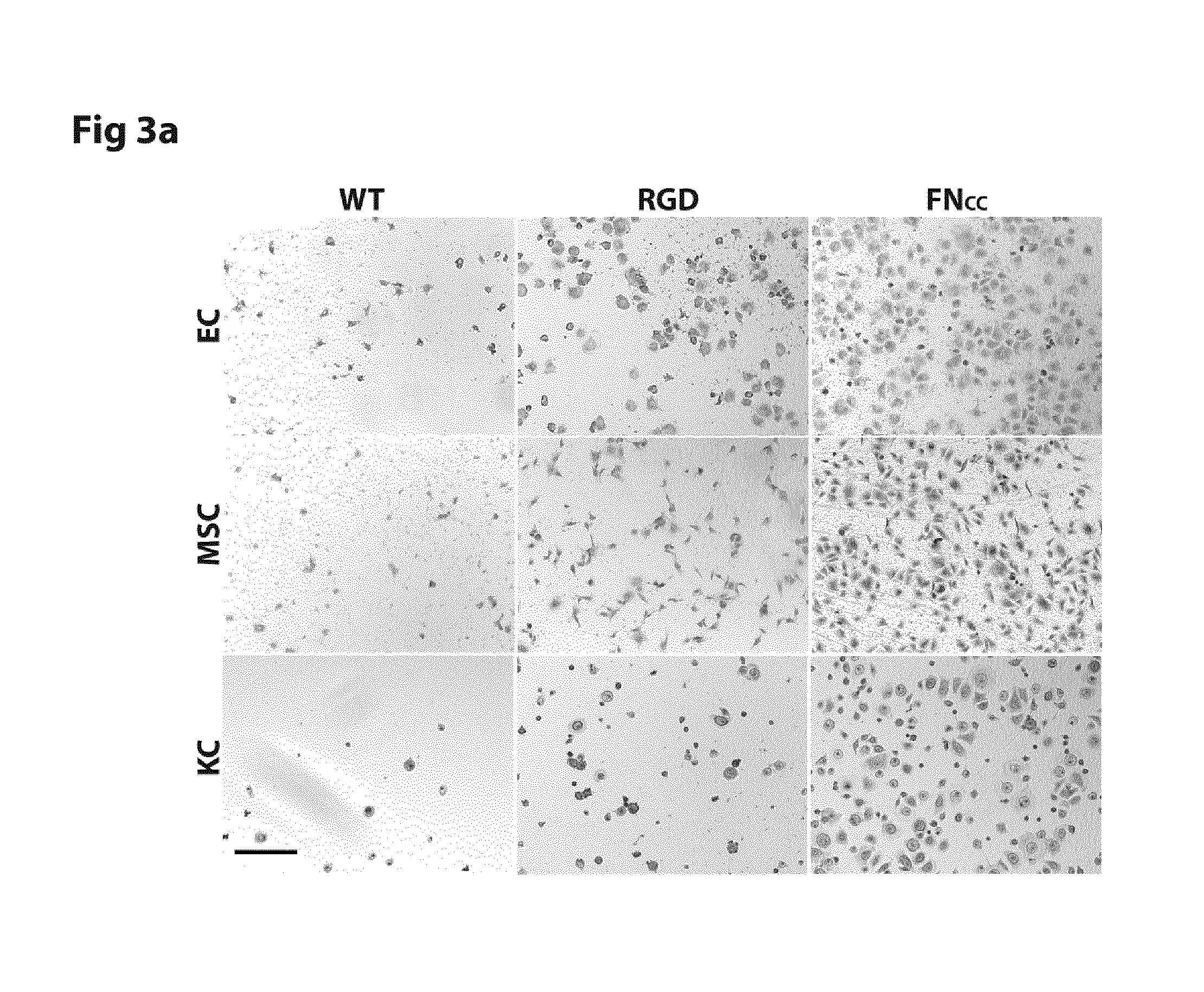
FIG. 3 shows micrographs and coverage density of endothelial cells (EC), mesenchymal stem cells (MSC) and keratinocytes (KC) after 1 h adhesion to film of WT silk (SEQ ID NO: 2) or silk functionalized with RGD (SEQ ID NO: 16) or FN.sub.CC (SEQ ID NO: 13).
FIG. 4 shows micrographs and cell coverage area of keratinocytes (KC) after 1 h adhesion to either silk functionalized with FN.sub.CC (SEQ ID NO: 13), a bovine fibronectin coated surface (BFN) or tissue culture treated cell plastic (TCT).
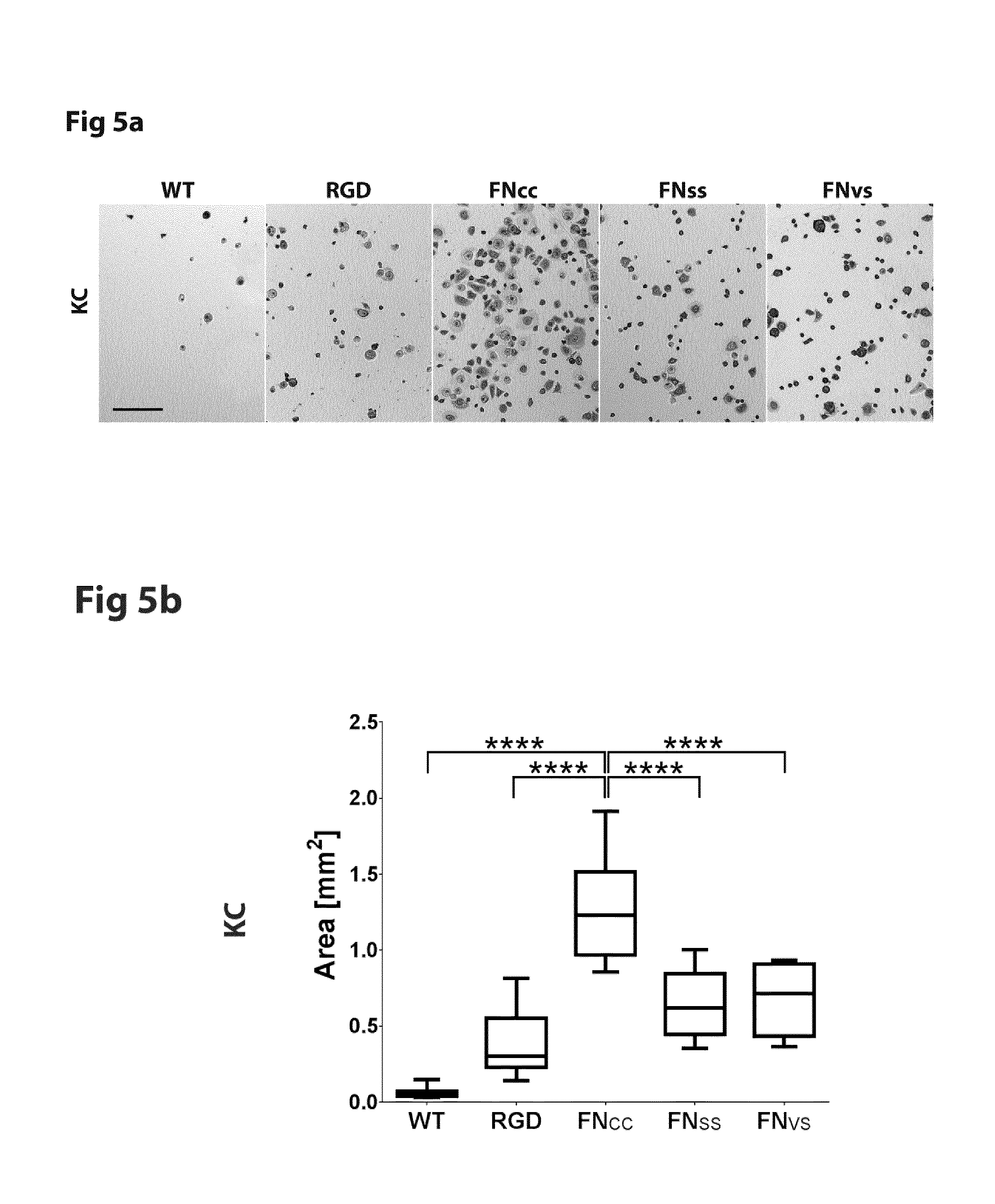
FIG. 5 shows micrographs and cell coverage area of keratinocytes (KC) after 1 h adhesion to WT-silk (SEQ ID NO: 2) or silk functionalized with FN.sub.CC (SEQ ID NO: 13), FN.sub.VS (SEQ ID NO: 15), FN.sub.SS (SEQ ID NO: 14) or RGD (SEQ ID NO: 16).
FIG. 6 shows cell coverage area and stress fiber ranking of keratinocytes (KC) after 3 h adhesion to WT-silk (SEQ ID NO: 2) or silk functionalized with FN.sub.CC (SEQ ID NO: 13), FN.sub.VS (SEQ ID NO: 15), FN.sub.SS (SEQ ID NO: 14) or RGD (SEQ ID NO: 16).
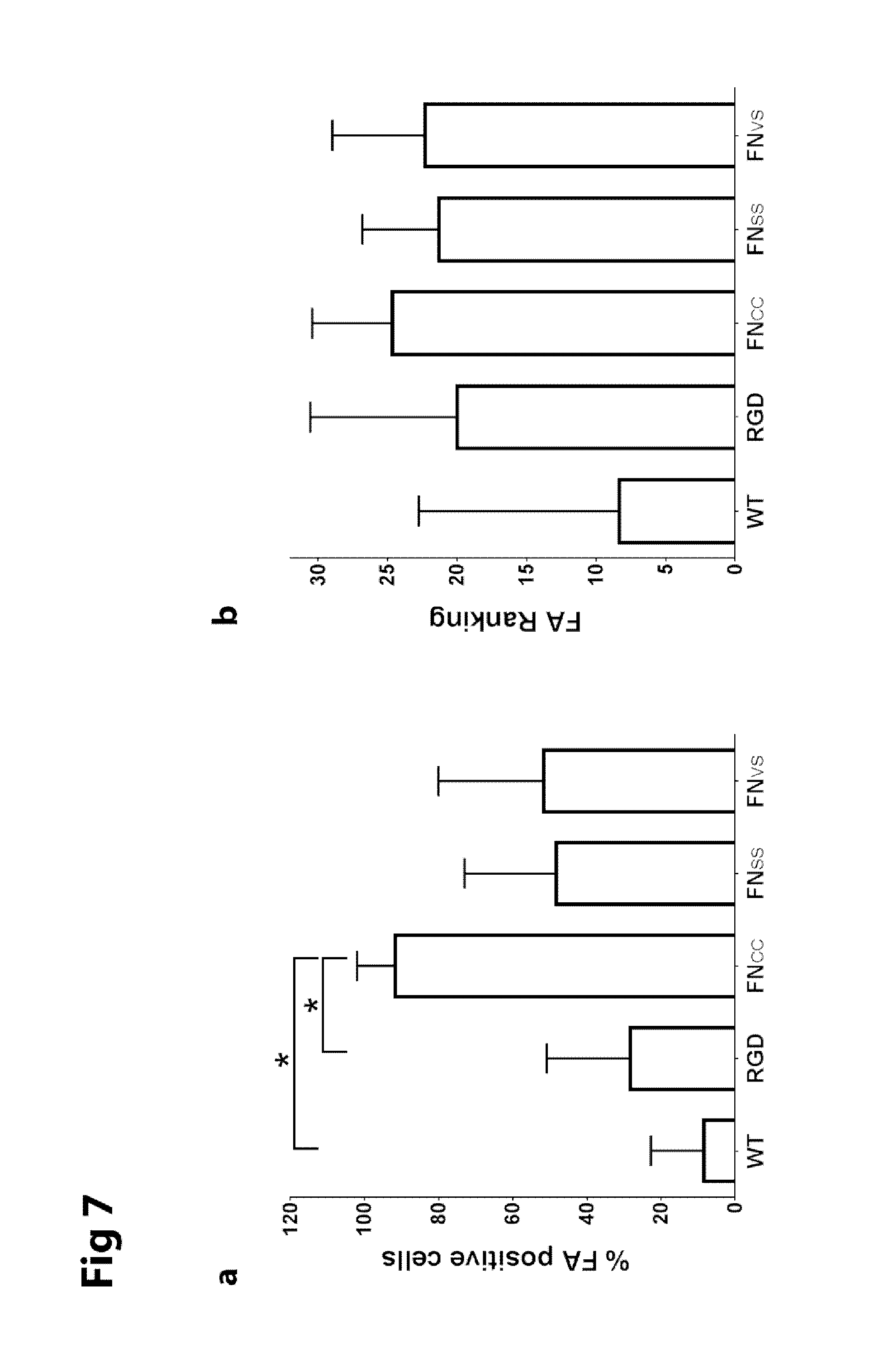
FIG. 7 shows graphs of formation of focal adhesions in keratinocytes after adherence for 3 h onto films of WT-silk (SEQ ID NO: 2) or silk functionalized with FN.sub.CC (SEQ ID NO: 13), FN.sub.VS (SEQ ID NO: 15), FN.sub.SS (SEQ ID NO: 14) or RGD (SEQ ID NO: 16).
FIG. 8 shows a graph of an Alamar blue viability assay of keratinocytes seeded on films of WT-silk (SEQ ID NO: 2) or FN.sub.CC silk (SEQ ID NO: 13).
FIG. 9 shows a sequence alignment of spidroin C-terminal domains.
LIST OF APPENDED SEQUENCES
TABLE-US-00001 SEQ ID NO: 1 RepCT (4RepCT, WT) (DNA) 2 RepCT (4RepCT, WT) 3 CT 4 consensus CT sequence 5 repetitive sequence from Euprosthenops australis MaSp1 6 consensus G segment sequence 1 7 consensus G segment sequence 2 8 consensus G segment sequence 3 9 FN.sub.VS, native fibronectin RGD cell-binding motif 10 FN.sub.CC 11 FN.sub.SS 12 linear RGD cell-binding motif, Widhe et al. (2013)* 13 FN.sub.CC-4RepCT 14 FN.sub.SS-4RepCT 15 FN.sub.VS-4RepCT 16 RGD-4RepCT, Widhe et al. (2013)* 17 FN.sub.CC-4RepCT (DNA) 18 FN.sub.SS-4RepCT (DNA) 19 FN.sub.VS-4RepCT (DNA) 20 RGD-4RepCT, Widhe et al. (2013) (DNA)* 21-24 RGD peptides with glycine spacer 25-28 Linker peptides 29 CT Euprosthenops sp MaSp1 30 CT Euprosthenops australis MaSp1 31 CT Argiope trifasciata MaSp1 32 CT Cyrtophora moluccensis Sp1 33 CT Latrodectus geometricus MaSp1 34 CT Latrodectus hesperus MaSp1 35 CT Macrothele holsti Sp1 36 CT Nephila clavipes MaSp1 37 CT Nephila pilipes MaSp1 38 CT Nephila madagascariensis MaSp1 39 CT Nephila senegalensis MaSp1 40 CT Octonoba varians Sp1 41 CT Psechrus sinensis Sp1 42 CT Tetragnatha kauaiensis MaSp1 43 CT Tetragnatha versicolor MaSp1 44 CT Araneus bicentenarius Sp2 45 CT Argiope amoena MaSp2 46 CT Argiope aurantia MaSp2 47 CT Argiope trifasciata MaSp2 48 CT Gasteracantha mammosa MaSp2 49 CT Latrodectus geometricus MaSp2 50 CT Latrodectus hesperus MaSp2 51 CT Nephila clavipes MaSp2 52 CT Nephila madagascariensis MaSp2 53 CT Nephila senegalensis MaSp2 54 CT Dolomedes tenebrosus Fb1 55 CT Dolomedes tenebrosus Fb2 56 CT Araneus diadematus ADF-1 57 CT Araneus diadematus ADF-2 58 CT Araneus diadematus ADF-3 59 CT Araneus diadematus ADF-4 60 STGRGDSPAV (FN1011) *Widhe M et al., Biomaterials 34(33): 8223-8234 (2013)
DETAILED DESCRIPTION OF THE INVENTION
Recombinantly produced spider silk and numerous other materials are useful as matrices for culture of mammalian cells. The inclusion of cell adhesion motifs derived from the extracellular matrix (ECM) into such materials increases cell attachment and proliferation by interaction with integrins on the cell surface. The integrins do not just confer the physical connection between cells and the surrounding, but also mediate signals controlling for example cell growth, polarity, proliferation and survival. Moreover, the integrins are essential for cell migration by acting as the cells' "feet".
The most widely characterized cell adhesion motif is the RGD peptide, first discovered in fibronectin. The RGD motif is found also in many other molecules of the natural ECM, for example in vitronectin, fibrinogen and in cryptic sites of both collagen I and several of the laminin .alpha. chains. Almost half of the known integrins, including .alpha.3.beta.1, .alpha.5.beta.1, .alpha.8.beta.1, .alpha.v.beta.1, .alpha.IIb.beta.3, .alpha.v.beta.3, .alpha.v.beta.5, .alpha.v.beta.6a and .alpha.v.beta.8, have been shown to bind ECM in a RGD-dependent manner. However, after initial proofs of RGD as general cell adhesion motif, it soon became clear that integrins in general bind with magnitudes higher affinity to larger RGD containing proteins than to short RGD peptides. The preferred conditions for binding also seem to vary between different integrins.
The present invention is based on a designed cell-binding motif. Without wishing to be bound to any specific theory, it is contemplated that the cell-binding motif presented herein imitates the .alpha.5.beta.1-specific RGD loop motif of fibronectin by positioning cysteines in precise positions adjacent to the RGD sequence to allow formation of a disulphide-bridge to constrain the chain into a similar type of turn loop. This cyclic RGD cell-binding motif increases the cell adhesion efficacy to a matrix made of a protein containing the cell-binding motif, such as a recombinantly produced spider silk protein or a synthetic peptide.
The term "cyclic" as used herein refers to a peptide wherein two amino acid residues are covalently bonded via their side chains, more specifically through a disulfide bond between two cysteine residues.
It is shown herein that the cell adhesive properties of a material is significantly enhanced by introducing the cyclic RGD cell-binding motif on a cysteine linked loop compared to when a linear RGD peptide is added. In addition, the cyclic RGD cell-binding motif presented herein promotes both proliferation of and migration by primary cells. Human primary cells cultured on a cell scaffold material containing the cyclic RGD cell-binding motif showed increased attachment, spreading, stress fiber formation and focal adhesions compared to the same material containing a linear RGD peptide.
The cyclic RGD cell-binding motif presented herein is also suitable for preparing free-standing matrices, in particular matrices containing spider silk, on which cells could readily form a monolayer culture. Such free-standing matrices are useful for cell sheet transfer. Thus, a material containing the cyclic RGD cell-binding motif presented herein, such as a spider silk material, is useful for both an in vitro setting, where adherent cells need to be expanded efficiently, and in an in vivo situation where cells need to be transferred as a cell sheet to e.g. a wound area. The results also support that a material containing the cyclic RGD cell-binding motif presented herein, such as a spider silk material, can efficiently attract inherent cells for migration into a wound area, e.g. from the wound edges from where dermal keratinocytes are usually recruited during wound healing. Cell binding to a cell scaffold containing the cyclic RGD cell-binding motif presented herein is demonstrated to involve the .alpha.5.beta.1 integrin, and to support proliferation and migration of keratinocytes.
The present inventor used DNA technology to modify the cell-binding motif of fibronectin, where the RGD motif is presented on a turn loop. This was accomplished with the amino acid sequence flanking RGD in the tenth type III domain of fibronectin as base (FIG. 1b). Firstly, the same decapeptide (VTGRGDSPAS; SEQ ID NO: 9) as in the turn loop of fibronectin was introduced N-terminally to a protein to yield a construct denoted FN.sub.VS (FIG. 1a). Without wishing to be bound to any specific theory, it was hypothezised that the cell-binding motif could be made more efficient by positioning the valine and serine residue situated 3 positions before and 4 positions after the RGD motif respectively, spatially very close to each other. The present inventor therefore mutated these two residues to cysteines (FIG. 1a, c), so that the RGD containing motif is flanked by one cysteine on each side. The cysteines are spatially less than 2 .ANG. apart, and thus connect the peptide chain into a disulphide bridged loop (denoted FN.sub.CC; SEQ ID NO: 10). As control, a variant with the two cysteines exchanged to serines was also constructed (denoted FN.sub.SS; SEQ ID NO: 11). The present inventor investigated the effect of these FN motifs, when introduced into protein matrices, on various mechanisms of early attachment (including spreading, stress fiber formation, focal adhesions and integrin binding) in primary adherent cells of human origin. It was found that the FN.sub.CC variant containing a cyclic RGD cell-binding motif increases the cell adhesion efficacy to a matrix made of a protein containing the cell-binding motif as compared to the controls FN.sub.VS and FN.sub.SS.
It can be seen from the crystal structure of the ninth and tenth domain of fibronectin determined by Leahy D J et al., Cell 84(1): 155-164 (1996), that the valine and serine residue situated 3 positions before and 4 positions after the RGD motif respectively, are located spatially very close to each other (FIG. 1c). Again without wishing to be bound to any specific theory, it is therefore considered that the cell-binding motif presented herein imitates the .alpha.5.beta.1-specific RGD loop motif of fibronectin by positioning cysteines adjacent to the RGD sequence to allow formation of a disulphide-bridge to constrain the chain into a similar type of turn loop. As a consequence, it is concluded that the cell-binding motif presented herein is in particular selective for .alpha.5.beta.1 integrins.
Thus, the relevant silk constructs with cell binding motifs derived from fibronectin are illustrated in FIG. 1. FIG. 1a schematically shows the silk protein 4RepCT with different RGD motifs genetically introduced to its N-terminus. "RGD" in FIG. 1a denotes the RGD containing peptide (SEQ ID NO 12) used in Widhe M et al., Biomaterials 34(33): 8223-8234 (2013). "FN.sub.VS" denotes the RGD-containing decapeptide from fibronectin (SEQ ID NO: 9). "FN.sub.CC" denotes the same peptide with V and S exchanged to C (SEQ ID NO: 10). "FN.sub.SS" denotes the same peptide with V and S exchanged to S (SEQ ID NO: 11). FIG. 1b shows the structure of the 9th and 10th domain of fibronectin, displaying the turn loop containing the RGD motif (SEQ ID NO: 60). FIG. 1c shows a structure model of the RGD loop taken from fibronectin, with the residues V and S mutated to C (adapted from 1FNF.pdb).
The cell-binding motif presented herein is selective for binding to integrins presented on the cell surface, such as and preferably to .alpha.5.beta.1 integrins. In the context of the present invention, "specific" or "selective" interaction of the cell-binding motif with its target integrin means that the interaction is such that a distinction between specific and non-specific, or between selective and non-selective, interaction becomes meaningful. The interaction between two proteins is sometimes measured by the dissociation constant. The dissociation constant describes the strength of binding (or affinity) between two molecules. Typically the dissociation constant between an antibody and its antigen is from 10.sup.-7 to 10.sup.-11 M. However, high specificity does not necessarily require high affinity. Molecules with low affinity (in the molar range) for its counterpart have been shown to be as specific as molecules with much higher affinity. In the case of the present invention, a specific or selective interaction refers to the extent to which a particular method can be used to preferentially bind to a specific protein or cell type, displaying the target integrin or a fragment thereof, under given conditions in the presence of other proteins or cells in a sample of a naturally occurring or processed biological or biochemical fluid. In other words, specificity or selectivity is the capacity to distinguish between related proteins and cell types displaying the related proteins. Specific and selective are sometimes used interchangeably in the present description.
The cyclic RGD cell-binding motif is comprising, or consisting of, the amino acid sequence C.sup.1X.sup.1X.sup.2RGDX.sup.3X.sup.4X.sup.5C.sup.2 wherein each of X.sup.1, X.sup.2, X.sup.3, X.sup.4 and X.sup.5 are independently selected from natural amino acid residues other than cysteine; and C.sup.1 and C.sup.2 are connected via a disulphide bond.
It is preferred that each of X.sup.1, X.sup.2, X.sup.3, X.sup.4 and X.sup.5 are independently selected from the group of amino acid residues consisting of: G, A, V, S, T, D, E, M, P, N and Q.
It is more preferred that each of X.sup.1 and X.sup.3 are independently selected from the group of amino acid residues consisting of: G, S, T, M, N and Q; and that each of X.sup.2, X.sup.4 and X.sup.5 are independently selected from the group of amino acid residues consisting of: G, A, V, S, T, P, N and Q. The resulting cell-binding motif does not contain any charged or bulky residues which could be disadvantageous for the cell-binding efficacy.
It is in particular preferred that: X.sup.1 is selected from the group of amino acid residues consisting of: G, S, T, N and Q; X.sup.3 is selected from the group of amino acid residues consisting of: S, T and Q; and each of X.sup.2, X.sup.4 and X.sup.5 are independently selected from the group of amino acid residues consisting of: G, A, V, S, T, P and N.
It is more preferred that X.sup.1 is S or T; X.sup.2 is G, A or V; preferably G or A; more preferably G; X.sup.3 is S or T; preferably S; X.sup.4 is G, A, V or P; preferably G or P; more preferably P; and X.sup.5 is G, A or V; preferably G or A; more preferably A.
A particularly preferred cyclic RGD cell-binding motif is comprising, or consisting of, the amino acid sequence CTGRGDSPAC (FN.sub.CC; SEQ ID NO: 10).
Further preferred cyclic RGD cell-binding motifs according to the invention display at least 60%, such as at least 70%, such as at least 80%, such as at least 90% identity to CTGRGDSPAC (FN.sub.CC; SEQ ID NO: 10), with the proviso that position 1 and 10 are always C; position 4 is always R; position 5 is always G; position 6 is always D; and positions 2-3 and 7-9 are never cysteine. It is understood that the non-identical positions among positions 2-3 and 7-9 can be freely selected as set out above.
The thus identified cyclic RGD cell-binding motif is useful in any recombinant or synthetic protein or peptide so as to provide selective binding to integrins, in particular .alpha.5.beta.1 integrins. Thus, there is provided a recombinant protein comprising the cell-binding motif with selectivity for integrins, such as for .alpha.5.beta.1 integrins. The recombinant protein is useful for the cultivation of cells, e.g. mammalian cells, displaying integrins, in particular .alpha.5.beta.1 integrins, on their cell surface.
Without limitation thereto, preferred cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines.
Fibronectin is recognized by at least ten of the cell surface receptors of the integrin family, among which five (.alpha.3.beta.1, .alpha.4.beta.1, .alpha.5.beta.1, .alpha.8.beta.1, .alpha.v.beta.1) include the .beta.1 subunit. The .alpha.5 subunit is found only in combination with .beta.1 and the .alpha.5.beta.1 integrin is unique since it is specialized for binding of fibronectin only, and therefore originally denoted the fibronectin receptor. The specific interaction between .alpha.5.beta.1 and fibronectin seem to be fundamental for vertebrate development since lack of either .alpha.5.beta.1 or fibronectin results in early embryonic lethality. Fibronectin and .alpha.5.beta.1 has also been shown important in the wound repair process of airway epithelium, where both have been observed to be exclusively expressed by the migratory cells in the wounded area, and to play a critical role in endothelial cell migration in vitro and angiogenesis in vivo.
There is provided a recombinant or synthetic protein or peptide comprising a cell-binding motif with selectivity for integrins, such as for .alpha.5.beta.1 integrins, wherein the cell-binding motif is as set out above.
A preferred recombinant protein is comprising a cell-binding motif with selectivity for integrins, such as for .alpha.5.beta.1 integrins, wherein the cell-binding motif has the amino acid sequence C.sup.1X.sup.1X.sup.2RGDX.sup.3X.sup.4X.sup.5C.sup.2 wherein X.sup.1 is selected from the group of amino acid residues consisting of: G, S, T, N and Q; X.sup.3 is selected from the group of amino acid residues consisting of: S, T and Q; and each of X.sup.2, X.sup.4 and X.sup.5 are independently selected from the group of amino acid residues consisting of: G, A, V, S, T, P and N; and C.sup.1 and C.sup.2 are connected via a disulphide bond.
Preferred embodiments of the cell-binding motif is presented herein. In particular, it is preferred that:
X.sup.1 is S or T; preferably T;
X.sup.2 is G, A or V; preferably G or A; more preferably G;
X.sup.3 is S or T; preferably S
X.sup.4 is G, A, V or P; preferably G or P; more preferably P;
X.sup.5 is G, A or V; preferably G or A; more preferably A.
A specific preferred cell-binding motif is comprising the amino acid sequence CTGRGDSPAC (FN.sub.CC; SEQ ID NO: 10).
The recombinant protein is useful in cell scaffold materials. It is also useful for the cultivation of cells displaying integrins on their cell surface, in particular, wherein the cells are displaying .alpha.5.beta.1 integrins on their cell surface.
Without limitation thereto, preferred cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines.
The recombinant or synthetic protein may also be constituted by a shorter peptide comprising or even consisting of the cell-binding motif, e.g. containing 10-50, or 10-30 amino acid residues. These peptides may be chemically coupled or immobilized to a surface as is well-known in the art. Advantageously, the peptide contains or is coupled to a spacer which allows greater accessibility to the cell-binding motif. The thus immobilized (i.e. not in solution) recombinant protein is surprisingly useful for the cultivation of cells displaying integrins on their cell surface, in particular, wherein the cells are displaying .alpha.5.beta.1 integrins on their cell surface.
The cell-binding motif is advantageously presented as part of a fusion protein together with a spider silk protein, in particular a miniature spider silk protein. The terms "spidroins" and "spider silk proteins" are used interchangeably throughout the description and encompass all known spider silk proteins, including major ampullate spider silk proteins which typically are abbreviated "MaSp", or "ADF" in the case of Araneus diadematus. These major ampullate spider silk proteins are generally of two types, 1 and 2. These terms furthermore include non-natural proteins with a high degree of identity and/or similarity to the known spider silk proteins.
There is provided a recombinant fusion protein comprising a spidroin fragment and the cell-binding motif with selectivity for integrins, such as for .alpha.5.beta.1 integrins, set out above. The spidroin fragment is preferably comprising, or consisting of, the protein moieties REP and CT, wherein REP is a repetitive fragment of from 70 to 300 amino acid residues, selected from the group consisting of L(AG).sub.nL, L(AG).sub.nAL, L(GA).sub.nL, and L(GA).sub.nGL, wherein n is an integer from 2 to 10; each individual A segment is an amino acid sequence of from 8 to 18 amino acid residues, wherein from 0 to 3 of the amino acid residues are not Ala, and the remaining amino acid residues are Ala; each individual G segment is an amino acid sequence of from 12 to 30 amino acid residues, wherein at least 40% of the amino acid residues are Gly; and each individual L segment is a linker amino acid sequence of from 0 to 30 amino acid residues; and
CT is a fragment of from 70 to 120 amino acid residues, having at least 70% identity to SEQ ID NO: 3.
The fusion protein according to the invention harbors both a desired selective cell-binding activity in the cell-binding motif and an internal solid support activity in the spidroin fragment. The binding activity of the fusion protein is maintained when it is structurally rearranged to form polymeric, solid structures. These protein structures, or protein polymers, also provides a high and predictable density of the cell-binding motif with selective interaction activity towards integrins, e.g. .alpha.5.beta.1 integrins. The thus immobilized cell-binding motif promotes integrin activation and cell binding. The way biomaterials functionalized with RGD stimulate different cell responses is not only affected by the type of RGD motif used, but also the resulting surface concentrations of ligands. Since the rather small silk proteins used in the present study self-assemble into multilayers where each molecule carries an RGD motif, a dense surface presentation is expected. However, if a more sparse surface concentration is desired, any possible surface density can be achieved simply by mixing silk proteins with and without the cyclic RGD cell-binding motif disclosed herein at different ratios, thereby directing the cellular response of interest.
In most of the proteins that have been engineered to contain RGD, the motif has been added as a linear extension either to the N- or C-terminus, thus with a high possibility of exposure and flexibility due to minimal constrain of the chain from the rest of the protein. Several constructs with the RGD motif placed within a protein fold have been made to reduce the flexibility of the RGD motif, but at the same time also reducing its exposure. The cyclic RGD cell-binding motif disclosed herein can advantageously be presented as a linear extension either to the N- or C-terminus, thus with a high possibility of exposure. At the same time, its cyclic properties limit the flexibility and is believed to contribute to highly useful cell binding properties. Furthermore, the covalent incorporation of the peptide into a folded protein chain might have contributed to the apparently efficient integrin-mediated cell binding, involving .alpha.5.beta.1.
The term "fusion protein" implies here a protein that is made by expression from a recombinant nucleic acid, i.e. DNA or RNA that is created artificially by combining two or more nucleic acid sequences that would not normally occur together (genetic engineering). The fusion proteins according to the invention are recombinant proteins, and they are therefore not identical to naturally occurring proteins. In particular, wildtype spidroins are not fusion proteins according to the invention, because they are not expressed from a recombinant nucleic acid as set out above. The combined nucleic acid sequences encode different proteins, partial proteins or polypeptides with certain functional properties. The resulting fusion protein, or recombinant fusion protein, is a single protein with functional properties derived from each of the original proteins, partial proteins or polypeptides. Furthermore, the fusion protein according to the invention and the corresponding genes are chimeric, i.e. the protein/gene moieties are derived from at least two different species.
The fusion protein typically consists of from 170 to 2000 amino acid residues, such as from 170 to 1000 amino acid residues, such as from 170 to 600 amino acid residues, preferably from 170 to 500 amino acid residues, such as from 170 to 400 amino acid residues. The small size is advantageous because longer proteins containing spider silk protein fragments may form amorphous aggregates, which require use of harsh solvents for solubilisation and polymerisation.
The fusion protein may contain one or more linker peptides, or L segments. The linker peptide(s) may be arranged between any moieties of the fusion protein, e.g. between the REP and CT moieties, at either terminal end of the fusion protein or between the spidroin fragment and the cell-binding motif. The linker(s) may provide a spacer between the functional units of the fusion protein, but may also constitute a handle for identification and purification of the fusion protein, e.g. a His and/or a Trx tag. If the fusion protein contains two or more linker peptides for identification and purification of the fusion protein, it is preferred that they are separated by a spacer sequence, e.g. His.sub.6-spacer-His.sub.6-. The linker may also constitute a signal peptide, such as a signal recognition particle, which directs the fusion protein to the membrane and/or causes secretion of the fusion protein from the host cell into the surrounding medium. The fusion protein may also include a cleavage site in its amino acid sequence, which allows for cleavage and removal of the linker(s) and/or other relevant moieties. Various cleavage sites are known to the person skilled in the art, e.g. cleavage sites for chemical agents, such as CNBr after Met residues and hydroxylamine between Asn-Gly residues, cleavage sites for proteases, such as thrombin or protease 3C, and self-splicing sequences, such as intein self-splicing sequences.
The spidroin fragment and the cell-binding motif are linked directly or indirectly to one another. A direct linkage implies a direct covalent binding between the moieties without intervening sequences, such as linkers. An indirect linkage also implies that the moieties are linked by covalent bonds, but that there are intervening sequences, such as linkers and/or one or more further moieties, e.g. 1-2 NT moieties.
The cell-binding motif may be arranged internally or at either end of the fusion protein, i.e. C-terminally arranged or N-terminally arranged. It is preferred that the cell-binding motif is arranged at the N-terminal end of the fusion protein. If the fusion protein contains one or more linker peptide(s) for identification and purification of the fusion protein, e.g. a His or Trx tag(s), it is preferred that it is arranged at the N-terminal end of the fusion protein.
A preferred fusion protein has the form of an N-terminally arranged cell-bonding motif, coupled by a linker peptide of 0-30 amino acid residues, such as 0-10 amino acid residues, to a REP moiety. Optionally, the fusion protein has an N-terminal or C-terminal linker peptide, which may contain a purification tag, such as a His tag, and a cleavage site.
The recombinant protein is useful in cell scaffold materials. It is also useful for the cultivation of cells displaying integrins on their cell surface, in particular wherein the cells are displaying .alpha.5.beta.1 integrins on their cell surface.
Without limitation thereto, preferred cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines.
Without wishing to be bound to any specific theory, it is contemplated that the cell-binding motif is functionally displayed on the surface of the resulting cell scaffold material, which is herein surprisingly shown to be advantageous for the binding capacity with respect to mammalian cells, c.f. Examples 6-9.
The prominent positive effect of the cell scaffold material containing the cyclic RGD cell-binding motif presented herein is evident already at initial attachment (within 0.5-3 h) of primary cells. Strong and rapid attachment of cells onto a material has been suggested to be of considerable importance when it comes to various clinical applications, where the present environment for cells is far from optimal, and fast establishment is necessary for cell survival. One example is the stressful milieu of a chronic wound, often with high bacterial load and necrosis. Here, migrating keratinocytes might benefit from the support of a suitably designed biomaterial constituting containing the cyclic RGD cell-binding motif, such as as a spider silk fusion protein. Also in clinical settings where the close surroundings imply physical stress, like velocity of passing fluids, e.g. blood passing the stent in a heart or a vessel implant, a material that facilitates for the endothelial cells to rapidly and firmly attach to an implant could be critical, and thus even decisive for a successful outcome.
A scaffold intended for tissue engineering will obviously be subjected to harsher handling and environments than in a cell culture setting, why the observed improved stability of the spider silk material containing the cyclic RGD cell-binding motif is valuable. This increase in stability compared to the wild type silk allows preparation of transferable scaffolds, e.g. free-standing films as demonstrated herein.
The protein moiety REP is fragment with a repetitive character, alternating between alanine-rich stretches and glycine-rich stretches. The REP fragment generally contains more than 70, such as more than 140, and less than 300, preferably less than 240, such as less than 200, amino acid residues, and can itself be divided into several L (linker) segments, A (alanine-rich) segments and G (glycine-rich) segments, as will be explained in more detail below. Typically, said linker segments, which are optional, are located at the REP fragment terminals, while the remaining segments are in turn alanine-rich and glycine-rich. Thus, the REP fragment can generally have either of the following structures, wherein n is an integer:
L(AG).sub.nL, such as LA.sub.1G.sub.1A.sub.2G.sub.2A.sub.3G.sub.3A.sub.4G.sub.4A.sub.5G.sub.5L;
L(AG).sub.nAL, such as LA.sub.1G.sub.1A.sub.2G.sub.2A.sub.3G.sub.3A.sub.4G.sub.4A.sub.5G.sub.5A.- sub.6L;
L(GA).sub.nL, such as LG.sub.1A.sub.1G.sub.2A.sub.2G.sub.3A.sub.3G.sub.4A.sub.4G.sub.5A.sub.5L; or
L(GA).sub.nGL, such as LG.sub.1A.sub.1G.sub.2A.sub.2G.sub.3A.sub.3G.sub.4A.sub.4G.sub.5A.sub.5G.- sub.6L.
It follows that it is not critical whether an alanine-rich or a glycine-rich segment is adjacent to the N-terminal or C-terminal linker segments. It is preferred that n is an integer from 2 to 10, preferably from 2 to 8, also preferably from 4 to 8, more preferred from 4 to 6, i.e. n=4, n=5 or n=6.
In some embodiments, the alanine content of the REP fragment is above 20%, preferably above 25%, more preferably above 30%, and below 50%, preferably below 40%, more preferably below 35%. It is contemplated that a higher alanine content provides a stiffer and/or stronger and/or less extendible fiber.
In certain embodiments, the REP fragment is void of proline residues, i.e. there are no Pro residues in the REP fragment.
Turning now to the segments that constitute the REP fragment, it is emphasized that each segment is individual, i.e. any two A segments, any two G segments or any two L segments of a specific REP fragment may be identical or may not be identical. Thus, it is not a general feature of the spidroin that each type of segment is identical within a specific REP fragment. Rather, the following disclosure provides the skilled person with guidelines how to design individual segments and gather them into a REP fragment, which is a part of a functional spider silk protein useful in a cell scaffold material.
Each individual A segment is an amino acid sequence having from 8 to 18 amino acid residues. It is preferred that each individual A segment contains from 13 to 15 amino acid residues. It is also possible that a majority, or more than two, of the A segments contain from 13 to 15 amino acid residues, and that a minority, such as one or two, of the A segments contain from 8 to 18 amino acid residues, such as 8-12 or 16-18 amino acid residues. A vast majority of these amino acid residues are alanine residues. More specifically, from 0 to 3 of the amino acid residues are not alanine residues, and the remaining amino acid residues are alanine residues. Thus, all amino acid residues in each individual A segment are alanine residues, with no exception or with the exception of one, two or three amino acid residues, which can be any amino acid. It is preferred that the alanine-replacing amino acid(s) is (are) natural amino acids, preferably individually selected from the group of serine, glutamic acid, cysteine and glycine, more preferably serine. Of course, it is possible that one or more of the A segments are all-alanine segments, while the remaining A segments contain 1-3 non-alanine residues, such as serine, glutamic acid, cysteine or glycine.
In an embodiment, each A segment contains 13-15 amino acid residues, including 10-15 alanine residues and 0-3 non-alanine residues as described above. In a more preferred embodiment, each A segment contains 13-15 amino acid residues, including 12-15 alanine residues and 0-1 non-alanine residues as described above.
It is preferred that each individual A segment has at least 80%, preferably at least 90%, more preferably 95%, most preferably 100% identity to an amino acid sequence selected from the group of amino acid residues 7-19, 43-56, 71-83, 107-120, 135-147, 171-183, 198-211, 235-248, 266-279, 294-306, 330-342, 357-370, 394-406, 421-434, 458-470, 489-502, 517-529, 553-566, 581-594, 618-630, 648-661, 676-688, 712-725, 740-752, 776-789, 804-816, 840-853, 868-880, 904-917, 932-945, 969-981, 999-1013, 1028-1042 and 1060-1073 of SEQ ID NO: 5. Each sequence of this group corresponds to a segment of the naturally occurring sequence of Euprosthenops australis MaSp1 protein, which is deduced from cloning of the corresponding cDNA, see WO2007/078239. Alternatively, each individual A segment has at least 80%, preferably at least 90%, more preferably 95%, most preferably 100% identity to an amino acid sequence selected from the group of amino acid residues 25-36, 55-69, 84-98, 116-129 and 149-158 of SEQ ID NO: 2. Each sequence of this group corresponds to a segment of expressed, non-natural spider silk proteins, which proteins have the capacity to form silk fibers under appropriate conditions. Thus, in certain embodiments of the spidroin, each individual A segment is identical to an amino acid sequence selected from the above-mentioned amino acid segments. Without wishing to be bound by any particular theory, it is envisaged that A segments according to the invention form helical structures or beta sheets.
Furthermore, it has been concluded from experimental data that each individual G segment is an amino acid sequence of from 12 to 30 amino acid residues. It is preferred that each individual G segment consists of from 14 to 23 amino acid residues. At least 40% of the amino acid residues of each G segment are glycine residues. Typically the glycine content of each individual G segment is in the range of 40-60%.
It is preferred that each individual G segment has at least 80%, preferably at least 90%, more preferably 95%, most preferably 100% identity to an amino acid sequence selected from the group of amino acid residues 20-42, 57-70, 84-106, 121-134, 148-170, 184-197, 212-234, 249-265, 280-293, 307-329, 343-356, 371-393, 407-420, 435-457, 471-488, 503-516, 530-552, 567-580, 595-617, 631-647, 662-675, 689-711, 726-739, 753-775, 790-803, 817-839, 854-867, 881-903, 918-931, 946-968, 982-998, 1014-1027, 1043-1059 and 1074-1092 of SEQ ID NO: 5. Each sequence of this group corresponds to a segment of the naturally occurring sequence of Euprosthenops australis MaSp1 protein, which is deduced from cloning of the corresponding cDNA, see WO2007/078239. Alternatively, each individual G segment has at least 80%, preferably at least 90%, more preferably 95%, most preferably 100% identity to an amino acid sequence selected from the group of amino acid residues 1-24, 37-54, 70-83, 99-115 and 130-148 of SEQ ID NO: 2. Each sequence of this group corresponds to a segment of expressed, non-natural spider silk proteins, which proteins have the capacity to form silk fibers under appropriate conditions. Thus, in certain embodiments of the spidroin in the cell scaffold material, each individual G segment is identical to an amino acid sequence selected from the above-mentioned amino acid segments.
In certain embodiments, the first two amino acid residues of each G segment are not -Gln-Gln-.
There are three subtypes of the G segment. This classification is based upon careful analysis of the Euprosthenops australis MaSp1 protein sequence (see WO2007/078239), and the information has been employed and verified in the construction of novel, non-natural spider silk proteins.
The first subtype of the G segment is represented by the amino acid one letter consensus sequence GQG(G/S)QGG(Q/Y)GG (L/Q)GQGGYGQGA GSS (SEQ ID NO: 6). This first, and generally the longest, G segment subtype typically contains 23 amino acid residues, but may contain as little as 17 amino acid residues, and lacks charged residues or contain one charged residue. Thus, it is preferred that this first G segment subtype contains 17-23 amino acid residues, but it is contemplated that it may contain as few as 12 or as many as 30 amino acid residues. Without wishing to be bound by any particular theory, it is envisaged that this subtype forms coil structures or 3.sub.1-helix structures. Representative G segments of this first subtype are amino acid residues 20-42, 84-106, 148-170, 212-234, 307-329, 371-393, 435-457, 530-552, 595-617, 689-711, 753-775, 817-839, 881-903, 946-968, 1043-1059 and 1074-1092 of SEQ ID NO: 5. In certain embodiments, the first two amino acid residues of each G segment of this first subtype according to the invention are not -Gln-Gln-.
The second subtype of the G segment is represented by the amino acid one letter consensus sequence GQGGQGQG(G/R)Y GQG(A/S)G(S/G)S (SEQ ID NO: 7). This second, generally mid-sized, G segment subtype typically contains 17 amino acid residues and lacks charged residues or contain one charged residue. It is preferred that this second G segment subtype contains 14-20 amino acid residues, but it is contemplated that it may contain as few as 12 or as many as 30 amino acid residues. Without wishing to be bound by any particular theory, it is envisaged that this subtype forms coil structures. Representative G segments of this second subtype are amino acid residues 249-265, 471-488, 631-647 and 982-998 of SEQ ID NO: 5.
The third subtype of the G segment is represented by the amino acid one letter consensus sequence G(R/Q)GQG(G/R)YGQG (A/S/V)GGN (SEQ ID NO: 8). This third G segment subtype typically contains 14 amino acid residues, and is generally the shortest of the G segment subtypes. It is preferred that this third G segment subtype contains 12-17 amino acid residues, but it is contemplated that it may contain as many as 23 amino acid residues. Without wishing to be bound by any particular theory, it is envisaged that this subtype forms turn structures. Representative G segments of this third subtype are amino acid residues 57-70, 121-134, 184-197, 280-293, 343-356, 407-420, 503-516, 567-580, 662-675, 726-739, 790-803, 854-867, 918-931, 1014-1027 of SEQ ID NO: 5.
Thus, in preferred embodiments of the spidroin in the cell scaffold material, each individual G segment has at least 80%, preferably 90%, more preferably 95%, identity to an amino acid sequence selected from SEQ ID NO: 6, SEQ ID NO: 7 and SEQ ID NO: 8.
In an embodiment of the alternating sequence of A and G segments of the REP fragment, every second G segment is of the first subtype, while the remaining G segments are of the third subtype, e.g. . . . A.sub.1G.sub.shortA.sub.2G.sub.longA.sub.3G.sub.shortA.sub.4G.sub.longA.s- ub.5G.sub.short . . . . In another embodiment of the REP fragment, one G segment of the second subtype interrupts the G segment regularity via an insertion, e.g. . . . A.sub.1G.sub.shortA.sub.2G.sub.longA.sub.3G.sub.midA.sub.4G.sub.shortA.su- b.5G.sub.long . . . .
Each individual L segment represents an optional linker amino acid sequence, which may contain from 0 to 30 amino acid residues, such as from 0 to 20 amino acid residues. While this segment is optional and not critical for the function of the spider silk protein, its presence still allows for fully functional spider silk proteins and polymers thereof which form fibers, films, foams and other structures. There are also linker amino acid sequences present in the repetitive part (SEQ ID NO: 5) of the deduced amino acid sequence of the MaSp1 protein from Euprosthenops australis. In particular, the amino acid sequence of a linker segment may resemble any of the described A or G segments, but usually not sufficiently to meet their criteria as defined herein.
As shown in WO 2007/078239, a linker segment arranged at the C-terminal part of the REP fragment can be represented by the amino acid one letter consensus sequences ASASAAASAA STVANSVS (SEQ ID NO: 22) and ASAASAAA (SEQ ID NO: 23), which are rich in alanine. In fact, the second sequence can be considered to be an A segment according to the definition herein, whereas the first sequence has a high degree of similarity to A segments according to this definition. Another example of a linker segment has the one letter amino acid sequence GSAMGQGS (SEQ ID NO: 24), which is rich in glycine and has a high degree of similarity to G segments according to the definition herein. Another example of a linker segment is SASAG (SEQ ID NO: 25).
Representative L segments are amino acid residues 1-6 and 1093-1110 of SEQ ID NO: 5; and amino acid residues 159-165 of SEQ ID NO: 2, but the skilled person will readily recognize that there are many suitable alternative amino acid sequences for these segments. In one embodiment of the REP fragment, one of the L segments contains 0 amino acids, i.e. one of the L segments is void. In another embodiment of the REP fragment, both L segments contain 0 amino acids, i.e. both L segments are void. Thus, these embodiments of the REP fragments according to the invention may be schematically represented as follows: (AG).sub.nL, (AG).sub.nAL, (GA).sub.nL, (GA).sub.nGL; L(AG).sub.n, L(AG).sub.nA, L(GA).sub.n, L(GA).sub.nG; and (AG).sub.n, (AG).sub.nA, (GA).sub.n, (GA).sub.nG. Any of these REP fragments are suitable for use with any CT fragment as defined below.
The CT fragment of the spidroin in the cell scaffold material has a high degree of similarity to the C-terminal amino acid sequence of spider silk proteins. As shown in WO2007/078239, this amino acid sequence is well conserved among various species and spider silk proteins, including MaSp1 and MaSp2. A consensus sequence of the C-terminal regions of MaSp1 and MaSp2 is provided as SEQ ID NO: 4. In FIG. 9, the following MaSp proteins are aligned, denoted with GenBank accession entries where applicable:
TABLE-US-00002 TABLE 1 Spidroin CT fragments Species and spidroin Entry Euprosthenops sp MaSp1 (Pouchkina-Stantcheva*) Cthyb_Esp Euprosthenops australis MaSp1 (SEQ ID NO: 3) CTnat_Eau Argiope trifasciata MaSp1 AF350266_At1 Cyrtophora moluccensis Sp1 AY666062_Cm1 Latrodectus geometricus MaSp1 AF350273_Lg1 Latrodectus hesperus MaSp1 AY953074_Lh1 Macrothele holsti Sp1 AY666068_Mh1 Nephila clavipes MaSp1 U20329_Nc1 Nephila pilipes MaSp1 AY666076_Np1 Nephila madagascariensis MaSp1 AF350277_Nm1 Nephila senegalensis MaSp1 AF350279_Ns1 Octonoba varians Sp1 AY666057_Ov1 Psechrus sinensis Sp1 AY666064_Ps1 Tetragnatha kauaiensis MaSp1 AF350285_Tk1 Tetragnatha versicolor MaSp1 AF350286_Tv1 Araneus bicentenarius Sp2 ABU20328_Ab2 Argiope amoena MaSp2 AY365016_Aam2 Argiope aurantia MaSp2 AF350263_Aau2 Argiope trifasciata MaSp2 AF350267_At2 Gasteracantha mammosa MaSp2 AF350272_Gm2 Latrodectus geometricus MaSp2 AF350275_Lg2 Latrodectus hesperus MaSp2 AY953075_Lh2 Nephila clavipes MaSp2 AY654293_Nc2 Nephila madagascariensis MaSp2 AF350278_Nm2 Nephila senegalensis MaSp2 AF350280_Ns2 Dolomedes tenebrosus Fb1 AF350269_DtFb1 Dolomedes tenebrosus Fb2 AF350270_DtFb2 Araneus diadematus ADF-1 U47853_ADF1 Araneus diadematus ADF-2 U47854_ADF2 Araneus diadematus ADF-3 U47855_ADF3 Araneus diadematus ADF-4 U47856_ADF4 *Comparative Biochemistry and Physiology, Part B 138: 371-376 (2004)
It is not critical which specific CT fragment is present in the spider silk protein in the cell scaffold material. Thus, the CT fragment can be selected from any of the amino acid sequences shown in FIG. 9 and Table 1 or sequences with a high degree of similarity. A wide variety of C-terminal sequences can be used in the spider silk protein.
The sequence of the CT fragment has at least 50% identity, preferably at least 60%, more preferably at least 65% identity, or even at least 70% identity, to the consensus amino acid sequence SEQ ID NO: 4, which is based on the amino acid sequences of FIG. 9.
A representative CT fragment is the Euprosthenops australis sequence SEQ ID NO: 3 or amino acid residues 180-277 of SEQ ID NO: 13. Thus, in one embodiment, the CT fragment has at least 70%, such as at least 80%, such as at least 85%, preferably at least 90%, such as at least 95%, identity to SEQ ID NO: 3, amino acid residues 180-277 of SEQ ID NO: 13, or any individual amino acid sequence of FIG. 9 and Table 1. For example, the CT fragment may be identical to SEQ ID NO: 3, amino acid residues 180-277 of SEQ ID NO: 13, or any individual amino acid sequence of FIG. 9 and Table 1.
The CT fragment typically consists of from 70 to 120 amino acid residues. It is preferred that the CT fragment contains at least 70, or more than 80, preferably more than 90, amino acid residues. It is also preferred that the CT fragment contains at most 120, or less than 110 amino acid residues. A typical CT fragment contains approximately 100 amino acid residues.
The term "% identity", as used herein, is calculated as follows. The query sequence is aligned to the target sequence using the CLUSTAL W algorithm (Thompson et al, Nucleic Acids Research, 22:4673-4680 (1994)). A comparison is made over the window corresponding to the shortest of the aligned sequences. The amino acid residues at each position are compared, and the percentage of positions in the query sequence that have identical correspondences in the target sequence is reported as % identity.
The term "% similarity", as used herein, is calculated as described above for "% identity", with the exception that the hydrophobic residues Ala, Val, Phe, Pro, Leu, Ile, Trp, Met and Cys are similar; the basic residues Lys, Arg and His are similar; the acidic residues Glu and Asp are similar; and the hydrophilic, uncharged residues Gln, Asn, Ser, Thr and Tyr are similar. The remaining natural amino acid Gly is not similar to any other amino acid in this context.
Throughout this description, alternative embodiments according to the invention fulfill, instead of the specified percentage of identity, the corresponding percentage of similarity. Other alternative embodiments fulfill the specified percentage of identity as well as another, higher percentage of similarity, selected from the group of preferred percentages of identity for each sequence. For example, a sequence may be 70% similar to another sequence; or it may be 70% identical to another sequence; or it may be 70% identical and 90% similar to another sequence.
In a preferred fusion protein according to the invention, the REP-CT fragment has at least 70%, such as at least 80%, such as at least 85%, preferably at least 90%, such as at least 95%, identity to SEQ ID NO: 2 or to amino acid residues 18-277 of SEQ ID NO: 13.
In one preferred fusion protein according to the invention, the protein has at least 70%, such as at least 80%, such as at least 85%, preferably at least 90%, such as at least 95%, identity to SEQ ID NO: 13. In a particularly preferred embodiment, the fusion protein according to the invention is SEQ ID NO: 13.
The cell scaffold material according to the invention comprises a protein or peptide according to the invention displaying the cyclic RGD cell-binding motif. The cyclic RGD cell-binding motif may be exposed from short synthetic peptides or longer synthetic or recombinant proteins, which may in turn be attached to or associated with a matrix or support.
The cell scaffold material preferably comprises a protein polymer, which protein polymer in turn is containing the recombinant fusion protein according to the invention as a repeating structural unit, i.e. the protein polymer contains or consists of a polymer of the recombinant fusion protein according to the invention. This implies that the protein polymer contains or consists of an ordered plurality of fusion proteins according to the invention, typically well above 100 fusion protein units, e.g. 1000 fusion protein units or more. In a preferred embodiment, the cell scaffold material according to the invention consists of the protein polymer.
The magnitude of fusion protein units in the polymer implies that the protein polymer obtains a significant size. In a preferred embodiment, the protein polymer has a size of at least 0.01 .mu.m in at least two dimensions. Thus, the term "protein polymer" as used herein relates to fusion protein polymers having a thickness of at least 0.01 .mu.m, preferably macroscopic polymers that are visible to the human eye, i.e. having a thickness of at least 1 .mu.m. The term "protein polymer" does not encompass unstructured aggregates or precipitates. While monomers/dimers of the fusion protein are water soluble, it is understood that the protein polymers according to the invention are solid structures, i.e. not soluble in water. The protein polymers are comprising monomers of the recombinant fusion proteins according to the invention as a repeating structural unit.
The protein polymer according to the invention is typically provided in a physical form selected from the group consisting of fiber, film, coating, foam, net, fiber-mesh, sphere and capsule. According to one embodiment, it is preferable that the protein polymer according to the invention is a fiber, film or fiber-mesh. According to certain embodiments, it is preferable that the protein polymer has a three-dimensional form, such as a foam or a fiber-mesh. One preferred embodiment involves thin (typically 0.01-0.1 .mu.m thickness) coatings made of the protein polymer, which are useful for coating of stents and other medical devices. The term "foam" is comprising a porous foam with channels connecting the bubbles of the foam, sometimes to the extent that it can even be regarded as a three-dimensional net or mesh of fibers.
In a preferred embodiment, the protein polymer is in a physical form of a free-standing matrix, such as a free-standing film. This is highly useful as it allows for transfer of a cell sheet where needed, e.g. in an in vivo situation where cells need to be transferred as a cell sheet to e.g. a wound area.
The fiber, film or fiber-mesh typically has a thickness of at least 0.1 .mu.m, preferably at least 1 .mu.m. It is preferred that the fiber, film or fiber-mesh has a thickness in the range of 1-400 .mu.m, preferably 60-120 .mu.m. It is preferred that fibers have a length in the range of 0.5-300 cm, preferably 1-100 cm. Other preferred ranges are 0.5-30 cm and 1-20 cm. The fiber has the capacity to remain intact during physical manipulation, i.e. can be used for spinning, weaving, twisting, crocheting and similar procedures. The film is advantageous in that it is coherent and adheres to solid structures, e.g. the plastics in microtiter plates. This property of the film facilitates washing and regeneration procedures and is very useful for separation purposes.
The fusion protein according to the invention harbors both the desired cell-binding activity in the cyclic RGD cell-binding motif and an internal solid support activity in the REP-CT moieties, and these activities are employed in the cell scaffold material. The cell scaffold material provides a high and predictable density of the selective interaction activity towards an organic target. Losses of valuable protein moieties with selective interaction activity are minimized, since all expressed protein moieties are associated with the cell scaffold material.
The polymers which are formed from the fusion proteins according to the invention are solid structures and are useful for their physical properties, especially the useful combination of high strength, elasticity and light weight. A particularly useful feature is that the REP-CT moieties of the fusion protein are biochemically robust and suitable for regeneration, e.g. with acid, base or chaotropic agents, and suitable for heat sterilization, e.g. autoclaving at 120.degree. C. for 20 min. The polymers are also useful for their ability to support cell adherence and growth.
The properties derived from the REP-CT moities are attractive in development of new materials for medical or technical purposes. In particular, the cell scaffold materials according to the invention are useful as scaffolds for cell immobilization, cell culture, cell differentiation, tissue engineering and guided cell regeneration. They are also useful in preparative and analytical separation procedures, such as chromatography, cell capture, selection and culture, active filters, and diagnostics. The cell scaffold materials according to the invention are also useful as in medical devices, such as implants and stents, e.g. as coatings.
In a preferred embodiment, the cell scaffold material comprises a protein polymer, which is consisting of a recombinant fusion protein according to the invention as a repeating structural unit. And in a further preferred embodiment, the cell scaffold material is a protein polymer, which is consisting of a recombinant fusion protein according to the invention as a repeating structural unit.
According to a further aspect, the present invention provides a method for the cultivation of cells, comprising the steps of providing a sample of cells; applying the sample to a cell scaffold material; and maintaining the cell scaffold material having the cells applied thereto under conditions suitable for cell culture; wherein the cell scaffold material comprises a protein polymer, which is containing a recombinant protein, such as recombinant fusion protein, according to the invention as a repeating structural unit.
In a preferred embodiment, the cells are displaying .alpha.5.beta.1 integrins on their cell surface; and the cell-binding motif of the recombinant fusion protein has selectivity for .alpha.5.beta.1 integrins.
In preferred embodiments, the recombinant protein containing this cyclic RGD cell-binding motif is immobilized, such as to a solid support (i.e. not in solution), e.g. to the surface of a cell cultivation device or any type of surface where cell binding and growth is desirable. The resulting exposure of the thus immobilized cyclic RGD cell-binding motif surprisingly promotes integrin activation and cell binding to the immobilized recombinant protein containing this cyclic RGD cell-binding motif.
Recombinant fusion proteins containing this cyclic RGD cell-binding motif are particularly useful for the cultivation of cells displaying integrins on their cell surface, since the internal spidroin fragment allows the fusion protein to be brought into ordered polymers and thereby provides an internal solid support to the immobilized (i.e. not in solution) cell-binding motif. The resulting exposure of the immobilized cyclic RGD cell-binding motif surprisingly promotes integrin activation and cell binding to polymers of the recombinant fusion proteins.
Without limitation thereto, preferred cells are selected from skeletal muscle cells, endothelial cells, stem cells, fibroblasts, keratinocytes and cell lines, in particular of human origin.
Without being limited thereto, the method is useful for cultivation of endothelial cells, human mesenchymal stem cells and keratinocytes, in particular of human origin. It is particularly useful for cultivation of keratinocytes.
The cell cultivation method may advantageously be performed both in vitro and in vivo.
The present invention will in the following be further illustrated by the following non-limiting examples.
EXAMPLES
Statistics
One-way ANOVA followed by Tukey's multiple comparisons test was performed using GraphPad Prism version 6.05 for Windows, GraphPad Software, La Jolla Calif. USA, www.graphpad.com.
Example 1--Genetic Incorporation of Fibronectin-Derived Cell-Binding Motifs into Recombinant Spider Silk
The recombinant spider silk protein 4RepCT (SEQ ID NO: 2, herein denoted WT) was genetically functionalized with the RGD containing cell binding motif from the fibronectin type III module 10, in four slightly different versions (FIG. 1). In the first (FN.sub.CC-4RepCT; SEQ ID NO: 13), two amino acids flanking the RGD sequence were substituted for cysteines to enable loop formation of the motif (CTGRGDSPAC; SEQ ID NO: 10). In the second (FN.sub.SS-4RepCT; SEQ ID NO: 14), the introduced cysteines were substituted for serines to create a linear control (STGRGDSPAS; SEQ ID NO: 11). Here the amino acid serine was selected due to its resemblance to cysteine, while lacking the ability to form disulfide bonds. In the third (FN.sub.VS-4RepCT; SEQ ID NO: 15), the original sequence of the fibronectin motif (VTGRGDSPAS; SEQ ID NO: 9) was used as a linear, native control. In the fourth (RGD-4RepCT; SEQ ID NO: 16), the RGD containing peptide (SEQ ID NO 12) used in Widhe M et al., Biomaterials 34(33): 8223-8234 (2013) was used as a further linear control.
The genes encoding the functionalized variants (FN.sub.CC-4RepCT DNA-SEQ ID NO: 17; FN.sub.SS-4RepCT DNA-SEQ ID NO: 18; FN.sub.VS-4RepCT DNA-SEQ ID NO: 19; and RGD-4RepCT DNA-SEQ ID NO: 20) were made by cloning of oligos encoding the different motifs into the vector encoding 4RepCT (4RepCT DNA-SEQ ID NO: 1) and using restriction enzymes. The new sequences were introduced N-terminally to 4RepCT and confirmed by sequencing.
Example 2--Expression of Fusion Proteins Containing Fibronectin-Derived Cell-binding Motifs
Protein production in E. coli of the genetic constructs obtained in Example 1 and the following purification were done essentially as described in Hedhammar M et al., Biochemistry 47(11):3407-3417 (2008) and Hedhammar M et al., Biomacromolecules 11: 953-959 (2010).
Briefly, Escherichia coli BL21(DE3) cells (Merck Biosciences) with the expression vector for the target protein were grown at 30.degree. C. in Luria-Bertani medium containing kanamycin to an OD.sub.600 of 0.8-1 and then induced with isopropyl .beta.-D-thiogalactopyranoside and further incubated for at least 2 h. Thereafter, cells were harvested and resuspended in 20 mM Tris-HCl (pH 8.0) supplemented with lysozyme and DNase I. After complete lysis, the supernatants from centrifugation at 15,000 g were loaded onto a column packed with Ni Sepharose (GE Healthcare, Uppsala, Sweden). The column was washed extensively before elution of bound proteins with 300 mM imidazole. Fractions containing the target proteins were pooled and dialyzed against 20 mM Tris-HCl (pH 8.0). The target protein was released from the tags by proteolytic cleavage. To remove the released HisTrxHis tag, the cleavage mixture was loaded onto a second Ni Sepharose column and the flowthrough was collected. The protein content was determined from the absorbance at 280 nm.
The protein solutions obtained were purified from lipopolysaccharides (lps) as described in Hedhammar et al., Biomacromolecules 11:953-959 (2010). The protein solutions were sterile filtered (0.22 .mu.m) before being used to prepare scaffolds (film, foam, coatings or fibers).
The recombinant spider silk proteins were successfully expressed in E coli and purified with similar yield and purity as the original 4RepCT (WT; SEQ ID NO: 2).
Example 3--Fabrication of Cell Culture Matrices
After purification, the protein solutions obtained in Example 2 were filter sterilized (0.22 .mu.m) and concentrated by centrifugal filtration (Amicon Ultra, Millipore) before preparation of films, as described in Widhe M et al., Biomaterials 31(36): 9575-9585 (2010) and Widhe M et al., Biomaterials 34(33): 8223-8234 (2013).
Briefly, petri dishes were coated at room temperature with recombinant spider silk solution at a concentration of 0.3 mg/ml to generate films. Foams were made by rapid pipetting of the silk solution, and fibers were formed by gentle wagging in 15 ml tube followed by cutting into smaller pieces.
For studies of early attachment and repopulation, solutions of a protein concentration of 0.3 mg/ml were casted into films in 96- and 24 well cell culture plates respectively (Sarstedt, suspension cells) precoated with 1 pluronic to limit cell adhesion to the plastic surface. In control experiments, a reducing agent (either 5 mM Dithiothreitol, 20 mM .beta.-mercaptoethanol or 10 mM Tris(2-carboxyethyl)phosphine HCl) were added to the protein solutions directly before films were prepared.
For microscopic studies, the proteins were casted as films in chamber glass slides (LabTekII). For Alamar blue experiments, where whole well coverage is desired, the cell culture wells were coated with a covering protein solution of 0.3 mg/ml for 2 h before the liquid was removed. Films and coated surfaces were allowed to dry over night at 25.degree. C. and 30% rh under sterile conditions, then washed twice with sterile 20 mM phosphate buffer, pH 7.4, and pre-incubated with complete cell culture medium for 1 h at 37.degree. C. with 5% CO.sub.2 before cell seeding.
Free-standing films were prepared by applying of a droplet of protein solution (3 mg/ml) onto a .about.3 mm wide frame of metal wire hanging hooked up in a well of a 96-well plate and allowed to dry over night at 25.degree. C. and 30% rh under sterile conditions.
The control Bovine Fibronectin (Sigma-Aldrich F1137) was coated at recommended concentration (5 .mu.g/cm.sup.2) overnight at 37.degree. C.
It was observed that a spider silk protein functionalized with a disulfide-looped RGD motif self-assembles into stable matrices. As shown by the micrographs in FIG. 2a, the FN.sub.CC-4RepCT (SEQ ID NO: 13) protein could be presented as matrices in the format of fiber (upper), film (middle) and free standing film (lower). Scale bars in FIG. 2a indicate 500 .mu.m (upper & middle) and 1000 .mu.m (lower). Surprisingly, the FN.sub.CC-4RepCT protein could form fibers, film and foam with appeared higher stability and integrity than noted for linear RGD silk proteins (RGD-4RepCT, SEQ ID NO: 16) and WT silk proteins (4RepCT, SEQ ID NO: 2). With the FN.sub.CC-4RepCT protein, it was even possible to form free-standing films. The smooth film formats (casted and free-standing) were used in the subsequent cell adhesion experiments to rule out the effects of matrix morphology.
Example 4--Structural Analysis of Matrices
Fourier Transform Infrared Spectroscopy (FTIR) spectra of the fibers, casted films and free-standing films obtained in Example 3 were recorded on a FTIR spectrometer (Bruker). The films were placed on a crystal for measuring IR spectra by attenuated total reflection. For each spectrum 100 scans were averaged. The amide I region was further analyzed to compare the peak height of .alpha.-helical (1654 cm.sup.-1) and .beta.-sheet (1629 cm.sup.-1) structures, respectively.
FIG. 2b shows FTIR spectra of FN.sub.CC-4RepCT (SEQ ID NO: 13) silk matrices in the format of fiber (upper), film (middle) and free standing film (lower). Peaks for typical signal of .alpha.-helix and .beta.-sheet respectively are indicated by lines. Interestingly, the FTIR data in FIG. 2b show that the free-standing films have, oppositely to the casted films, completely converted to .beta.-sheet structure.
Example 5--Cell Culture
Human dermal microvascular endothelial cells (EC), (HDMEC, PromoCell GmbH, Germany) isolated from dermis from adult donor were grown in culture flasks coated with gelatin (Sigma Aldrich) in complete endothelial cell media MV, containing 5% fetal bovine serum (PromoCell GmbH, Germany).
Human mesenchymal stem cells (hMSC, Gibco) from bone marrow were grown in culture flasks coated with CELLstart (Gibco) in complete StemPro MSC serum free medium CTS (Gibco) containing 25 ng/.mu.l fibroblast growth factor .beta. (Gibco) and 2 mM Glutamax (Gibco).
Normal human epidermal keratinocytes from adult skin (NHEK-ad) were purchased from Lonza. Subculture, proliferation and migration experiments were done in KGM-Gold (Lonza), containing bovine pituitary extract, whereas adhesion experiments were performed in KGM-CD (chemically defined), supplemented with CaCl.sub.2 to give 1.2 mM Ca.sup.2+.
Keratinocyte and mesenchymal stem cell cultures, as well as experiments, were performed under serum-free conditions to avoid possible interactions between the matrices and serum proteins that potentially could give rise to increased cell adherence.
Medium was changed every 2-3 days. Cells were harvested with TrypLE (Life Technologies) when reaching a confluency of 80% for subculture or experiments. All experiments were performed at 37.degree. C. with 5% CO2 and 95% humidity.
Example 6--Effect of Matrices on Early Attachment of Adherent Cells
A. Early Attachment Assay
Cells were harvested at passage 3-8, seeded at 20 000/cm.sup.2 and allowed to adhere to the films or controls for 1 h in a cell incubator before gentle washing twice with pre-warmed phosphate buffered saline (PBS) followed by 10 min fixation with 96% ethanol. After three washings in water, cells were stained for 30 min with 0.1% Crystal Violet in H.sub.2O. Plates were dried after extensive washing in water.
Attachment and morphology of cells bound to the films obtained in Example 3 were documented by taking micrographs at 2.times. and 10.times. magnification in an inverted bright field microscope. The color was then dissolved in 40 .mu.L 20% acetic acid for 10 min, and 35 .mu.L of the solution was transferred to a 384-well plate for optical density measurement at 595 nm (TECAN Infinite M200). Wells with cells fixed without pre-washing was used as positive control. Wells with no cells were used as blank. Experiments were run in hexaplicates and repeated three times.
Determination of cell coverage area within a defined region (9.12 mm.sup.2) of the micrographs (at 2.times. magnification) was done using the software NIS elements BR (Nikon).
B. Cellular Stainings
Cells were harvested at passage 3-8, seeded at 3500/cm.sup.2 and allowed to adhere onto films for 20 min, 1 or 3 hours in chamber slides. After gentle washing, cells were fixed with 4% paraformaldehyde, permeabilized with 0.1% Triton X-100 in PBS, and blocked with 1% bovine serum albumin (BSA, AppliChem) in PBS.
Primary antibody were used at the following concentrations in 1% BSA: mouse anti human vinculin (Sigma V9131) at 9.5 .mu.g/ml, mouse-anti human beta1-integrin (activated conformation, clone HUTS-4) at 3.3 .mu.g/mL, or mouse-anti human alpha5-integrin (ligand bound conformation, clone SNAKA-51) at 2.5 .mu.g/mL, both Millipore.
Secondary antibody was AlexaFlour488 goat anti mouse IgG (H+L), cross adsorbed (Invitrogen), used at 1:500. Phalloidin-AlexaFluor594 (Life Technologies) were used at 1:40 to detect filamentous actin. DAPI was used for nuclear staining. Slides were mounted in Fluorescence mounting medium (Dako, Copenhagen).
The stained cells were analyzed using an inverted microscope (Nikon Eclipse Ti) at 4.times. and 10.times. magnification. Excitation at 563/45 nm and detection at 625/50 nm was used for red fluorescence, whereas excitation with 387/11 nm and detection at 447/60 nm was used to monitor blue fluorescence.
For microscopic analysis of cell adhesion (formation of focal adhesions and stress fibers), a confocal microscope was used (Carl Zeiss LSM 710) at 10.times. and 63.times. magnification.
Presence of stress fibers were defined as strongly stained prominent and thick f-actin filaments, and graded from 0-4, where 0=none, 1=few-some, 2=many, 3=most, and 4=all cells exhibit stress fibers.
Presence of focal adhesions were estimated as percent of cells exhibiting focal adhesions. Quality of focal adhesions were graded from 1-4 regarding presence of small and dim focal adhesion (=1p), small and distinct focal adhesions (=2p), abundant focal adhesions (=3p) and large and bright focal adhesions (=4p), and multiplied with the portion of positive cells expressing this specific type of focal adhesions (0-4, where 0.apprxeq.none of, 1.apprxeq.one fourth of, 2.apprxeq.half of, 3.apprxeq.three fourths of, and 4.apprxeq.all of the focal adhesion-positive cells.)
C. FN.sub.CC-Silk Promotes Early Attachment of Adherent Cells
First, we wanted to investigate how well adherent cells attach and spread on the FN.sub.CC-silk (FN.sub.CC-4RepCT, SEQ ID NO: 13) compared to linear RGD proteins (RGD-4RepCT, SEQ ID NO: 16) and WT silk proteins (4RepCT, SEQ ID NO: 2) obtained in Example 3. Silk films of the three different variants were prepared in cell culture plates, and human primary endothelial cells (EC), mesenchymal stem cells (MSC) or keratinocytes (KC) were allowed to adhere for 1 h before fixation and staining.
FIG. 3a shows micrographs of EC, MSC and KC after 1 h adhesion to a film of WT silk (SEQ ID NO: 2) or silk functionalized with RGD (SEQ ID NO: 16) or FN.sub.CC (SEQ ID NO: 13), followed by staining with crystal violet (10.times. magnification). Scale bar 50 .mu.m.
FIG. 3b shows the OD of crystal violet dissolved from cells adhered to different silk variants for EC (upper panel), and MSC (middle panel), and cell coverage area by KC (lower panel) within a defined region (9.12 mm.sup.2). EC and MSC: triplicates or duplicates, KC: quadruplicates. All cell types n=3. Seeding density 20 000/cm2. Boxplot: line=median, box: 25%-75%, whiskers=mean and max. Statistics: *P<0.05, **P<0.01, ****P<0.0001.
From the micrographs shown in FIG. 3a, a clear improvement of attachment is seen on the FN.sub.CC films compared to both RGD and WT for all three cell types. After imaging, the color trapped by the EC and MSC respectively was dissolved and OD was captured and used as a measure of the number of bound cells (FIG. 3b, upper and middle panel). For both cell types significantly more cells had bound to the FN.sub.CC-silk after 1 h compared to WT silk (P<0.01 for EC and P<0.05 for MSC). Significantly more EC had attached to FN.sub.CC also compared to RGD silk (p<0.01). This colorimetric method was less suitable for KC since with this cell type, some cells also attached outside the film surface, thus contributing to the OD value although not bound to the silk film. Instead, the area of cells bound to the film was measured by image analysis at 2.times. magnification, as shown in FIG. 3b (lower panel). The area of KC bound to FN.sub.CC was significantly larger than on both WT- and RGD-silk (P<0.0001).
D. Primary KC Adhere Equally Well to FN.sub.CC-Silk and Bovine Fibronectin
After seeing this positive effect of the introduced FN.sub.CC motif, we wanted to find out how well the FNCC-silk would compare to native, full length fibronectin, where the RGD is presented on a turn loop constrained by the structure. We therefore used fibronectin from bovine plasma (BFN) to coat cell culture wells, as well as naked cell culture treated plastic (TCT), on which KC can be cultured, as a control. Serum-free experimental conditions was chosen to avoid possible interactions between the matrices and serum proteins that potentially could give rise to increased cell adherence.
The results for KC after 1 h adhesion to either silk functionalized with FN.sub.CC, a bovine fibronectin coated surface (BFN) or tissue culture treated cell plastic (TCT) are presented in FIG. 4. FIG. 4a shows micrographs at 10.times. magnification after staining with crystal violet. Seeding density 40 000/cm.sup.2. Scale bar 50 .mu.m. FIG. 4b shows cell coverage area within a defined region (9.12 mm.sup.2), (Quadruplicates, n=3). Seeding density 20 000/cm2. Boxplot: line=median, box: 25%-75%, whiskers=mean and max. Statistics (vs TCT): ****P<0.0001.
When comparing cell coverage area, it was evident that KC bound equally well to the BFN and the FN.sub.CC-silk after 1 h adherence, and importantly both significantly better than TCT (P<0.0001) (FIG. 4).
Example 7--Impact of a Cysteine-Looped Conformation for RGD Presentation in FN.sub.CC-Silk
Encouraged by these results, we wanted to go further and look into the mechanism by which FN.sub.CC-silk (SEQ ID NO: 13) creates an attractive surface for the cells. For this purpose, we used two FN-silk variants where a linear RGD presentation is expected (FIG. 1a). The first variant (FN.sub.VS; SEQ ID NO: 15), contains the original sequence of the RGD-containing motif in fibronectin, to show the effect of the native flanking amino acids without influence of the loop conformation. In the second variant (FN.sub.SS; SEQ ID NO: 14), the two flanking cysteines in FN.sub.CC were substituted for serine, which resembles cysteine but lacks the --SH-group and is therefore unable to form disulfide bridges. The different FN-silk variants, as well as RGD-silk (SEQ ID NO: 16) and WT-silk (SEQ ID NO: 2), were evaluated with primary KC. Cells were analyzed both for early attachment (FIG. 5), spreading and formation of stress fibers (FIG. 6), and focal adhesions (FIG. 7). Early attachment assay and cellular stainings were performed as detailed in Example 6.
A. Early Attachment
The results for KC after 1 h adhesion to films of WT-silk (SEQ ID NO: 2) or silk functionalized with FN.sub.CC (SEQ ID NO: 13), FN.sub.VS (SEQ ID NO: 15), FN.sub.SS (SEQ ID NO: 14) or RGD (SEQ ID NO: 16) are presented in FIG. 5. FIG. 5a shows micrographs at 10.times. magnification after staining with crystal violet. Seeding density 20 000/cm.sup.2. FIG. 5b shows cell coverage area within a defined region (9.12 mm.sup.2), (Quadruplicates, n=3). Boxplot: line=median, box: 25%-75%, whiskers=mean and max. Statistics: ****P<0.0001.
In initial experiments, KC were allowed to adhere for 1 h onto films of WT, RGD- and FN-silk variants, and stained with crystal violet for detection and morphology (FIG. 5a). When pooling data from image analysis of 3 experiments (hexaplicates), FN.sub.CC-silk showed increased attachment (i.e. area covered by cells) compared to both FN.sub.SS and FN.sub.VS (P<0.0001, FIG. 5b). FN.sub.CC-silk also gave significantly higher adhesion of KC compared to RGD-silk (P<0.0001). All FN-silk variants showed significantly increased adhesion compared to WT-silk (P<0.0001).
Moreover, pooled data from 8 experiments, where the crystal violet was dissolved from the cells and the OD thereof measured in a plate reader, showed very similar results (FNCC versus FN-controls, P<0.0001), despite that cells in these experiments to some degree also adhered to the cell plastic outside the silk-films (data not shown).
B. Cell Spreading and Formation of Stress-Fibers
The results for KC after 3 h adhesion to films of WT-silk (SEQ ID NO: 2) or silk functionalized with FN.sub.CC (SEQ ID NO: 13), FN.sub.VS (SEQ ID NO: 15), FN.sub.SS (SEQ ID NO: 14) or RGD (SEQ ID NO: 16) are presented in FIG. 6. FIG. 6a shows cell coverage area, duplicates, n=4. Boxplot: line=median, box: 25%-75%, whiskers=mean and max. FIG. 6b shows stress fiber ranking (mean and standard deviation, single wells, n=3). Seeding density 3 500/cm.sup.2. Statistics: ***P<0.001, *P<0.05.
By staining for F-actin, cell spreading and formation of stress fibers in KC after 3 h adhesion were investigated (FIG. 6). The results show that FN.sub.CC film, but not FN.sub.SS or FN.sub.VS films, gave a significantly increased spreading of KC compared to RGD-film (p<0.05) and WT-film (p<0.001), when measuring total cell area in 4.times. micrographs (n=4, duplicates), (FIG. 6a). The spreading of KC on FN.sub.CC-silk was also significantly increased compared to FN.sub.SS-silk (p<0.05). The KC on RGD, FN.sub.SS- and FN.sub.VS-silk showed a higher proportion of cells with a rounded appearance, whereas on FN.sub.CC-silk most cells had a nice spread-out morphology with distinct actin filaments.
KC stained for F-actin were also analyzed for the presence of stress fibers, as an indicator of established attachment (FIG. 6b). Presence of stress fibers was defined as thick and brightly stained actin filaments (bundles), and the analysis was done by inspection at 63.times. magnification (n=3). This analysis showed similar results as the area measurement, but no statistically significant differences were found.
C. Formation of Focal Adhesions
Formation of focal adhesions within the cells was analyzed after 3 h by staining for F-actin in combination with vinculin, which is one of the major components of the focal adhesion complex. Co-staining of F-actin and vinculin is thus a sign of integrin-involved, well established binding of cells to the underlying substrate. Focal adhesions appear as yellow-greenish elongations of the F-actin filaments, often situated close to the cell membrane.
The results from the analysis of formation and characterization of focal adhesions in KC after adherence for 3 h onto films of WT-silk (SEQ ID NO: 2) or silk functionalized with FN.sub.CC (SEQ ID NO: 13), FN.sub.VS (SEQ ID NO: 15), FN.sub.SS (SEQ ID NO: 14) or RGD (SEQ ID NO: 16) is presented in FIG. 7. Slides were scanned with a confocal microscope at 10.times. for overview (FIG. 7a) and at 63.times. for details (FIG. 7b). Two types of grading of the focal adhesions in the cells were performed.
Firstly, the percentage of cells exhibiting focal adhesions was assessed by visual examination of the entire film in each well at 10.times. magnification. Pooled data from three experiments showed a significant increase in percentage of cells expressing focal adhesions on FN.sub.CC-silk, compared to RGD and WT (p<0.05), (FIG. 7a). FIG. 7a is a graph showing percentage of cells exhibiting focal adhesions (mean and standard deviation). Experiments were run in duplicates, n=3. Statistics: *P<0.05.
Secondly, since not only the abundance of cells exhibiting focal adhesions, but also the characteristics of the focal adhesions seen in the cells appeared to differ between the different silk variants, we decided to examine this further. The appearance of the focal adhesions within each positive cell was therefore evaluated according to a grading system. Briefly, grading spanned from small and dim, appearing sparsely within the cell ("subtle"), to large and bright, appearing abundantly within the cell ("prominent"). In this way we could judge the quality of the focal adhesions independently of how many of the cells on the film that exhibited these structures. The outcome of this analysis showed a tendency of more prominent focal adhesions in cells attached to FN.sub.CC-silk compared to the other silk types. FIG. 7b is a graph showing grading of the focal adhesions, independently of total number of positive cells found, (mean and standard deviation). Grading was done in single wells, n=3.
The results show that the variation of focal adhesion quality is larger in cells on RGD, FN.sub.SS and FN.sub.VS than on FN.sub.CC-silk, reflecting the presence of both prominent and subtle focal adhesions in cells on FN.sub.SS and FN.sub.VS, but almost only prominent focal adhesions on FN.sub.CC-silk. Interestingly, such prominent focal adhesions appeared as early as 20 minutes after seeding onto FN.sub.CC.
In each individual experiment, FN.sub.CC-silk, without exception, gave the most efficient adhesion of the tested films. In contrast, the attachment onto FN.sub.SS- and FN.sub.VS-silk varied from being similar to RGD-silk to being only somewhat lower than on FN.sub.CC-silk.
With the aim to further elucidate the role of the cysteine linked loop for presentation of the RGD motif, we performed experiments where reducing agents were added to the FN.sub.CC-silk solution directly before films were casted. The idea was to prevent disulphide formation in these films, generating a linear, non-looped motif. However, no differences compared to non-reduced FN.sub.CC film were detected. When considering that the films are completely dried during the production process, one can assume that the reducing agent, in the lack of buffer, can no longer prevent disulfide formation to occur. We therefore consider FN.sub.SS the most proper non-looped control accomplishable.
Example 8--Engagement of Integrin .alpha.5.beta.1 in KC Adhering to FN.sub.CC-Silk
Since the integrin .alpha.5.beta.1 is known to selectively bind to fibronectin, we decided to investigate if this integrin is involved in the binding of KC to FN.sub.CC-silk (SEQ ID NO: 13). To do this, we selected two monoclonal antibodies, developed to specifically recognize the ligand bound conformation of .alpha.5 integrin (SNAKA-51) and the activated conformation of .beta.1 integrin (HUTS-4), respectively, and used them in combination for staining of KC adhering to FN.sub.CC-silk for 3 h, in combination with staining with phalloidin for F-actin, as set out in Examples 6-7. Analysis of the cells revealed a week but distinct staining pattern resembling the pattern seen when staining for vinculin.
Example 9--Applications of FN.sub.CC-Silk
Intrigued by the findings of such excellent binding properties of FN.sub.CC-silk regarding early attachment of adherent cells, we performed a few pilot studies to get a picture of its ability to support various cell culture applications. Firstly, we wanted to evaluate the effect of the FN.sub.CC motif on cell proliferation.
A. Cell Viability Analysis with Alamar Blue
Cell growth of primary keratinocytes (NHEK) after initial low seeding density 3 500 cells/cm.sup.2 on wells coated with films of WT-silk (SEQ ID NO: 2) or FN.sub.CC-silk (SEQ ID NO: 13) in 96-well plates was monitored with Alamar Blue cell viability assay (Molecular Probes) every third day during the culture period. After 4 h incubation with Alamar blue (diluted 1:10 in cell culture medium), fluorescence intensity of 90 .mu.L supernatants from the cultures was measured with a fluorescence plate reader (CLARIOstar, BMG Labtech) using excitation at 544 nm and emission at 595 nm. Two independent experiments were performed where films were analysed in hexaplicates. Fluorescence intensities, correlating the number of living cells in each well, were plotted over time to yield growth profiles of cells seeded on silk with different cell binding motifs. The results presented in FIG. 8 show an increased level of viable cells on FN.sub.CC-silk compared to WT-silk (P<0.001 day 3 and P<0.0001 day 6; ****P<0.0001, ***P<0.001), indicating an improved ability to support cell proliferation conveyed by the FN.sub.CC motif.
B. Repopulation Assay
To evaluate the ability of the different silk variants to support repopulation of an open would field, dermal keratinocytes (NHEK) were stained with Oregon green cell trace (Life Technologies) before seeded onto films of FN.sub.CC-silk (SEQ ID NO: 13) and WT-silk (SEQ ID NO: 2) at 20 000 cells/cm.sup.2 in 24 well plates. Wound field inserts (CytoSelect.TM. Wound healing assay, Cell Biolabs) were added into the wells before cell seeding to generate a 0.9 mm wide open would field in the cell monolayer, while keeping the film intact. After 16 h the inserts were removed, and the repopulation process were followed each day and documented by inverted fluorescence microscopy at day 0 (insert removal), day 2 and 4. At day 6 cells were fixed and stained according to the assay protocol, and imaged by inverted bright field microscopy.
Thus, green-traced cells were seeded at high density into wells with inserts preventing cells to reach a defined part of the silk-film, the "wound field". After monolayer formation outside the wound field, the insert was removed, and repopulation of the gap was documented during 6 days of culture, thus allowing both migration and proliferation of cells. Keratinocytes efficiently repopulated the wound field on FN.sub.CC-silk, which was almost completely covered with cells at the end of the experiment.
C. Transferable Cell Monolayers
NHEK were harvested and traced with AMCA orange cell tracker (Life Technologies) before seeded onto free-standing films of FN.sub.CC-silk silk (SEQ ID NO: 13) mounted on metal frames at 20 000 cells/cm2. The formed monolayer was documented by inverted fluorescence microscopy.
Primary keratinocytes seeded onto such free-standing films formed a monolayer that could easily be transferred between culture wells.
Example 10--Cell Adherence to Surfaces with Immobilized Peptides
A silicon (SiO) surface is activated using an organosilane (e.g. 3-am inopropyltriethoxysilane APTES) to thereafter immobilize aminoreactive peptides (via their N-terminus) using e.g. EDC/NHS chemistry.
The peptides used for immobilization are designed with a glycine spacer, as follows:
TABLE-US-00003 1. (SEQ ID NO: 21) GGGGGCTGRGDSPAC 2. (SEQ ID NO: 22) GGGGGVTGRGDSPAS 3. (SEQ ID NO: 23) GGGGGSTGRGDSPAS 4. (SEQ ID NO: 24) GGGGGCDWRGDNQFC
Early attachment to the surfaces with immobilized peptides is analyzed using human keratinocytes (HaCAT) seeded at 20 000/cm.sup.2. The cells are then allowed to adhere for 1 h in a cell incubator before gentle washing twice with pre-warmed phosphate buffered saline (PBS) followed by 10 min fixation with 96% ethanol. After three washings in water, cells are stained for 30 min with 0.1% Crystal Violet in H.sub.2O.
Attachment and morphology of cells are documented by taking micrographs at 2.times. and 10.times. magnification in an inverted bright field microscope. The Crystal violet color is then dissolved in 40 .mu.L 20% acetic acid for 10 min, and 35 .mu.L of the solution is transferred to a 384-well plate for optical density measurement at 595 nm (TECAN Infinite M200). Cells fixed without pre-washing are used as positive control (reference).
Example 11--Cell Culture on FNcc Silk Matrices
After purification, solutions of FNcc-silk protein (SEQ ID NO: 13) were used to coat cell culture plates (Sarstedt, hydrophobic plates for suspension cells). Briefly, the protein solutions were diluted to 0.1 mg/ml in Tris buffer, and allowed to incubate at room temperature for 30 minutes before removal and wash.
Cells were harvested using trypsination (TrpLE) and seeded onto the FNcc-silk coatings at suitable cell density (3-10 000 cells/cm2). Cell growth was monitored with Alamar Blue cell viability assay (Molecular Probes) regularly (every 2-3 day). At the end point, after 7-14 days, Live/dead staining was performed. The following cell types showed positive growth profile and a majority (>80%) of viable cells at the end point:
Human Skeletal Muscle Satellite Cells
Human Dermal Microvascular Endothelial Cells
Human Mesenchymal stem cells
Mouse Mesenchymal stem cells
Human Dermal fibroblasts
HaCaT Keratinocytes
MING-m9 pancreatic cell line
SEQUENCE LISTINGS
1
601789DNAEuprosthenops australis 1ggtccgaatt caggtcaagg aggatatggt ggactaggtc aaggagggta tggacaaggt 60gcaggaagtt ctgcagccgc tgccgccgcc gcagcagccg ccgcagcagg tggacaaggt 120ggacaaggtc aaggaggata tggacaaggt tcaggaggtt ctgcagccgc cgccgccgcc 180gcagcagcag cagcagctgc agcagctgga cgaggtcaag gaggatatgg ccaaggttct 240ggaggtaatg ctgctgccgc agccgctgcc gccgccgccg ccgctgcagc agccggacag 300ggaggtcaag gtggatatgg tagacaaagc caaggtgctg gttccgctgc tgctgctgct 360gctgctgctg ccgctgctgc tgctgcagga tctggacaag gtggatacgg tggacaaggt 420caaggaggtt atggtcagag tagtgcttct gcttcagctg ctgcgtcagc tgctagtact 480gtagctaatt cggtgagtcg cctctcatcg ccttccgcag tatctcgagt ttcttcagca 540gtttctagct tggtttcaaa tggtcaagtg aatatggcag cgttacctaa tatcatttcc 600aacatttctt cttctgtcag tgcatctgct cctggtgctt ctggatgtga ggtcatagtg 660caagctctac tcgaagtcat cactgctctt gttcaaatcg ttagttcttc tagtgttgga 720tatattaatc catctgctgt gaaccaaatt actaatgttg ttgctaatgc catggctcaa 780gtaatgggc 7892263PRTEuprosthenops australisDOMAIN(1)..(158)REP fragmentDOMAIN(159)..(165)Spacer fragmentDOMAIN(166)..(263)CT fragment 2Gly Pro Asn Ser Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly Gly 1 5 10 15 Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala 20 25 30 Ala Ala Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr Gly 35 40 45 Gln Gly Ser Gly Gly Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 50 55 60 Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ser 65 70 75 80 Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 85 90 95 Ala Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Arg Gln Ser Gln Gly 100 105 110 Ala Gly Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 115 120 125 Ala Gly Ser Gly Gln Gly Gly Tyr Gly Gly Gln Gly Gln Gly Gly Tyr 130 135 140 Gly Gln Ser Ser Ala Ser Ala Ser Ala Ala Ala Ser Ala Ala Ser Thr 145 150 155 160 Val Ala Asn Ser Val Ser Arg Leu Ser Ser Pro Ser Ala Val Ser Arg 165 170 175 Val Ser Ser Ala Val Ser Ser Leu Val Ser Asn Gly Gln Val Asn Met 180 185 190 Ala Ala Leu Pro Asn Ile Ile Ser Asn Ile Ser Ser Ser Val Ser Ala 195 200 205 Ser Ala Pro Gly Ala Ser Gly Cys Glu Val Ile Val Gln Ala Leu Leu 210 215 220 Glu Val Ile Thr Ala Leu Val Gln Ile Val Ser Ser Ser Ser Val Gly 225 230 235 240 Tyr Ile Asn Pro Ser Ala Val Asn Gln Ile Thr Asn Val Val Ala Asn 245 250 255 Ala Met Ala Gln Val Met Gly 260 398PRTEuprosthenops australis 3Ser Arg Leu Ser Ser Pro Ser Ala Val Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Ser Leu Val Ser Asn Gly Gln Val Asn Met Ala Ala Leu Pro Asn 20 25 30 Ile Ile Ser Asn Ile Ser Ser Ser Val Ser Ala Ser Ala Pro Gly Ala 35 40 45 Ser Gly Cys Glu Val Ile Val Gln Ala Leu Leu Glu Val Ile Thr Ala 50 55 60 Leu Val Gln Ile Val Ser Ser Ser Ser Val Gly Tyr Ile Asn Pro Ser 65 70 75 80 Ala Val Asn Gln Ile Thr Asn Val Val Ala Asn Ala Met Ala Gln Val 85 90 95 Met Gly 4100PRTArtificial SequenceConsensus sequence derived from known MaSp1 and MaSp2 proteinsMISC_FEATURE(1)..(71)Sequence length present in known species variantsVARIANT(7)..(7)Glu 4Ser Arg Leu Ser Ser Pro Gln Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Asn Ser Ala Ala Leu Ser Asn 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Ser Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Val His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Ile Val Gly Gln Ser Val Ala Gln Ala 85 90 95 Leu Gly Glu Phe 100 51110PRTEuprosthenops australisREPEAT(7)..(19)REPEAT(20)..(42)REPEAT(43)..(56)REPEAT(57)..(70)R- EPEAT(71)..(83)REPEAT(84)..(106)REPEAT(107)..(120)REPEAT(121)..(134)REPEAT- (135)..(147)REPEAT(148)..(170)REPEAT(171)..(183)REPEAT(184)..(197)REPEAT(1- 98)..(211)REPEAT(212)..(234)REPEAT(235)..(248)REPEAT(249)..(265)REPEAT(266- )..(279)REPEAT(280)..(293)REPEAT(294)..(306)REPEAT(307)..(329)REPEAT(330).- .(342)REPEAT(343)..(356)REPEAT(357)..(370)REPEAT(371)..(393)REPEAT(394)..(- 406)REPEAT(407)..(420)REPEAT(421)..(434)REPEAT(435)..(457)REPEAT(458)..(47- 0)REPEAT(471)..(488)REPEAT(489)..(502)REPEAT(503)..(516)REPEAT(517)..(529)- REPEAT(530)..(552)REPEAT(553)..(566)REPEAT(567)..(580)REPEAT(581)..(594)RE- PEAT(595)..(617)REPEAT(618)..(630)REPEAT(631)..(647)REPEAT(648)..(661)REPE- AT(662)..(675)REPEAT(676)..(688)REPEAT(689)..(711)REPEAT(712)..(725)REPEAT- (726)..(739)REPEAT(740)..(752)REPEAT(753)..(775)REPEAT(776)..(789)REPEAT(7- 90)..(803)REPEAT(804)..(816)REPEAT(817)..(839)REPEAT(840)..(853)REPEAT(854- )..(867)REPEAT(868)..(880)REPEAT(881)..(903)REPEAT(904)..(917)REPEAT(918).- .(931)REPEAT(932)..(945)REPEAT(946)..(968)REPEAT(969)..(981)REPEAT(982)..(- 998)REPEAT(999)..(1013)REPEAT(1014)..(1027)REPEAT(1028)..(1042)REPEAT(1043- )..(1059)REPEAT(1060)..(1073)REPEAT(1074)..(1092) 5Gln Gly Ala Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 1 5 10 15 Ala Ala Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln 20 25 30 Gly Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala 35 40 45 Ala Ala Ala Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly 50 55 60 Gln Gly Ser Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 65 70 75 80 Ala Ala Ser Gly Gln Gly Gly Gln Gly Gly Gln Gly Gly Gln Gly Gln 85 90 95 Gly Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala 100 105 110 Ala Ala Ala Ala Ala Ala Ala Ala Gly Gln Gly Gln Gly Arg Tyr Gly 115 120 125 Gln Gly Ala Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 130 135 140 Ala Ala Ala Gly Gln Gly Gly Gln Gly Gly Gln Gly Gly Leu Gly Gln 145 150 155 160 Gly Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala 165 170 175 Ser Ala Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln 180 185 190 Gly Ala Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 195 200 205 Ala Ala Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln 210 215 220 Gly Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala 225 230 235 240 Ala Ala Ala Ala Ala Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly 245 250 255 Arg Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala 260 265 270 Ala Ala Ala Ala Ala Ala Ala Gly Gln Gly Gln Gly Gly Tyr Gly Gln 275 280 285 Gly Ala Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 290 295 300 Ala Ala Gly Gln Gly Gly Gln Gly Gly Gln Gly Gly Leu Gly Gln Gly 305 310 315 320 Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala 325 330 335 Ala Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly 340 345 350 Ala Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Glu Ala Ala 355 360 365 Ala Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly 370 375 380 Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala 385 390 395 400 Ala Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly 405 410 415 Ala Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 420 425 430 Ala Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly 435 440 445 Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala 450 455 460 Ala Ala Ala Ala Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly Arg 465 470 475 480 Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala 485 490 495 Ala Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly 500 505 510 Ser Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 515 520 525 Ser Gly Gln Gly Ser Gln Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly 530 535 540 Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala 545 550 555 560 Ala Ala Ala Ala Ala Ser Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly 565 570 575 Ala Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 580 585 590 Ala Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly 595 600 605 Gly Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala 610 615 620 Ala Ala Ala Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr 625 630 635 640 Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala 645 650 655 Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ser 660 665 670 Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ser 675 680 685 Gly Gln Gly Gly Gln Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr 690 695 700 Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala 705 710 715 720 Ala Ala Ala Ala Ala Gly Gln Gly Gln Gly Gly Tyr Gly Gln Gly Ala 725 730 735 Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 740 745 750 Gly Gln Gly Gly Gln Gly Gly Gln Gly Gly Leu Gly Gln Gly Gly Tyr 755 760 765 Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala 770 775 780 Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Val 785 790 795 800 Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 805 810 815 Gly Gln Gly Gly Gln Gly Gly Gln Gly Gly Leu Gly Gln Gly Gly Tyr 820 825 830 Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala 835 840 845 Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ser 850 855 860 Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ser 865 870 875 880 Gly Gln Gly Ser Gln Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr 885 890 895 Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala 900 905 910 Ala Ala Ala Ala Ser Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ala 915 920 925 Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 930 935 940 Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly Gly 945 950 955 960 Tyr Gly Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala 965 970 975 Ala Ala Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr Gly 980 985 990 Gln Gly Ser Gly Gly Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 995 1000 1005 Ala Ala Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly 1010 1015 1020 Ser Gly Gly Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 1025 1030 1035 Ala Ala Ala Ala Gly Gln Gly Gly Gln Gly Gly Tyr Gly Arg Gln 1040 1045 1050 Ser Gln Gly Ala Gly Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala 1055 1060 1065 Ala Ala Ala Ala Ala Gly Ser Gly Gln Gly Gly Tyr Gly Gly Gln 1070 1075 1080 Gly Gln Gly Gly Tyr Gly Gln Ser Ser Ala Ser Ala Ser Ala Ala 1085 1090 1095 Ala Ser Ala Ala Ser Thr Val Ala Asn Ser Val Ser 1100 1105 1110 623PRTArtificial SequenceConsensus sequence derived from internal repeats of Euprosthenops australis MaSp1VARIANT(4)..(4)SerVARIANT(8)..(8)TyrVARIANT(11)..(11)Gln 6Gly Gln Gly Gly Gln Gly Gly Gln Gly Gly Leu Gly Gln Gly Gly Tyr 1 5 10 15 Gly Gln Gly Ala Gly Ser Ser 20 717PRTArtificial SequenceConsensus sequence derived from internal repeats of Euprosthenops australis MaSp1VARIANT(9)..(9)ArgVARIANT(14)..(14)SerVARIANT(16)..(16)Gly 7Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln Gly Ala Gly Ser 1 5 10 15 Ser 814PRTArtificial SequenceConsensus sequence derived from internal repeats of Euprosthenops australis MaSp1VARIANT(2)..(2)GlnVARIANT(6)..(6)ArgVARIANT(11)..(11)SerVARIANT(11).- .(11)Val 8Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ala Gly Gly Asn 1 5 10 910PRTHomo sapiens 9Val Thr Gly Arg Gly Asp Ser Pro Ala Ser 1 5 10 1010PRTArtificial SequenceModified from Homo sapiensDISULFID(1)..(10) 10Cys Thr Gly Arg Gly Asp Ser Pro Ala Cys 1 5 10 1110PRTArtificial Sequencemodified from Homo sapiens 11Ser Thr Gly Arg Gly Asp Ser Pro Ala Ser 1 5 10 1212PRTArtificial Sequencelinear RGD cell-binding motif 12Gly Pro Asn Ser Arg Gly Asp Ala Gly Ala Ala Ser 1 5 10 13277PRTArtificial SequenceFusion proteinDISULFID(5)..(14) 13Gly Pro Asn Ser Cys Thr Gly Arg Gly Asp Ser Pro Ala Cys Gly Ser 1 5 10 15 Ala Ser Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly Gly Tyr Gly 20 25 30 Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 35 40 45 Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln Gly 50 55 60 Ser Gly Gly Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 65 70 75 80 Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ser Gly Gly 85 90 95 Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 100 105 110 Gly Gln Gly Gly Gln Gly Gly Tyr Gly Arg Gln Ser Gln Gly Ala Gly 115 120 125 Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Gly 130 135 140 Ser Gly Gln Gly Gly Tyr Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln 145 150 155 160 Ser Ser Ala Ser Ala Ser Ala Ala Ala Ser Ala Ala Ser Thr Val Ala 165 170
175 Asn Ser Val Ser Arg Leu Ser Ser Pro Ser Ala Val Ser Arg Val Ser 180 185 190 Ser Ala Val Ser Ser Leu Val Ser Asn Gly Gln Val Asn Met Ala Ala 195 200 205 Leu Pro Asn Ile Ile Ser Asn Ile Ser Ser Ser Val Ser Ala Ser Ala 210 215 220 Pro Gly Ala Ser Gly Cys Glu Val Ile Val Gln Ala Leu Leu Glu Val 225 230 235 240 Ile Thr Ala Leu Val Gln Ile Val Ser Ser Ser Ser Val Gly Tyr Ile 245 250 255 Asn Pro Ser Ala Val Asn Gln Ile Thr Asn Val Val Ala Asn Ala Met 260 265 270 Ala Gln Val Met Gly 275 14277PRTArtificial SequenceFusion protein 14Gly Pro Asn Ser Ser Thr Gly Arg Gly Asp Ser Pro Ala Ser Gly Ser 1 5 10 15 Ala Ser Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly Gly Tyr Gly 20 25 30 Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 35 40 45 Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln Gly 50 55 60 Ser Gly Gly Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 65 70 75 80 Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ser Gly Gly 85 90 95 Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 100 105 110 Gly Gln Gly Gly Gln Gly Gly Tyr Gly Arg Gln Ser Gln Gly Ala Gly 115 120 125 Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Gly 130 135 140 Ser Gly Gln Gly Gly Tyr Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln 145 150 155 160 Ser Ser Ala Ser Ala Ser Ala Ala Ala Ser Ala Ala Ser Thr Val Ala 165 170 175 Asn Ser Val Ser Arg Leu Ser Ser Pro Ser Ala Val Ser Arg Val Ser 180 185 190 Ser Ala Val Ser Ser Leu Val Ser Asn Gly Gln Val Asn Met Ala Ala 195 200 205 Leu Pro Asn Ile Ile Ser Asn Ile Ser Ser Ser Val Ser Ala Ser Ala 210 215 220 Pro Gly Ala Ser Gly Cys Glu Val Ile Val Gln Ala Leu Leu Glu Val 225 230 235 240 Ile Thr Ala Leu Val Gln Ile Val Ser Ser Ser Ser Val Gly Tyr Ile 245 250 255 Asn Pro Ser Ala Val Asn Gln Ile Thr Asn Val Val Ala Asn Ala Met 260 265 270 Ala Gln Val Met Gly 275 15277PRTArtificial SequenceFusion protein 15Gly Pro Asn Ser Val Thr Gly Arg Gly Asp Ser Pro Ala Ser Gly Ser 1 5 10 15 Ala Ser Gly Gln Gly Gly Tyr Gly Gly Leu Gly Gln Gly Gly Tyr Gly 20 25 30 Gln Gly Ala Gly Ser Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 35 40 45 Ala Ala Gly Gly Gln Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln Gly 50 55 60 Ser Gly Gly Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 65 70 75 80 Ala Ala Ala Gly Arg Gly Gln Gly Gly Tyr Gly Gln Gly Ser Gly Gly 85 90 95 Asn Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala 100 105 110 Gly Gln Gly Gly Gln Gly Gly Tyr Gly Arg Gln Ser Gln Gly Ala Gly 115 120 125 Ser Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Gly 130 135 140 Ser Gly Gln Gly Gly Tyr Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln 145 150 155 160 Ser Ser Ala Ser Ala Ser Ala Ala Ala Ser Ala Ala Ser Thr Val Ala 165 170 175 Asn Ser Val Ser Arg Leu Ser Ser Pro Ser Ala Val Ser Arg Val Ser 180 185 190 Ser Ala Val Ser Ser Leu Val Ser Asn Gly Gln Val Asn Met Ala Ala 195 200 205 Leu Pro Asn Ile Ile Ser Asn Ile Ser Ser Ser Val Ser Ala Ser Ala 210 215 220 Pro Gly Ala Ser Gly Cys Glu Val Ile Val Gln Ala Leu Leu Glu Val 225 230 235 240 Ile Thr Ala Leu Val Gln Ile Val Ser Ser Ser Ser Val Gly Tyr Ile 245 250 255 Asn Pro Ser Ala Val Asn Gln Ile Thr Asn Val Val Ala Asn Ala Met 260 265 270 Ala Gln Val Met Gly 275 16271PRTArtificial SequenceFusion protein 16Gly Pro Asn Ser Arg Gly Asp Ala Gly Ala Ala Ser Gly Gln Gly Gly 1 5 10 15 Tyr Gly Gly Leu Gly Gln Gly Gly Tyr Gly Gln Gly Ala Gly Ser Ser 20 25 30 Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Gly Gly Gln Gly 35 40 45 Gly Gln Gly Gln Gly Gly Tyr Gly Gln Gly Ser Gly Gly Ser Ala Ala 50 55 60 Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Gly Arg Gly 65 70 75 80 Gln Gly Gly Tyr Gly Gln Gly Ser Gly Gly Asn Ala Ala Ala Ala Ala 85 90 95 Ala Ala Ala Ala Ala Ala Ala Ala Ala Ala Gly Gln Gly Gly Gln Gly 100 105 110 Gly Tyr Gly Arg Gln Ser Gln Gly Ala Gly Ser Ala Ala Ala Ala Ala 115 120 125 Ala Ala Ala Ala Ala Ala Ala Ala Ala Gly Ser Gly Gln Gly Gly Tyr 130 135 140 Gly Gly Gln Gly Gln Gly Gly Tyr Gly Gln Ser Ser Ala Ser Ala Ser 145 150 155 160 Ala Ala Ala Ser Ala Ala Ser Thr Val Ala Asn Ser Val Ser Arg Leu 165 170 175 Ser Ser Pro Ser Ala Val Ser Arg Val Ser Ser Ala Val Ser Ser Leu 180 185 190 Val Ser Asn Gly Gln Val Asn Met Ala Ala Leu Pro Asn Ile Ile Ser 195 200 205 Asn Ile Ser Ser Ser Val Ser Ala Ser Ala Pro Gly Ala Ser Gly Cys 210 215 220 Glu Val Ile Val Gln Ala Leu Leu Glu Val Ile Thr Ala Leu Val Gln 225 230 235 240 Ile Val Ser Ser Ser Ser Val Gly Tyr Ile Asn Pro Ser Ala Val Asn 245 250 255 Gln Ile Thr Asn Val Val Ala Asn Ala Met Ala Gln Val Met Gly 260 265 270 17831DNAArtificial SequenceFusion protein 17ggtccgaatt catgcacagg tcgtggtgat tctccggcgt gcggatccgc tagcggtcaa 60ggaggatatg gtggactagg tcaaggaggg tatggacaag gtgcaggaag ttctgcagcc 120gctgccgccg ccgcagcagc cgccgcagca ggtggacaag gtggacaagg tcaaggagga 180tatggacaag gttcaggagg ttctgcagcc gccgccgccg ccgcagcagc agcagcagct 240gcagcagctg gacgaggtca aggaggatat ggccaaggtt ctggaggtaa tgctgctgcc 300gcagccgctg ccgccgccgc cgccgctgca gcagccggac agggaggtca aggtggatat 360ggtagacaaa gccaaggtgc tggttccgct gctgctgctg ctgctgctgc tgccgctgct 420gctgctgcag gatctggaca aggtggatac ggtggacaag gtcaaggagg ttatggtcag 480agtagtgctt ctgcttcagc tgctgcgtca gctgctagta ctgtagctaa ttcggtgagt 540cgcctctcat cgccttccgc agtatctcga gtttcttcag cagtttctag cttggtttca 600aatggtcaag tgaatatggc agcgttacct aatatcattt ccaacatttc ttcttctgtc 660agtgcatctg ctcctggtgc ttctggatgt gaggtcatag tgcaagctct actcgaagtc 720atcactgctc ttgttcaaat cgttagttct tctagtgttg gatatattaa tccatctgct 780gtgaaccaaa ttactaatgt tgttgctaat gccatggctc aagtaatggg c 83118831DNAArtificial SequenceFusion protein 18ggtccgaatt caagcacagg tcgtggtgat tctccggcga gcggatccgc tagcggtcaa 60ggaggatatg gtggactagg tcaaggaggg tatggacaag gtgcaggaag ttctgcagcc 120gctgccgccg ccgcagcagc cgccgcagca ggtggacaag gtggacaagg tcaaggagga 180tatggacaag gttcaggagg ttctgcagcc gccgccgccg ccgcagcagc agcagcagct 240gcagcagctg gacgaggtca aggaggatat ggccaaggtt ctggaggtaa tgctgctgcc 300gcagccgctg ccgccgccgc cgccgctgca gcagccggac agggaggtca aggtggatat 360ggtagacaaa gccaaggtgc tggttccgct gctgctgctg ctgctgctgc tgccgctgct 420gctgctgcag gatctggaca aggtggatac ggtggacaag gtcaaggagg ttatggtcag 480agtagtgctt ctgcttcagc tgctgcgtca gctgctagta ctgtagctaa ttcggtgagt 540cgcctctcat cgccttccgc agtatctcga gtttcttcag cagtttctag cttggtttca 600aatggtcaag tgaatatggc agcgttacct aatatcattt ccaacatttc ttcttctgtc 660agtgcatctg ctcctggtgc ttctggatgt gaggtcatag tgcaagctct actcgaagtc 720atcactgctc ttgttcaaat cgttagttct tctagtgttg gatatattaa tccatctgct 780gtgaaccaaa ttactaatgt tgttgctaat gccatggctc aagtaatggg c 83119831DNAArtificial SequenceFusion protein 19ggtccgaatt cagttacagg tcgtggtgat tctccggcga gcggatccgc tagcggtcaa 60ggaggatatg gtggactagg tcaaggaggg tatggacaag gtgcaggaag ttctgcagcc 120gctgccgccg ccgcagcagc cgccgcagca ggtggacaag gtggacaagg tcaaggagga 180tatggacaag gttcaggagg ttctgcagcc gccgccgccg ccgcagcagc agcagcagct 240gcagcagctg gacgaggtca aggaggatat ggccaaggtt ctggaggtaa tgctgctgcc 300gcagccgctg ccgccgccgc cgccgctgca gcagccggac agggaggtca aggtggatat 360ggtagacaaa gccaaggtgc tggttccgct gctgctgctg ctgctgctgc tgccgctgct 420gctgctgcag gatctggaca aggtggatac ggtggacaag gtcaaggagg ttatggtcag 480agtagtgctt ctgcttcagc tgctgcgtca gctgctagta ctgtagctaa ttcggtgagt 540cgcctctcat cgccttccgc agtatctcga gtttcttcag cagtttctag cttggtttca 600aatggtcaag tgaatatggc agcgttacct aatatcattt ccaacatttc ttcttctgtc 660agtgcatctg ctcctggtgc ttctggatgt gaggtcatag tgcaagctct actcgaagtc 720atcactgctc ttgttcaaat cgttagttct tctagtgttg gatatattaa tccatctgct 780gtgaaccaaa ttactaatgt tgttgctaat gccatggctc aagtaatggg c 83120813DNAArtificial SequenceFusion protein 20ggtccgaatt cacgcggcga tgcaggagcg gctagcggtc aaggaggata tggtggacta 60ggtcaaggag ggtatggaca aggtgcagga agttctgcag ccgctgccgc cgccgcagca 120gccgccgcag caggtggaca aggtggacaa ggtcaaggag gatatggaca aggttcagga 180ggttctgcag ccgccgccgc cgccgcagca gcagcagcag ctgcagcagc tggacgaggt 240caaggaggat atggccaagg ttctggaggt aatgctgctg ccgcagccgc tgccgccgcc 300gccgccgctg cagcagccgg acagggaggt caaggtggat atggtagaca aagccaaggt 360gctggttccg ctgctgctgc tgctgctgct gctgccgctg ctgctgctgc aggatctgga 420caaggtggat acggtggaca aggtcaagga ggttatggtc agagtagtgc ttctgcttca 480gctgctgcgt cagctgctag tactgtagct aattcggtga gtcgcctctc atcgccttcc 540gcagtatctc gagtttcttc agcagtttct agcttggttt caaatggtca agtgaatatg 600gcagcgttac ctaatatcat ttccaacatt tcttcttctg tcagtgcatc tgctcctggt 660gcttctggat gtgaggtcat agtgcaagct ctactcgaag tcatcactgc tcttgttcaa 720atcgttagtt cttctagtgt tggatatatt aatccatctg ctgtgaacca aattactaat 780gttgttgcta atgccatggc tcaagtaatg ggc 8132115PRTArtificial SequenceSynthetic peptideDISULFID(6)..(15) 21Gly Gly Gly Gly Gly Cys Thr Gly Arg Gly Asp Ser Pro Ala Cys 1 5 10 15 2215PRTArtificial SequenceSynthetic peptide 22Gly Gly Gly Gly Gly Val Thr Gly Arg Gly Asp Ser Pro Ala Ser 1 5 10 15 2315PRTArtificial SequenceSynthetic peptide 23Gly Gly Gly Gly Gly Ser Thr Gly Arg Gly Asp Ser Pro Ala Ser 1 5 10 15 2415PRTArtificial SequenceSynthetic peptideDISULFID(6)..(15) 24Gly Gly Gly Gly Gly Cys Asp Trp Arg Gly Asp Asn Gln Phe Cys 1 5 10 15 2518PRTEuprosthenops australis 25Ala Ser Ala Ser Ala Ala Ala Ser Ala Ala Ser Thr Val Ala Asn Ser 1 5 10 15 Val Ser 268PRTEuprosthenops australis 26Ala Ser Ala Ala Ser Ala Ala Ala 1 5 278PRTEuprosthenops australis 27Gly Ser Ala Met Gly Gln Gly Ser 1 5 285PRTEuprosthenops australis 28Ser Ala Ser Ala Gly 1 5 29100PRTEuprosthenops sp 29Ser Arg Leu Ser Ser Pro Glu Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Asn Ser Ala Ala Leu Ser Ser 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Gly Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Ile His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Leu Val Gly Gln Ser Val Tyr Gln Ala 85 90 95 Leu Gly Glu Phe 100 3098PRTEuprosthenops australis 30Ser Arg Leu Ser Ser Pro Ser Ala Val Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Ser Leu Val Ser Asn Gly Gln Val Asn Met Ala Ala Leu Pro Asn 20 25 30 Ile Ile Ser Asn Ile Ser Ser Ser Val Ser Ala Ser Ala Pro Gly Ala 35 40 45 Ser Gly Cys Glu Val Ile Val Gln Ala Leu Leu Glu Val Ile Thr Ala 50 55 60 Leu Val Gln Ile Val Ser Ser Ser Ser Val Gly Tyr Ile Asn Pro Ser 65 70 75 80 Ala Val Asn Gln Ile Thr Asn Val Val Ala Asn Ala Met Ala Gln Val 85 90 95 Met Gly 3199PRTArgiope trifasciata 31Ser Arg Leu Ser Ser Pro Gly Ala Ala Ser Arg Val Ser Ser Ala Val 1 5 10 15 Thr Ser Leu Val Ser Ser Gly Gly Pro Thr Asn Ser Ala Ala Leu Ser 20 25 30 Asn Thr Ile Ser Asn Val Val Ser Gln Ile Ser Ser Ser Asn Pro Gly 35 40 45 Leu Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Ile Val Ser 50 55 60 Ala Leu Val His Ile Leu Gly Ser Ala Asn Ile Gly Gln Val Asn Ser 65 70 75 80 Ser Gly Val Gly Arg Ser Ala Ser Ile Val Gly Gln Ser Ile Asn Gln 85 90 95 Ala Phe Ser 3289PRTCyrtophora moluccensis 32Ser His Leu Ser Ser Pro Glu Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Ser Thr Asn Ser Ala Ala Leu Pro Asn 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Ser Ser Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Ile His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Ile Val 85 3398PRTLatrodectus geometricus 33Ser Ala Leu Ala Ala Pro Ala Thr Ser Ala Arg Ile Ser Ser His Ala 1 5 10 15 Ser Thr Leu Leu Ser Asn Gly Pro Thr Asn Pro Ala Ser Ile Ser Asn 20 25 30 Val Ile Ser Asn Ala Val Ser Gln Ile Ser Ser Ser Asn Pro Gly Ala 35 40 45 Ser Ser Cys Asp Val Leu Val Gln Ala Leu Leu Glu Leu Val Thr Ala 50 55 60 Leu Leu Thr Ile Ile Gly Ser Ser Asn Val Gly Asn Val Asn Tyr Asp 65 70 75 80 Ser Ser Gly Gln Tyr Ala Gln Val Val Ser Gln Ser Val Gln Asn Ala 85 90 95 Phe Val 3498PRTLatrodectus hesperus 34Ser Ala Leu Ser Ala Pro Ala Thr Ser Ala Arg Ile Ser Ser His Ala 1 5 10 15 Ser Ala Leu Leu Ser Ser Gly Pro Thr Asn Pro Ala Ser Ile Ser Asn 20 25 30 Val Ile Ser Asn Ala Val Ser Gln Ile Ser Ser Ser Asn Pro Gly Ala 35 40 45 Ser Ala Cys Asp Val Leu Val Gln Ala Leu Leu Glu Leu Val Thr Ala 50 55 60 Leu Leu Thr Ile Ile Gly Ser Ser Asn Ile Gly Ser Val Asn Tyr Asp 65 70 75 80 Ser Ser Gly Gln Tyr Ala Gln Val Val Thr Gln Ser Val Gln Asn Val 85 90 95 Phe Gly 3593PRTMacrothele holsti 35Ser His Leu Ser Ser Pro Glu Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Gly Gly Ser Thr Asn Ser Ala Ala Leu Pro Asn 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Ser Ser Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Ile His Ile Leu Gly
Ser Ser Ser Ile Gly Gln Val Asp Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Ile Val Gly Gln Ser Ala 85 90 3698PRTNephila clavipes 36Ser Arg Leu Ser Ser Pro Gln Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ala Ser Gly Pro Thr Asn Ser Ala Ala Leu Ser Ser 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Gly Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Ile Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Ile Gln Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Ile Val Gly Gln Ser Val Tyr Gln Ala 85 90 95 Leu Gly 3789PRTNephila pilipes 37Ser Arg Leu Ser Ser Pro Glu Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Asn Ser Ala Ala Leu Ser Asn 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Ser Ser Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Ile His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Ile Val 85 3887PRTNephila madagascariensis 38Ser Arg Leu Ser Ser Pro Gln Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ala Ser Gly Pro Thr Asn Ser Ala Ala Leu Ser Ser 20 25 30 Thr Ile Ser Asn Ala Val Ser Gln Ile Gly Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Ile Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Ile His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln 85 3987PRTNephila senegalensis 39Ser Arg Leu Ser Ser Pro Glu Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Asn Ser Ala Ala Leu Ser Ser 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Gly Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Ile Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Val His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln 85 4089PRTOctonoba varians 40Ser Arg Leu Ser Ser Pro Glu Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Asn Ser Ala Ala Leu Ser Asn 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Ser Ser Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Pro Ile His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Ile Val 85 4189PRTPsechrus sinensis 41Ser Arg Leu Ser Ser Pro Glu Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Asn Ser Ala Ala Leu Pro Asn 20 25 30 Thr Ile Ser Asn Val Val Ser Gln Ile Ser Ser Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Ile His Ile Leu Gly Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ala Gly Gln Ala Thr Gln Ile Val 85 4288PRTTetragnatha kauaiensis 42Ser Leu Leu Ser Ser Pro Ala Ser Asn Ala Arg Ile Ser Ser Ala Val 1 5 10 15 Ser Ala Leu Ala Ser Gly Ala Ala Ser Gly Pro Gly Tyr Leu Ser Ser 20 25 30 Val Ile Ser Asn Val Val Ser Gln Val Ser Ser Asn Ser Gly Gly Leu 35 40 45 Val Gly Cys Asp Thr Leu Val Gln Ala Leu Leu Glu Ala Ala Ala Ala 50 55 60 Leu Val His Val Leu Ala Ser Ser Ser Gly Gly Gln Val Asn Leu Asn 65 70 75 80 Thr Ala Gly Tyr Thr Ser Gln Leu 85 4388PRTTetragnatha versicolor 43Ser Arg Leu Ser Ser Pro Ala Ser Asn Ala Arg Ile Ser Ser Ala Val 1 5 10 15 Ser Ala Leu Ala Ser Gly Gly Ala Ser Ser Pro Gly Tyr Leu Ser Ser 20 25 30 Ile Ile Ser Asn Val Val Ser Gln Val Ser Ser Asn Asn Asp Gly Leu 35 40 45 Ser Gly Cys Asp Thr Val Val Gln Ala Leu Leu Glu Val Ala Ala Ala 50 55 60 Leu Val His Val Leu Ala Ser Ser Asn Ile Gly Gln Val Asn Leu Asn 65 70 75 80 Thr Ala Gly Tyr Thr Ser Gln Leu 85 4489PRTAraneus bicentenarius 44Ser Arg Leu Ser Ser Ser Ala Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Ser Leu Val Ser Ser Gly Pro Thr Thr Pro Ala Ala Leu Ser Asn 20 25 30 Thr Ile Ser Ser Ala Val Ser Gln Ile Ser Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Val His Ile Leu Gly Ser Ser Ser Val Gly Gln Ile Asn Tyr Gly 65 70 75 80 Ala Ser Ala Gln Tyr Ala Gln Met Val 85 4597PRTArgiope amoena 45Arg Leu Ser Ser Pro Gln Ala Ser Ser Arg Val Ser Ser Ala Val Ser 1 5 10 15 Thr Leu Val Ser Ser Gly Pro Thr Asn Pro Ala Ser Leu Ser Asn Ala 20 25 30 Ile Gly Ser Val Val Ser Gln Val Ser Ala Ser Asn Pro Gly Leu Pro 35 40 45 Ser Cys Asp Val Leu Val Gln Ala Leu Leu Glu Ile Val Ser Ala Leu 50 55 60 Val His Ile Leu Gly Ser Ser Ser Ile Gly Gln Ile Asn Tyr Ser Ala 65 70 75 80 Ser Ser Gln Tyr Ala Arg Leu Val Gly Gln Ser Ile Ala Gln Ala Leu 85 90 95 Gly 4682PRTArgiope aurantia 46Ser Arg Leu Ser Ser Pro Gln Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Thr Leu Val Ser Ser Gly Pro Thr Asn Pro Ala Ala Leu Ser Asn 20 25 30 Ala Ile Ser Ser Val Val Ser Gln Val Ser Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Leu Val Ser Ala 50 55 60 Leu Val His Ile Leu Gly Ser Ser Ser Ile Gly Gln Ile Asn Tyr Ala 65 70 75 80 Ala Ser 4798PRTArgiope trifasciata 47Ser Arg Leu Ser Ser Pro Gln Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Thr Leu Val Ser Ser Gly Pro Thr Asn Pro Ala Ser Leu Ser Asn 20 25 30 Ala Ile Ser Ser Val Val Ser Gln Val Ser Ser Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Ile Val Ser Ala 50 55 60 Leu Val His Ile Leu Gly Ser Ser Ser Ile Gly Gln Ile Asn Tyr Ala 65 70 75 80 Ala Ser Ser Gln Tyr Ala Gln Leu Val Gly Gln Ser Leu Thr Gln Ala 85 90 95 Leu Gly 4889PRTGasteracantha mammosa 48Ser Arg Leu Ser Ser Pro Gln Ala Gly Ala Arg Val Ser Ser Ala Val 1 5 10 15 Ser Ala Leu Val Ala Ser Gly Pro Thr Ser Pro Ala Ala Val Ser Ser 20 25 30 Ala Ile Ser Asn Val Ala Ser Gln Ile Ser Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Ile Val Ser Ala 50 55 60 Leu Val Ser Ile Leu Ser Ser Ala Ser Ile Gly Gln Ile Asn Tyr Gly 65 70 75 80 Ala Ser Gly Gln Tyr Ala Ala Met Ile 85 4990PRTLatrodectus geometricus 49Ser Ala Leu Ser Ser Pro Thr Thr His Ala Arg Ile Ser Ser His Ala 1 5 10 15 Ser Thr Leu Leu Ser Ser Gly Pro Thr Asn Ser Ala Ala Ile Ser Asn 20 25 30 Val Ile Ser Asn Ala Val Ser Gln Val Ser Ala Ser Asn Pro Gly Ser 35 40 45 Ser Ser Cys Asp Val Leu Val Gln Ala Leu Leu Glu Leu Ile Thr Ala 50 55 60 Leu Ile Ser Ile Val Asp Ser Ser Asn Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ser Gly Gln Tyr Ala Gln Met Val Gly 85 90 5098PRTLatrodectus hesperus 50Ser Ala Leu Ser Ser Pro Thr Thr His Ala Arg Ile Ser Ser His Ala 1 5 10 15 Ser Thr Leu Leu Ser Ser Gly Pro Thr Asn Ala Ala Ala Leu Ser Asn 20 25 30 Val Ile Ser Asn Ala Val Ser Gln Val Ser Ala Ser Asn Pro Gly Ser 35 40 45 Ser Ser Cys Asp Val Leu Val Gln Ala Leu Leu Glu Ile Ile Thr Ala 50 55 60 Leu Ile Ser Ile Leu Asp Ser Ser Ser Val Gly Gln Val Asn Tyr Gly 65 70 75 80 Ser Ser Gly Gln Tyr Ala Gln Ile Val Gly Gln Ser Met Gln Gln Ala 85 90 95 Met Gly 5197PRTNephila clavipes 51Ser Arg Leu Ala Ser Pro Asp Ser Gly Ala Arg Val Ala Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Ser Ser Ala Ala Leu Ser Ser 20 25 30 Val Ile Ser Asn Ala Val Ser Gln Ile Gly Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Ile Gln Ala Leu Leu Glu Ile Val Ser Ala 50 55 60 Cys Val Thr Ile Leu Ser Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ala Ala Ser Gln Phe Ala Gln Val Val Gly Gln Ser Val Leu Ser Ala 85 90 95 Phe 5282PRTNephila madagascariensis 52Ser Arg Leu Ala Ser Pro Asp Ser Gly Ala Arg Val Ala Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Ser Ser Ala Ala Leu Ser Ser 20 25 30 Val Ile Ser Asn Ala Val Ser Gln Ile Gly Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Ile Gln Ala Leu Leu Glu Ile Val Ser Ala 50 55 60 Cys Val Thr Ile Leu Ser Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ala Ala 5382PRTNephila senegalensismisc_feature(35)..(35)Xaa can be any naturally occurring amino acidmisc_feature(56)..(56)Xaa can be any naturally occurring amino acid 53Ser Arg Leu Ala Ser Pro Asp Ser Gly Ala Arg Val Ala Ser Ala Val 1 5 10 15 Ser Asn Leu Val Ser Ser Gly Pro Thr Ser Ser Ala Ala Leu Ser Ser 20 25 30 Val Ile Xaa Asn Ala Val Ser Gln Ile Gly Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Ile Xaa Ala Leu Leu Glu Ile Val Ser Ala 50 55 60 Cys Val Thr Ile Leu Ser Ser Ser Ser Ile Gly Gln Val Asn Tyr Gly 65 70 75 80 Ala Ala 5471PRTDolomedes tenebrosus 54Ser Arg Leu Ser Ser Pro Glu Ala Ala Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Ser Leu Val Ser Asn Gly Gln Val Asn Val Asp Ala Leu Pro Ser 20 25 30 Ile Ile Ser Asn Leu Ser Ser Ser Ile Ser Ala Ser Ala Thr Thr Ala 35 40 45 Ser Asp Cys Glu Val Leu Val Gln Val Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Val Gln Ile Val Cys Ser 65 70 5597PRTDolomedes tenebrosus 55Ser Arg Leu Ser Ser Pro Gln Ala Ala Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Ser Leu Val Ser Asn Gly Gln Val Asn Val Ala Ala Leu Pro Ser 20 25 30 Ile Ile Ser Ser Leu Ser Ser Ser Ile Ser Ala Ser Ser Thr Ala Ala 35 40 45 Ser Asp Cys Glu Val Leu Val Gln Val Leu Leu Glu Ile Val Ser Ala 50 55 60 Leu Val Gln Ile Val Ser Ser Ala Asn Val Gly Tyr Ile Asn Pro Glu 65 70 75 80 Ala Ser Gly Ser Leu Asn Ala Val Gly Ser Ala Leu Ala Ala Ala Met 85 90 95 Gly 5693PRTAraneus diadematus 56Asn Arg Leu Ser Ser Ala Gly Ala Ala Ser Arg Val Ser Ser Asn Val 1 5 10 15 Ala Ala Ile Ala Ser Ala Gly Ala Ala Ala Leu Pro Asn Val Ile Ser 20 25 30 Asn Ile Tyr Ser Gly Val Leu Ser Ser Gly Val Ser Ser Ser Glu Ala 35 40 45 Leu Ile Gln Ala Leu Leu Glu Val Ile Ser Ala Leu Ile His Val Leu 50 55 60 Gly Ser Ala Ser Ile Gly Asn Val Ser Ser Val Gly Val Asn Ser Ala 65 70 75 80 Leu Asn Ala Val Gln Asn Ala Val Gly Ala Tyr Ala Gly 85 90 5798PRTAraneus diadematus 57Ser Arg Leu Ser Ser Pro Ser Ala Ala Ala Arg Val Ser Ser Ala Val 1 5 10 15 Ser Leu Val Ser Asn Gly Gly Pro Thr Ser Pro Ala Ala Leu Ser Ser 20 25 30 Ser Ile Ser Asn Val Val Ser Gln Ile Ser Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Ile Leu Val Gln Ala Leu Leu Glu Ile Ile Ser Ala 50 55 60 Leu Val His Ile Leu Gly Ser Ala Asn Ile Gly Pro Val Asn Ser Ser 65 70 75 80 Ser Ala Gly Gln Ser Ala Ser Ile Val Gly Gln Ser Val Tyr Arg Ala 85 90 95 Leu Ser 5898PRTAraneus diadematus 58Ser Arg Leu Ser Ser Pro Ala Ala Ser Ser Arg Val Ser Ser Ala Val 1 5 10 15 Ser Ser Leu Val Ser Ser Gly Pro Thr Lys His Ala Ala Leu Ser Asn 20 25 30 Thr Ile Ser Ser Val Val Ser Gln Val Ser Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Val Leu Val Gln Ala Leu Leu Glu Val Val Ser Ala 50 55 60 Leu Val Ser Ile Leu Gly Ser Ser Ser Ile Gly Gln Ile Asn Tyr Gly 65 70 75 80 Ala Ser Ala Gln Tyr Thr Gln Met Val Gly Gln Ser Val Ala Gln Ala 85 90 95 Leu Ala 5994PRTAraneus diadematus 59Ser Val Tyr Leu Arg Leu Gln Pro Arg Leu Glu Val Ser Ser Ala Val 1 5 10 15 Ser Ser Leu Val Ser Ser Gly Pro Thr Asn Gly Ala Ala Val Ser Gly 20 25 30 Ala Leu Asn Ser Leu Val Ser Gln Ile Ser Ala Ser Asn Pro Gly Leu 35 40 45 Ser Gly Cys Asp Ala Leu Val Gln Ala Leu Leu Glu Leu Val Ser Ala 50 55 60 Leu Val Ala Ile Leu Ser Ser Ala Ser Ile Gly Gln Val Asn Val Ser 65 70 75 80 Ser Val Ser Gln Ser Thr Gln Met Ile Ser Gln Ala Leu Ser 85 90 6010PRTHomo sapiens 60Ser Thr
Gly Arg Gly Asp Ser Pro Ala Val 1 5 10
* * * * *
References
D00001

D00002

D00003

D00004

D00005

D00006

D00007

D00008

D00009

D00010

D00011

S00001
XML
uspto.report is an independent third-party trademark research tool that is not affiliated, endorsed, or sponsored by the United States Patent and Trademark Office (USPTO) or any other governmental organization. The information provided by uspto.report is based on publicly available data at the time of writing and is intended for informational purposes only.
While we strive to provide accurate and up-to-date information, we do not guarantee the accuracy, completeness, reliability, or suitability of the information displayed on this site. The use of this site is at your own risk. Any reliance you place on such information is therefore strictly at your own risk.
All official trademark data, including owner information, should be verified by visiting the official USPTO website at www.uspto.gov. This site is not intended to replace professional legal advice and should not be used as a substitute for consulting with a legal professional who is knowledgeable about trademark law.